

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
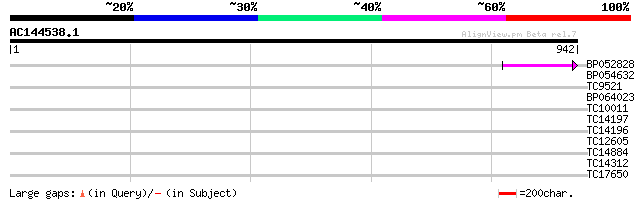
Query= AC144538.1 - phase: 0 /pseudo
(942 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP052828 87 9e-18
BP054632 31 1.0
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 30 1.4
BP064023 30 2.3
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 30 2.3
TC14197 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein,... 29 4.0
TC14196 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein,... 29 4.0
TC12605 similar to UP|AAR99871 (AAR99871) Strubbelig receptor fa... 28 6.7
TC14884 homologue to UP|Q93XV7 (Q93XV7) Hydroxypyruvate reductas... 28 6.7
TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%) 28 6.7
TC17650 28 6.7
>BP052828
Length = 523
Score = 87.4 bits (215), Expect = 9e-18
Identities = 43/127 (33%), Positives = 74/127 (57%), Gaps = 3/127 (2%)
Frame = +2
Query: 819 PCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLAGYIEDI 878
PC IG + + LGA ++++P S++ + G L+ T +I++LA+RS G +ED+
Sbjct: 71 PCTIGDNKYENCMLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDV 250
Query: 879 PVKIEGIYIPTDFVVVDIE---EDHDVPIILGRPFLATAGAIIDVQSGRIVFQASDAMIG 935
V++ + P DF ++D+E + PIILGRPF+ TA IDV +G + + D +
Sbjct: 251 LVQVNELIFPADFYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAK 430
Query: 936 FELENVM 942
F +++ M
Sbjct: 431 FNIDDAM 451
>BP054632
Length = 514
Score = 30.8 bits (68), Expect = 1.0
Identities = 23/87 (26%), Positives = 42/87 (47%), Gaps = 4/87 (4%)
Frame = +3
Query: 579 NQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLENCMMNQNKQLQELKNQTGSLNDS 638
NQ+ YK P +Y Q + Y++ LE+ + +++Q L+++ LN+
Sbjct: 261 NQDAYKKP--NYVQISVESYSHLTG----------LEDQVKTYEEKVQTLEDEINELNEK 404
Query: 639 LS----KLNTKVDSIATHTKMLETQIS 661
LS ++NTK + H K+ E +S
Sbjct: 405 LSAANSEINTKEGLVKQHAKVAEEAVS 485
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/28 (46%), Positives = 14/28 (49%)
Frame = +1
Query: 536 PSPASPTCEVCGITGHTGVDCQLGSAAN 563
P P S C CGI GH DC+ G N
Sbjct: 370 PPPGSGRCFNCGIDGHWARDCKAGDWKN 453
>BP064023
Length = 442
Score = 29.6 bits (65), Expect = 2.3
Identities = 17/46 (36%), Positives = 22/46 (46%)
Frame = -1
Query: 297 KGLGADLVRKYEDSETKIRLPKVRSLARPFSGSPTEDPNLHISSFL 342
K LG RK E ++ R PK+ SL +P G L + SFL
Sbjct: 325 KDLGTQTARKIEGRKSPERAPKIASLEKPGLGPSGTALTLWLRSFL 188
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 29.6 bits (65), Expect = 2.3
Identities = 12/28 (42%), Positives = 14/28 (49%)
Frame = +2
Query: 536 PSPASPTCEVCGITGHTGVDCQLGSAAN 563
P P S C CG+ GH DC+ G N
Sbjct: 350 PPPGSGRCFNCGLDGHWARDCKAGDWKN 433
>TC14197 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein, partial
(95%)
Length = 976
Score = 28.9 bits (63), Expect = 4.0
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -2
Query: 812 DPGSFAIPCVIGKETVDKALCHLGASVSLLPL 843
DPG +P V+G +DK LC+L S S L L
Sbjct: 348 DPGVVGVPGVVGVGGMDKFLCNLLPSPSSLML 253
>TC14196 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein, partial
(95%)
Length = 1040
Score = 28.9 bits (63), Expect = 4.0
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -2
Query: 812 DPGSFAIPCVIGKETVDKALCHLGASVSLLPL 843
DPG +P V+G +DK LC+L S S L L
Sbjct: 205 DPGVVGVPGVVGVGGMDKFLCNLLPSPSSLML 110
>TC12605 similar to UP|AAR99871 (AAR99871) Strubbelig receptor family 3,
partial (22%)
Length = 680
Score = 28.1 bits (61), Expect = 6.7
Identities = 12/19 (63%), Positives = 16/19 (84%)
Frame = +1
Query: 106 LREQLVTRRSVSFLSW*AV 124
+RE L+TRR +SFL+W* V
Sbjct: 247 IREPLLTRRMMSFLNW*IV 303
>TC14884 homologue to UP|Q93XV7 (Q93XV7) Hydroxypyruvate reductase ,
partial (66%)
Length = 921
Score = 28.1 bits (61), Expect = 6.7
Identities = 12/36 (33%), Positives = 21/36 (58%)
Frame = -1
Query: 646 VDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQT 681
++S A+ + ++S+ A +S +TPGVFP T
Sbjct: 585 MNSSASTILLAAAKVSEAASSAVVSVRTPGVFPTAT 478
>TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%)
Length = 622
Score = 28.1 bits (61), Expect = 6.7
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = +1
Query: 583 YKNPQGSYGQTAPPGYTNN 601
Y PQGSYGQ APP Y N+
Sbjct: 199 YGAPQGSYGQ-APPSYGNS 252
>TC17650
Length = 826
Score = 28.1 bits (61), Expect = 6.7
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = +1
Query: 410 FNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIV 443
FN + * R +EM + C H ++KWLI+
Sbjct: 163 FNSHSMFQYFST*RRMEEMGKFCRHAVMQKWLIL 264
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.332 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,049,407
Number of Sequences: 28460
Number of extensions: 200669
Number of successful extensions: 1264
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1252
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1263
length of query: 942
length of database: 4,897,600
effective HSP length: 99
effective length of query: 843
effective length of database: 2,080,060
effective search space: 1753490580
effective search space used: 1753490580
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144538.1


