

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.13 + phase: 0 /pseudo
(1574 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
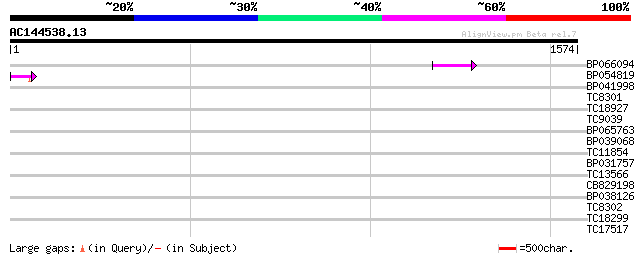
Score E
Sequences producing significant alignments: (bits) Value
BP066094 64 1e-10
BP054819 53 4e-07
BP041998 38 0.011
TC8301 35 0.12
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 34 0.21
TC9039 32 0.60
BP065763 32 0.78
BP039068 31 1.7
TC11854 similar to UP|Q84TU4 (Q84TU4) Arm repeat-containing prot... 30 3.9
BP031757 29 5.1
TC13566 weakly similar to UP|Q86RM4 (Q86RM4) Merozoite surface p... 29 5.1
CB829198 29 5.1
BP038126 29 5.1
TC8302 29 6.6
TC18299 28 8.6
TC17517 28 8.6
>BP066094
Length = 532
Score = 64.3 bits (155), Expect = 1e-10
Identities = 51/123 (41%), Positives = 64/123 (51%)
Frame = +3
Query: 1173 KVLS*MVF*MKRFMFINLLVLKI*KNQITCSN*PKLYMD*NKLQEPGMKD*AFF*LKMGF 1232
+V *MV ++ MFIN V K+ + SN* L M *+KL E GM+D A KM
Sbjct: 162 RVPF*MVTSQRKSMFINPQVXKMRRTLTIFSN*RNLSMV*SKLPEHGMRDSAHSFWKMSX 341
Query: 1233 HEEKLTPHFLEKLTKMIY*SYKCTSMISFLELQMKKCVMIFQNSWKVNLK*V*WGNLNSF 1292
K KLTKMIY* K MI +L+L + F ++NLK V*W N N+F
Sbjct: 342 *GVK*IQLSFAKLTKMIY*LCKFMLMILYLDLLIHLYAKSFLR*CRLNLKCV*WENSNTF 521
Query: 1293 WDY 1295
W Y
Sbjct: 522 WVY 530
>BP054819
Length = 537
Score = 52.8 bits (125), Expect = 4e-07
Identities = 30/79 (37%), Positives = 42/79 (52%), Gaps = 7/79 (8%)
Frame = -3
Query: 3 EGGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLN-------EK 55
EGG NRPP+ +GT+Y Y K +M FL+S DN W + G P ++ EK
Sbjct: 301 EGG*VNRPPILDGTNYDY*KVRMTAFLKSIDNKTWKAIVKGWKHPTKVEVEGTSTTVVEK 122
Query: 56 KKEDWSEEEGKRMLLNFKA 74
+E+W+ E + L N KA
Sbjct: 121 PEEEWTSVEDEAALGNSKA 65
>BP041998
Length = 485
Score = 38.1 bits (87), Expect = 0.011
Identities = 14/20 (70%), Positives = 17/20 (85%)
Frame = -2
Query: 3 EGGSSNRPPLFEGTDYYYWK 22
EGGS NRPP+ +GT+Y YWK
Sbjct: 313 EGGSVNRPPILDGTNYDYWK 254
>TC8301
Length = 1243
Score = 34.7 bits (78), Expect = 0.12
Identities = 28/93 (30%), Positives = 44/93 (47%)
Frame = +2
Query: 11 PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEEEGKRMLL 70
P F G+++ WK K++ L+ DN + + + + D ++++ K M
Sbjct: 521 PKFSGSNFSLWKLKIKAILRK-DNCLPAI--------------DGRPADITDDKWKEMDD 655
Query: 71 NFKAKLFLTMALSREEYDRVQECKNAKEIWDTL 103
N A L L +A S + E K AKEIWDTL
Sbjct: 656 NAVANLHLAVADS--VLSSIAEKKTAKEIWDTL 748
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 33.9 bits (76), Expect = 0.21
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = -2
Query: 281 TELDKSQVTCYGCYKLGHYKNECP 304
T+ D S++ C+ C K GH+ N CP
Sbjct: 356 TQSDTSEIVCHRCSKKGHFANRCP 285
>TC9039
Length = 1218
Score = 32.3 bits (72), Expect = 0.60
Identities = 30/120 (25%), Positives = 54/120 (45%)
Frame = +2
Query: 184 AKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELD 243
A LK L+LE L L H VLL ++ K I+ +E + ++F Q++E
Sbjct: 539 ASKLKALKLE-LSDDLLIHLVLLSLPAQFSQFK-ISYNCPKEKWSLNELISFCVQEEERL 712
Query: 244 EEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNEC 303
+++ ++ ++ + +++ +N A K + K TCY C GH K +C
Sbjct: 713 KQERKESAHFVSTSKDKGKRKKTVEPKNEAADAPAPKKQ--KEDDTCYFCNVSGHMKKKC 886
>BP065763
Length = 537
Score = 32.0 bits (71), Expect = 0.78
Identities = 16/42 (38%), Positives = 19/42 (45%), Gaps = 7/42 (16%)
Frame = -1
Query: 273 PARKENAKTELDK-------SQVTCYGCYKLGHYKNECPLNK 307
P RKE K + K + CY C K GH+K CP K
Sbjct: 453 PGRKEQKKDQALKVKEGRIHKEHVCYFCKKAGHFKKYCPKRK 328
>BP039068
Length = 467
Score = 30.8 bits (68), Expect = 1.7
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 3/40 (7%)
Frame = +3
Query: 287 QVTCYGCYKLGHYKNECP---LNKRKSSNLQTNQKSFMTT 323
Q C C + GH N+CP L R + + + NQ+S T+
Sbjct: 303 QTVCMNCQETGHASNDCPSLTLRSRNAQHPRMNQQSGETS 422
>TC11854 similar to UP|Q84TU4 (Q84TU4) Arm repeat-containing protein, partial
(9%)
Length = 615
Score = 29.6 bits (65), Expect = 3.9
Identities = 16/64 (25%), Positives = 31/64 (48%)
Frame = -2
Query: 1257 SMISFLELQMKKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSFVKKNISKIFL 1316
S + + +K +F SW L+* +SFW+++ + + ++KN S IFL
Sbjct: 215 SSFTLIPKMTEKLWCLFSCSWGTRLR*CN*RRYSSFWEHQGTKLITMKA*LQKNASSIFL 36
Query: 1317 KNIK 1320
++
Sbjct: 35 SPLR 24
>BP031757
Length = 507
Score = 29.3 bits (64), Expect = 5.1
Identities = 19/66 (28%), Positives = 31/66 (46%)
Frame = +2
Query: 49 DEDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHE 108
+ED +K ED+ EEE K+ + K +EE + Q ++AK I D ++ +
Sbjct: 164 EEDNEDKDGEDYEEEEEKKKKKKSRKKKEEEEEEEQEEPESPQPPEDAKPIGDPVRFSGK 343
Query: 109 GTSHVK 114
G K
Sbjct: 344 GRGRKK 361
>TC13566 weakly similar to UP|Q86RM4 (Q86RM4) Merozoite surface protein 3
alpha (Fragment), partial (3%)
Length = 509
Score = 29.3 bits (64), Expect = 5.1
Identities = 19/73 (26%), Positives = 28/73 (38%), Gaps = 13/73 (17%)
Frame = -1
Query: 519 KFKNNYTNFHKKETLNNTNHQGPKKIWVPKALLTSTAGMS-------------HNSQEKA 565
K+K N E T+H ++ + LTST GM+ H SQ++
Sbjct: 419 KYKKCLNNGDNAEWGGGTDHPRVDRLDPQQRYLTSTGGMNALKVP*PPSHPPQHGSQQRQ 240
Query: 566 LVLGQWMLKTYDW 578
GQW + W
Sbjct: 239 SSGGQWAASAFGW 201
>CB829198
Length = 562
Score = 29.3 bits (64), Expect = 5.1
Identities = 17/69 (24%), Positives = 32/69 (45%)
Frame = +3
Query: 397 EKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMGS 456
E+ D ++ QHS+SKE +E + + +N K K N +S ++ +++
Sbjct: 255 EEPQNDDVEESQHSDSKEETEEEEDEDGDNENNKPKRAKAAKKKKNKERSNKSDNEVVKK 434
Query: 457 QVSVFDKAC 465
+ F K C
Sbjct: 435 KGGGFCKLC 461
>BP038126
Length = 548
Score = 29.3 bits (64), Expect = 5.1
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = +2
Query: 289 TCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPE 328
TC+ C++ GH+ PLN + L ++ S T P+
Sbjct: 284 TCFKCHQQGHWSWFWPLNSPNKNPLSSSSSSHSPTLHSPQ 403
>TC8302
Length = 494
Score = 28.9 bits (63), Expect = 6.6
Identities = 29/119 (24%), Positives = 51/119 (42%), Gaps = 4/119 (3%)
Frame = +3
Query: 196 IGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQEL----EMNFDNQQQELDEEDHQDQI 251
IG + E+LL+ S +I +K N + L E+ + Q ++ED D
Sbjct: 105 IGENERAELLLESLPDSYDQLVINIKNNNIVNHLPLMMLSEVLLEEDSQR*NKEDR*DS- 281
Query: 252 ILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKS 310
++ ++ + R ++K +RK K + CY C + H K +C NK+ S
Sbjct: 282 ---SKPMEALTMTRGRSK----SRK--------KINLKCYHCGQR*HLKKDCWFNKKSS 413
>TC18299
Length = 569
Score = 28.5 bits (62), Expect = 8.6
Identities = 19/71 (26%), Positives = 33/71 (45%)
Frame = +3
Query: 350 VNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQH 409
V +D CE+ LF +CE R + EL++ E LE++ KDL + +
Sbjct: 111 VAVDKNKICEENIQLF-----------MKCEEFRKKETELKQTVENLEEDLKDLPRKCRQ 257
Query: 410 SESKEVSENKN 420
+ V++ K+
Sbjct: 258 LSAFLVAKKKH 290
>TC17517
Length = 505
Score = 28.5 bits (62), Expect = 8.6
Identities = 24/79 (30%), Positives = 39/79 (48%), Gaps = 7/79 (8%)
Frame = +3
Query: 461 FDKACIGFKTNQKQKLYEN--FFIPKKNKNTCERY---KCTYCEKYGHLEPFC--FRKRK 513
F +CI F Q+QKL +N F I ++ Y + T E G + FC ++K+K
Sbjct: 138 FYHSCITFIPQQQQKLKKNKLFIIIHIIPDSTTHYNQ*QKTRIEDDGKVHVFCTTWKKKK 317
Query: 514 RLASQKFKNNYTNFHKKET 532
+ Q+ +TN K++T
Sbjct: 318 KKKRQEPXGRFTNNWKEKT 374
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.355 0.157 0.539
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,450,163
Number of Sequences: 28460
Number of extensions: 345447
Number of successful extensions: 4204
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 3093
Number of HSP's successfully gapped in prelim test: 118
Number of HSP's that attempted gapping in prelim test: 1019
Number of HSP's gapped (non-prelim): 3346
length of query: 1574
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1472
effective length of database: 1,994,680
effective search space: 2936168960
effective search space used: 2936168960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144538.13


