

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
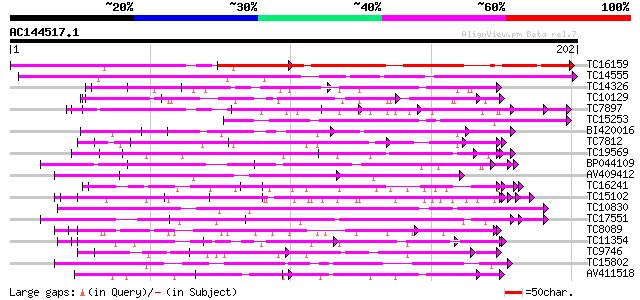
Query= AC144517.1 + phase: 0
(202 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, parti... 100 1e-22
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 90 2e-19
TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific... 80 2e-16
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 78 9e-16
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 78 1e-15
TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pro... 72 7e-14
BI420016 72 9e-14
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 71 1e-13
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 70 2e-13
BP044109 70 2e-13
AV409412 70 3e-13
TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%) 69 4e-13
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 69 6e-13
TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, ... 69 7e-13
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 69 7e-13
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 69 7e-13
TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II ... 69 7e-13
TC9746 68 1e-12
TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber... 68 1e-12
AV411518 67 2e-12
>TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, partial (19%)
Length = 605
Score = 100 bits (250), Expect = 1e-22
Identities = 63/128 (49%), Positives = 83/128 (64%), Gaps = 1/128 (0%)
Frame = +2
Query: 75 SPAS-PVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSP 133
SPAS P +P TP++ S S +PSPA++PS A +P AS P PV S PSPSP+ +NSP
Sbjct: 113 SPASSPALSPTKTPVAASPRKSLSPSPAVSPS--AQTPAASPPTPVESGPSPSPATVNSP 286
Query: 134 PSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAV 193
P + + APA+TPSSIS+PP++A S PS NGA +N F+VAG +AAV
Sbjct: 287 P------ASSDAPAVTPSSISSPPSEAQS---------PS--QNGAALNRFSVAG-SAAV 412
Query: 194 VFIAALIL 201
VF A L++
Sbjct: 413 VFAAVLLM 436
Score = 82.8 bits (203), Expect = 4e-17
Identities = 55/116 (47%), Positives = 67/116 (57%), Gaps = 15/116 (12%)
Frame = +2
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATP---------------SPAGSPLAISPAV 45
MAS V ML+A+LLVSS A+S +SSP +P SPA SP A +PA
Sbjct: 47 MASSTAVFMLLAALLVSSAVAQSPASSPALSPTKTPVAASPRKSLSPSPAVSPSAQTPAA 226
Query: 46 SSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPA 101
S P P+ + PSPSP VN SPP +S APAVTP SIS+PPS+A SP+
Sbjct: 227 SPPTPVESGPSPSPATVN---SPPASS--------DAPAVTPSSISSPPSEAQSPS 361
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 90.1 bits (222), Expect = 2e-19
Identities = 65/214 (30%), Positives = 106/214 (49%), Gaps = 15/214 (7%)
Frame = +2
Query: 4 YMVVLMLVASLLVSSTFARSLSSSPVA--------TPSPAGSPLAISPAVSSPVPLMNAP 55
++V+++ + ++++S A+S S+SP TP P + + PA SSP P+ ++P
Sbjct: 131 HVVLVLGLICIVIASVGAQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSP 310
Query: 56 SPSPFAVNYPTSPPPASFGSP--ASPVAAPAVTPLS-----ISTPPSQAPSPAIAPSTSA 108
P+ +SPPP S P ++P APA TP S PP+ P PA P
Sbjct: 311 PPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATP---- 478
Query: 109 NSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPP 168
P A +PVP T SP+P+P+ + S SP P +PA AP + S+S PS S P
Sbjct: 479 --PPALTPVPAT-SPAPAPAKVKS-KSPAPAPAPALAPVV---SLSPSEAPGPSLSSLSP 637
Query: 169 TKAPSTPTNGATMNGFNVAGYAAAVVFIAALILM 202
+P+ + +++ ++VF +A + +
Sbjct: 638 ALSPAASDDSGVEKSWSMQKMIGSLVFGSAFLYL 739
>TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific
extensin-like protein precursor (PELP), partial (11%)
Length = 630
Score = 80.5 bits (197), Expect = 2e-16
Identities = 55/151 (36%), Positives = 72/151 (47%), Gaps = 5/151 (3%)
Frame = +2
Query: 30 ATPSPAGSPLAISPAVSSPVP-LMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPL 88
A PSP P +PAV+ P P L+ AP+P P SPP P P A +P
Sbjct: 143 APPSPVKPPTPSTPAVTVPTPPLVKAPTPPPQPPVKAPSPPIVK-SPPYFPPPVKAPSPP 319
Query: 89 SISTPPSQAPSPAIAPSTSANSPVASSP-VPVTSSP---SPSPSAINSPPSPPPHASPAA 144
+ +PP +AP+P I S + P +P P+ SP +P+P + SPP PP P
Sbjct: 320 IVKSPPVKAPTPPIVKSPPSYPPPVKAPSPPIVKSPPVKAPTPPIVKSPPYYPP---PVK 490
Query: 145 APAITPSSISTPPTKAPSSISTPPTKAPSTP 175
AP TP + TPP P + P PSTP
Sbjct: 491 AP--TPPQVKTPPPH-PPVVKPPVAPIPSTP 574
Score = 67.4 bits (163), Expect = 2e-12
Identities = 51/144 (35%), Positives = 66/144 (45%), Gaps = 11/144 (7%)
Frame = +2
Query: 43 PAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAI 102
P+V +P + P+PS AV PT P V AP P PP +APSP I
Sbjct: 131 PSVPAPPSPVKPPTPSTPAVTVPTPPL----------VKAPTPPP----QPPVKAPSPPI 268
Query: 103 APSTSANSPVASSP-VPVTSSP---SPSPSAINSPPSPPPHASPAAAPAITPSSISTPPT 158
S P +P P+ SP +P+P + SPPS PP P AP +P + +PP
Sbjct: 269 VKSPPYFPPPVKAPSPPIVKSPPVKAPTPPIVKSPPSYPP---PVKAP--SPPIVKSPPV 433
Query: 159 KAPS-------SISTPPTKAPSTP 175
KAP+ PP KAP+ P
Sbjct: 434 KAPTPPIVKSPPYYPPPVKAPTPP 505
Score = 60.5 bits (145), Expect = 2e-10
Identities = 46/124 (37%), Positives = 62/124 (49%), Gaps = 10/124 (8%)
Frame = +2
Query: 62 VNY--PTSPPPASFGSPASPVAAPAVTPLSISTPPS-QAPSPAIAPSTSANSPVASSPVP 118
VNY P+ P P S P +P + PAVT + TPP +AP+P P A SP P
Sbjct: 116 VNYAIPSVPAPPSPVKPPTP-STPAVT---VPTPPLVKAPTPPPQPPVKAPSP------P 265
Query: 119 VTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPT-----KAPSS--ISTPPTKA 171
+ SP P + +P P + P AP TP + +PP+ KAPS + +PP KA
Sbjct: 266 IVKSPPYFPPPVKAPSPPIVKSPPVKAP--TPPIVKSPPSYPPPVKAPSPPIVKSPPVKA 439
Query: 172 PSTP 175
P+ P
Sbjct: 440 PTPP 451
Score = 43.5 bits (101), Expect = 3e-05
Identities = 33/102 (32%), Positives = 43/102 (41%), Gaps = 14/102 (13%)
Frame = +2
Query: 28 PVATPSPAGSPLAISPAVSSPVP-LMNAPSPSPFAVNYPTSP-----------PPASFGS 75
PV PSP P+ SP V +P P ++ +P P V P+ P PP
Sbjct: 296 PVKAPSP---PIVKSPPVKAPTPPIVKSPPSYPPPVKAPSPPIVKSPPVKAPTPPIVKSP 466
Query: 76 PASPVAAPAVTPLSISTPPSQAP--SPAIAPSTSANSPVASS 115
P P A TP + TPP P P +AP S +P+ S
Sbjct: 467 PYYPPPVKAPTPPQVKTPPPHPPVVKPPVAPIPS--TPIVKS 586
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 78.2 bits (191), Expect = 9e-16
Identities = 58/161 (36%), Positives = 72/161 (44%), Gaps = 20/161 (12%)
Frame = +1
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAP-SPSPFAVNYPTSPPPASFGSPASP------- 79
P TPSP+ SP + +P P + P +P+P PTS PP P +P
Sbjct: 85 PPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPTPPKTPSPTSQPPYIPLPPKTPSPISQPP 264
Query: 80 -VAAPAVTPLSISTPP-----SQAPSPAIAPSTSANSPVASSPV---PVTSSPSPSPSAI 130
V P TP IS PP + PSP P + P SP+ P T +P +PS
Sbjct: 265 HVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPPYTPTPPKTPSPT 444
Query: 131 NSP---PSPPPHASPAAAPAITPSSISTPPTKAPSSISTPP 168
N P PSPP +SP + P TPS P K PS S PP
Sbjct: 445 NQPPHIPSPPNSSSPTSQPPNTPS-----PPKTPSPTSQPP 552
Score = 73.9 bits (180), Expect = 2e-14
Identities = 54/164 (32%), Positives = 75/164 (44%), Gaps = 14/164 (8%)
Frame = +1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPS-PSPFAVNYPTSPPPASFGSPASPVAAPAV 85
+P TPSP P +P P + P+ PSP P + PP + P +P +P
Sbjct: 34 TPPKTPSPGNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPTPPKTP--SPTS 207
Query: 86 TPLSISTPPSQAPSPAIAPSTSANSPVASSPV---PVTSSPSPSPSAINSP---PSPPPH 139
P I PP + PSP P P SP+ P T +P +PS + P P+PP
Sbjct: 208 QPPYIPLPP-KTPSPISQPPHVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNT 384
Query: 140 ASPAAAPAITPSSISTP-PTKAPSSISTPP------TKAPSTPT 176
SP + P TP+ TP PT P I +PP ++ P+TP+
Sbjct: 385 PSPISQPPYTPTPPKTPSPTNQPPHIPSPPNSSSPTSQPPNTPS 516
Score = 67.0 bits (162), Expect = 2e-12
Identities = 48/129 (37%), Positives = 59/129 (45%), Gaps = 7/129 (5%)
Frame = +1
Query: 55 PSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVAS 114
PSP + PT P S G+ P TP +PP+ PSP APS + + P
Sbjct: 1 PSPVYTPPSIPTPPKTPSPGNQPPHTPIPPKTPSPSYSPPN-VPSPPKAPSPNNHPPYTP 177
Query: 115 SPVPVTSSPSPSPSAINSPP------SPPPHASPAAAPAITPSSISTPP-TKAPSSISTP 167
+P P T SP+ P I PP S PPH P TPS IS PP T P +P
Sbjct: 178 TP-PKTPSPTSQPPYIPLPPKTPSPISQPPH---VPTPPSTPSPISQPPYTPTPPKTPSP 345
Query: 168 PTKAPSTPT 176
++ P TPT
Sbjct: 346 TSQPPHTPT 372
Score = 54.7 bits (130), Expect = 1e-08
Identities = 42/116 (36%), Positives = 55/116 (47%), Gaps = 2/116 (1%)
Frame = +1
Query: 26 SSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAV 85
S P P+P +P IS +P P PSP+ + PT P + SP++ P
Sbjct: 253 SQPPHVPTPPSTPSPISQPPYTPTP-PKTPSPTSQPPHTPTPP------NTPSPISQPPY 411
Query: 86 TPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSP--SPSPSAINSPPSPPPH 139
TP TPP + PSP P + P +SSP TS P +PSP SP S PP+
Sbjct: 412 TP----TPP-KTPSPTNQPPHIPSPPNSSSP---TSQPPNTPSPPKTPSPTSQPPN 555
Score = 37.7 bits (86), Expect = 0.001
Identities = 24/73 (32%), Positives = 34/73 (45%)
Frame = +1
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA 82
S +S P TP+P +P IS +P P PSP+ + P+ P +S S +
Sbjct: 340 SPTSQPPHTPTPPNTPSPISQPPYTPTP-PKTPSPTNQPPHIPSPPNSSSPTSQPPNTPS 516
Query: 83 PAVTPLSISTPPS 95
P TP S PP+
Sbjct: 517 PPKTPSPTSQPPN 555
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 77.8 bits (190), Expect = 1e-15
Identities = 72/203 (35%), Positives = 96/203 (46%), Gaps = 29/203 (14%)
Frame = +3
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVT 86
SP T P P SP SP P + PS SP PT+ P+ G+P S + +PA
Sbjct: 510 SPRGTQPPPSPPPKSSP---SPSPKASPPSGSP-----PTAGEPSPAGTPPSVLPSPAGA 665
Query: 87 PL-----SISTPPSQA----PSPAIAPSTSANSPVASSP--VPVTSSP-------SPSP- 127
P+ S TPPS + PSP AP ++ +P A SP P ++SP SPSP
Sbjct: 666 PVPAGGPSAGTPPSASTSPSPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTATSPSPA 845
Query: 128 --SAINSPPSP-------PPHAS-PAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
S PP+P PP AS P +P+ TPSS+ P PSS S T AP
Sbjct: 846 GDSPAGGPPAPSPTTGDTPPSASGPDVSPSGTPSSL---PADTPSSSSNNST-APPPGNG 1013
Query: 178 GATMNGFNVAGYAAAVVFIAALI 200
G+++ + Y+ +V AAL+
Sbjct: 1014GSSVVPSSFLHYSVTIVVGAALL 1082
Score = 57.0 bits (136), Expect = 2e-09
Identities = 39/119 (32%), Positives = 60/119 (49%), Gaps = 6/119 (5%)
Frame = +3
Query: 80 VAAPAVTPLSISTPPSQAPSPA-IAPSTSANSPVASSPVPVTSSPSPSPSAINSP-PSPP 137
V +P T S PP +PSP+ A S + P A P P + PS PS +P P+
Sbjct: 504 VISPRGTQPPPSPPPKSSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGG 683
Query: 138 PHAS--PAAAPAITPSSISTPPTKAPSSISTPPTKAP--STPTNGATMNGFNVAGYAAA 192
P A P+A+ + +PS +S PP +P+ + P+ +P ++P G T + AG + A
Sbjct: 684 PSAGTPPSASTSPSPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTATSPSPAGDSPA 860
Score = 56.2 bits (134), Expect = 4e-09
Identities = 45/125 (36%), Positives = 61/125 (48%)
Frame = +3
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA 82
S S+SP +PSP +P A SPA + P + S SP TSP PA SPA
Sbjct: 705 SASTSP--SPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTATSPSPAG-DSPAG--GP 869
Query: 83 PAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP 142
PA +P + TPPS A P ++PS + +S A +PS S+ NS PP +
Sbjct: 870 PAPSPTTGDTPPS-ASGPDVSPSGTPSSLPAD---------TPSSSSNNSTAPPPGNGGS 1019
Query: 143 AAAPA 147
+ P+
Sbjct: 1020SVVPS 1034
Score = 50.1 bits (118), Expect = 3e-07
Identities = 40/106 (37%), Positives = 52/106 (48%)
Frame = +3
Query: 21 ARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPV 80
A S SSSP + SPAG P A SP+ + P P+PSP + +PP AS P
Sbjct: 771 AESPSSSPTSN-SPAGGPTATSPSPAGDSPAGGPPAPSPTTGD---TPPSAS-----GPD 923
Query: 81 AAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPS 126
+P+ TP S+ PA PS+S+N+ A P SS PS
Sbjct: 924 VSPSGTPSSL---------PADTPSSSSNNSTAPPPGNGGSSVVPS 1034
Score = 40.4 bits (93), Expect = 2e-04
Identities = 30/75 (40%), Positives = 34/75 (45%), Gaps = 6/75 (8%)
Frame = +3
Query: 112 VASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAIT---PSSISTPPTKAPSSISTP- 167
V SP PSP P S PSP P ASP + T PS TPP+ PS P
Sbjct: 501 VVISPRGTQPPPSPPPK---SSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPV 671
Query: 168 PTKAPS--TPTNGAT 180
P PS TP + +T
Sbjct: 672 PAGGPSAGTPPSAST 716
>TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(ATAGP4) (AT5G10430/F12B17_220), partial (42%)
Length = 575
Score = 72.0 bits (175), Expect = 7e-14
Identities = 48/134 (35%), Positives = 68/134 (49%), Gaps = 10/134 (7%)
Frame = +3
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSP 136
A+P AP TP PP A +PA P+T P A++P P T+ P+ +P+ SPP+P
Sbjct: 159 AAPTQAPTTTP-----PPPPAAAPAPPPATP---PPAATPAPTTTPPAATPAPSASPPAP 314
Query: 137 PPHASPAAAPAITPSSISTPPTKAPSSISTPPTKA----------PSTPTNGATMNGFNV 186
P ASP AP TPS+ S+PP PS ++ P A P P+ ++N +
Sbjct: 315 TPTASPTGAP--TPSA-SSPPAPIPSGPASGPGPAAGPGPNSADTPPPPSAAFSLNQPII 485
Query: 187 AGYAAAVVFIAALI 200
AG A F A ++
Sbjct: 486 AGTALVATFFAMVL 527
>BI420016
Length = 559
Score = 71.6 bits (174), Expect = 9e-14
Identities = 46/127 (36%), Positives = 69/127 (54%), Gaps = 3/127 (2%)
Frame = +2
Query: 57 PSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSP 116
P P + PT+ + G SP PA TP +PPS SP + P S SP ++P
Sbjct: 155 PEPQTLKKPTTQ--CTHGGNPSPKPVPASTP----SPPS---SPTLTPPPSPPSPTPNAP 307
Query: 117 VPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPT---KAPSSISTPPTKAPS 173
+ +S PSP P+ + PPSP P+ +P+A + TPSS TP + +PSS+ +P + +P+
Sbjct: 308 LSPSSPPSP-PTPPSPPPSPAPNPTPSATGSTTPSSPLTPTSASLSSPSSLFSPASTSPA 484
Query: 174 TPTNGAT 180
+ T T
Sbjct: 485 STTATTT 505
Score = 65.9 bits (159), Expect = 5e-12
Identities = 44/118 (37%), Positives = 59/118 (49%), Gaps = 1/118 (0%)
Frame = +2
Query: 48 PVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTS 107
PVP PS + P SPP + +P SP + P S TPPS PSPA P+ S
Sbjct: 221 PVPASTPSPPSSPTLTPPPSPPSPTPNAPLSPSSPP-----SPPTPPSPPPSPAPNPTPS 385
Query: 108 A-NSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSI 164
A S SSP+ TS+ SPS++ SP S P ++ A T + +P +K SS+
Sbjct: 386 ATGSTTPSSPLTPTSASLSSPSSLFSPASTSPASTTATTTTTTMIRLRSPASKLFSSL 559
Score = 45.4 bits (106), Expect = 7e-06
Identities = 32/102 (31%), Positives = 48/102 (46%), Gaps = 11/102 (10%)
Frame = +2
Query: 26 SSPVATPSPA------GSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFG----- 74
SSP TP P+ +PL+ S S P P PSP+P +P P++ G
Sbjct: 251 SSPTLTPPPSPPSPTPNAPLSPSSPPSPPTPPSPPPSPAP-------NPTPSATGSTTPS 409
Query: 75 SPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSP 116
SP +P +A +P S+ +P S +P+ A +T+ SP
Sbjct: 410 SPLTPTSASLSSPSSLFSPASTSPASTTATTTTTTMIRLRSP 535
Score = 27.3 bits (59), Expect = 1.9
Identities = 17/58 (29%), Positives = 25/58 (42%)
Frame = +2
Query: 125 PSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
P P + P + H P+ P STP PSS + P +P +PT A ++
Sbjct: 155 PEPQTLKKPTTQCTHGGN---PSPKPVPASTP--SPPSSPTLTPPPSPPSPTPNAPLS 313
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 70.9 bits (172), Expect = 1e-13
Identities = 49/132 (37%), Positives = 71/132 (53%)
Frame = +1
Query: 44 AVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIA 103
A S VPL P+ +P + +S PP+S SPA +P + + PP A SP
Sbjct: 85 ATSPSVPLRRQPTQTP*GQPWLSSSPPSSSSSPAPVPESPTTNSPTTAPPPPPASSP--- 255
Query: 104 PSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSS 163
P + +SP +SSP P S+P+ SP+A ++PPSP +S +P+ SS S+ PSS
Sbjct: 256 PPSPTHSP-SSSPSP--SAPT-SPAATSTPPSPSASSSAVTSPSSAVSSTSSLSFSDPSS 423
Query: 164 ISTPPTKAPSTP 175
PP +PS+P
Sbjct: 424 ---PPCSSPSSP 450
Score = 61.2 bits (147), Expect = 1e-10
Identities = 40/101 (39%), Positives = 56/101 (54%), Gaps = 2/101 (1%)
Frame = +1
Query: 79 PVAAPAVTPLSISTPPSQAPSPAIAP-STSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
P P P S+PPS + SPA P S + NSP + P P SSP PSP+ +SP S P
Sbjct: 118 PTQTP*GQPWLSSSPPSSSSSPAPVPESPTTNSPTTAPPPPPASSPPPSPT--HSPSSSP 291
Query: 138 PHASPAAAPAITPSSISTPPT-KAPSSISTPPTKAPSTPTN 177
++P + P++ STPP+ A SS T P+ A S+ ++
Sbjct: 292 SPSAPTS-----PAATSTPPSPSASSSAVTSPSSAVSSTSS 399
Score = 57.8 bits (138), Expect = 1e-09
Identities = 52/138 (37%), Positives = 67/138 (47%), Gaps = 5/138 (3%)
Frame = +1
Query: 31 TPSPAGSP-LAISPAVSSPVPLMNAPSPSPFAVNYPTS---PPPASFGSPASPVAAPAVT 86
T +P G P L+ SP SS P AP P N PT+ PPPAS
Sbjct: 121 TQTP*GQPWLSSSPPSSSSSP---APVPESPTTNSPTTAPPPPPAS-------------- 249
Query: 87 PLSISTPPSQAPSPAIAPSTSA-NSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAA 145
S PPS SP+ +PS SA SP A+S P SPS S SA+ SP S + ++
Sbjct: 250 ----SPPPSPTHSPSSSPSPSAPTSPAATSTPP---SPSASSSAVTSPSS-----AVSST 393
Query: 146 PAITPSSISTPPTKAPSS 163
+++ S S+PP +PSS
Sbjct: 394 SSLSFSDPSSPPCSSPSS 447
Score = 55.1 bits (131), Expect = 8e-09
Identities = 44/113 (38%), Positives = 60/113 (52%), Gaps = 1/113 (0%)
Frame = +1
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS-PVAAP 83
SS P ++ SPA P+ SP +SP AP P P +SPPP+ SP+S P +
Sbjct: 154 SSPPSSSSSPA--PVPESPTTNSPT---TAPPPPP-----ASSPPPSPTHSPSSSPSPSA 303
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSP 136
+P + STPPS PS + + TS +S V+S+ S PS P +SP SP
Sbjct: 304 PTSPAATSTPPS--PSASSSAVTSPSSAVSSTSSLSFSDPSSPP--CSSPSSP 450
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 70.5 bits (171), Expect = 2e-13
Identities = 55/149 (36%), Positives = 70/149 (46%), Gaps = 5/149 (3%)
Frame = +1
Query: 33 SPAGSPLAISPAVSS----PVPLMNAPSPSPF-AVNYPTSPPPASFGSPASPVAAPAVTP 87
SP G L S + SS P P P+ SP + T+PPP+ +P AP+ +
Sbjct: 19 SPCGLLLHTSSSSSSSYSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTST 198
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPA 147
S ++ S+ PSP PSTS SP +SPSPSP S P ASP+ +P
Sbjct: 199 KSPTSSASEEPSPP-NPSTSTTQQPKHSPTSPPASPSPSPP---STKPPTAMASPSTSPL 366
Query: 148 ITPSSISTPPTKAPSSISTPPTKAPSTPT 176
S TPP APS+ STPP S T
Sbjct: 367 SATKSHPTPPA-APSASSTPPPTTTSLKT 450
Score = 68.2 bits (165), Expect = 1e-12
Identities = 49/147 (33%), Positives = 69/147 (46%), Gaps = 12/147 (8%)
Frame = +1
Query: 41 ISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQ---- 96
+ P ++SP L+ S S + +Y PPP+ P +P+ +P S +TPP
Sbjct: 1 VLPNLTSPCGLLLHTSSSSSS-SYSPPPPPSQ-----QPTHSPSTSPTSTTTPPPSPTKV 162
Query: 97 APSPAIAPSTSANSPVASS------PVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITP 150
+P APSTS SP +S+ P P TS+ + SPP+ P + P+ P
Sbjct: 163 TENPPTAPSTSTKSPTSSASEEPSPPNPSTSTTQQPKHSPTSPPASPSPSPPSTKPPTAM 342
Query: 151 SSISTPPTKAPSSISTPPT--KAPSTP 175
+S ST P A S TPP A STP
Sbjct: 343 ASPSTSPLSATKSHPTPPAAPSASSTP 423
Score = 67.8 bits (164), Expect = 1e-12
Identities = 49/146 (33%), Positives = 70/146 (47%), Gaps = 1/146 (0%)
Frame = +1
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA 82
S SSS + P P SP+ S PSP+ N PT+P ++ +S
Sbjct: 49 SSSSSSYSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASEE 228
Query: 83 PAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSP-SAINSPPSPPPHAS 141
P+ S ST SP P++ + SP ++ P +SPS SP SA S P+PP A+
Sbjct: 229 PSPPNPSTSTTQQPKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPP--AA 402
Query: 142 PAAAPAITPSSISTPPTKAPSSISTP 167
P+A+ P++ S +PSS STP
Sbjct: 403 PSASSTPPPTTTSLKTMSSPSS-STP 477
Score = 48.9 bits (115), Expect = 6e-07
Identities = 27/70 (38%), Positives = 39/70 (55%)
Frame = +1
Query: 111 PVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTK 170
P +SP + S S S+ SPP PPP P +P+ +P+S +TPP +P+ ++ P
Sbjct: 7 PNLTSPCGLLLHTSSSSSSSYSPP-PPPSQQPTHSPSTSPTSTTTPPP-SPTKVTENPPT 180
Query: 171 APSTPTNGAT 180
APST T T
Sbjct: 181 APSTSTKSPT 210
>BP044109
Length = 497
Score = 70.5 bits (171), Expect = 2e-13
Identities = 55/163 (33%), Positives = 82/163 (49%), Gaps = 1/163 (0%)
Frame = +2
Query: 12 ASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA 71
A+ +S + + SLSSS PSP P + ++P NA S S PT+PPP
Sbjct: 47 ATTSLSHSSSTSLSSSHFF-PSPPPPPPPPNKNKTTPSHASNATSKSTPRTQTPTTPPP- 220
Query: 72 SFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAIN 131
SP ++P P + +T PS SP PS+S++ P + P P +SS +P++
Sbjct: 221 ------SPTSSPKPPPSTFATKPS--TSPPETPSSSSHGPAPTLPSPPSSS---TPTSTP 367
Query: 132 SPPSPPPHASPAAAPAITPS-SISTPPTKAPSSISTPPTKAPS 173
SPPSPP ++ + P TP+ + S P + +S S T PS
Sbjct: 368 SPPSPPNGSTLPSPPRATPTPAPSLPAARRMTSASLCSTSKPS 496
Score = 53.5 bits (127), Expect = 2e-08
Identities = 41/140 (29%), Positives = 61/140 (43%), Gaps = 1/140 (0%)
Frame = +2
Query: 43 PAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAI 102
P ++ L ++ S S + ++ SPPP P P T PS A +
Sbjct: 35 PPHTATTSLSHSSSTSLSSSHFFPSPPP------------PPPPPNKNKTTPSHASNATS 178
Query: 103 APSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPS 162
+ +P P P TSSP P PS + PS P +P+++ S P +P
Sbjct: 179 KSTPRTQTPTTPPPSP-TSSPKPPPSTFATKPSTSPPETPSSS-----SHGPAPTLPSPP 340
Query: 163 SISTP-PTKAPSTPTNGATM 181
S STP T +P +P NG+T+
Sbjct: 341 SSSTPTSTPSPPSPPNGSTL 400
Score = 42.7 bits (99), Expect = 4e-05
Identities = 30/101 (29%), Positives = 42/101 (40%), Gaps = 9/101 (8%)
Frame = +2
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAA-- 145
L + P Q P S S +S + S SP P P N + P HAS A +
Sbjct: 5 LQSTRPEQQQPPHTATTSLSHSSSTSLSSSHFFPSPPPPPPPPNKNKTTPSHASNATSKS 184
Query: 146 ------PAITPSSISTPPTKAPSSIST-PPTKAPSTPTNGA 179
P P S ++ P PS+ +T P T P TP++ +
Sbjct: 185 TPRTQTPTTPPPSPTSSPKPPPSTFATKPSTSPPETPSSSS 307
Score = 25.8 bits (55), Expect = 5.4
Identities = 30/123 (24%), Positives = 43/123 (34%), Gaps = 19/123 (15%)
Frame = +3
Query: 47 SPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS-----------PVAAPAVTPLSISTPPS 95
SP N P+P +++ T PPP S +S P T ++
Sbjct: 9 SPPGQNNNNHPTPPLLHFLTLPPPHSPPLTSSLLRHHHHHHRTRTRQPHHTLPTLPPNQH 188
Query: 96 QAPSPAI--------APSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPA 147
+AP P + PS + P +P + P P P P PP H P P
Sbjct: 189 RAPKPRLHLRRLLPHHPSL-LHQPSPQNPPLLPRKPPPPPHMARLQPFPPLH-PPQLPPR 362
Query: 148 ITP 150
+ P
Sbjct: 363 LRP 371
>AV409412
Length = 426
Score = 70.1 bits (170), Expect = 3e-13
Identities = 44/134 (32%), Positives = 66/134 (48%), Gaps = 11/134 (8%)
Frame = +1
Query: 40 AISPAVSSPVPLMNAPSPSPFAVNY-------PTSPPPASFGSPASPVAAPAVTPLSIST 92
++ +S+ P + P P P + + PPP+S +P+ P P S
Sbjct: 37 SLQKPISAAAPPCHPPPPPPPPYFWCCHWLF*ASPPPPSSA*TPS*PT-----NPASTLA 201
Query: 93 PPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPS----AINSPPSPPPHASPAAAPAI 148
PP+ SP++ PS + + SP P S PSPSPS + SPP+PPP + P++ P
Sbjct: 202 PPTIPSSPSLPPSRNPSLLHHLSPSPSLSLPSPSPSPSTSSATSPPTPPPSSPPSSPPPP 381
Query: 149 TPSSISTPPTKAPS 162
P S S+PP+ PS
Sbjct: 382 PPPSTSSPPSTTPS 423
Score = 42.0 bits (97), Expect = 7e-05
Identities = 33/102 (32%), Positives = 49/102 (47%)
Frame = +1
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
SS S ++P +T +P P + S S L++ SPSP +++ P+ P S S
Sbjct: 151 SSA*TPS*PTNPASTLAPPTIPSSPSLPPSRNPSLLHHLSPSP-SLSLPSPSPSPSTSSA 327
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVP 118
SP P S+PPS P P P S +SP +++P P
Sbjct: 328 TSPPTPPP------SSPPSSPPPP---PPPSTSSPPSTTPSP 426
Score = 36.6 bits (83), Expect = 0.003
Identities = 26/95 (27%), Positives = 46/95 (48%)
Frame = +1
Query: 31 TPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSI 90
TPS +P + + P PS +P +++ + P S SP+ P+ + S
Sbjct: 163 TPS*PTNPASTLAPPTIPSSPSLPPSRNPSLLHHLSPSPSLSLPSPS-----PSPSTSSA 327
Query: 91 STPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSP 125
++PP+ P P+ PS+ P S+ P +++PSP
Sbjct: 328 TSPPT--PPPSSPPSSPPPPPPPSTSSPPSTTPSP 426
>TC16241 weakly similar to UP|Q91TH4 (Q91TH4) T122, partial (7%)
Length = 799
Score = 69.3 bits (168), Expect = 4e-13
Identities = 63/174 (36%), Positives = 82/174 (46%), Gaps = 19/174 (10%)
Frame = +1
Query: 29 VATPSPAGSPLAISPAVSSPVPLMNAPSPS--------PFAVNYPTSPPP-ASFGSPASP 79
+ATP SP + P +P N PSPS P N PTS PP A +P
Sbjct: 70 IATPPNTHSPTSQPPNTQTPA---NTPSPSSQPPYLSTPPKSNSPTSQPPVAQTPRAVTP 240
Query: 80 VAAPAVTPLSIS-----TPPSQAPS-----PAIAPSTSANSPVASSPVPVTSSPSPSPSA 129
+++P+ +P S S PPS APS PA+ P AS+PVP T+SP PS
Sbjct: 241 ISSPSSSPTSNSPYPSTNPPSLAPSISTSPPALTP--------ASTPVPATASP-PS--- 384
Query: 130 INSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNG 183
SP + P AS AA +PSS + T+ PSS + T A + AT +G
Sbjct: 385 -QSPSAESPGASLPAATTPSPSSTTPRSTQTPSS-NNETTPAQRNGASSATPSG 540
Score = 66.6 bits (161), Expect = 3e-12
Identities = 48/159 (30%), Positives = 80/159 (50%), Gaps = 4/159 (2%)
Frame = +1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASP---VAAP 83
+P+ PSP P P +++P P ++P+ P P + PA+ SP+S ++ P
Sbjct: 28 APIKPPSPTSQP----PYIATP-PNTHSPTSQP-----PNTQTPANTPSPSSQPPYLSTP 177
Query: 84 AVTPLSISTPPSQAPSPAIAP-STSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP 142
+ S PP A+ P S+ ++SP ++SP P T+ PS +PS SPP
Sbjct: 178 PKSNSPTSQPPVAQTPRAVTPISSPSSSPTSNSPYPSTNPPSLAPSISTSPP-------- 333
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATM 181
A+TP+S P T ++PP+++PS + GA++
Sbjct: 334 ----ALTPASTPVPAT------ASPPSQSPSAESPGASL 420
Score = 53.9 bits (128), Expect = 2e-08
Identities = 39/115 (33%), Positives = 55/115 (46%), Gaps = 22/115 (19%)
Frame = +1
Query: 83 PAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSA----------INS 132
P TP I P + P IA + +SP + P T + +PSPS+ NS
Sbjct: 13 PPYTPAPIKPPSPTSQPPYIATPPNTHSPTSQPPNTQTPANTPSPSSQPPYLSTPPKSNS 192
Query: 133 PPSPPPHA------SPAAAPAITPSSIS-----TPPTKAPSSISTPPTKAP-STP 175
P S PP A +P ++P+ +P+S S PP+ APS ++PP P STP
Sbjct: 193 PTSQPPVAQTPRAVTPISSPSSSPTSNSPYPSTNPPSLAPSISTSPPALTPASTP 357
Score = 47.4 bits (111), Expect = 2e-06
Identities = 32/108 (29%), Positives = 52/108 (47%), Gaps = 21/108 (19%)
Frame = +1
Query: 91 STPPSQAPSPAIAPSTSANSPVASSP---------VPVTSSPSPSPSAINSPP---SPPP 138
++ P P+P PS ++ P ++P P T +P+ +PS + PP +PP
Sbjct: 4 TSQPPYTPAPIKPPSPTSQPPYIATPPNTHSPTSQPPNTQTPANTPSPSSQPPYLSTPPK 183
Query: 139 HASPAAAP-------AITP--SSISTPPTKAPSSISTPPTKAPSTPTN 177
SP + P A+TP S S+P + +P + PP+ APS T+
Sbjct: 184 SNSPTSQPPVAQTPRAVTPISSPSSSPTSNSPYPSTNPPSLAPSISTS 327
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 68.9 bits (167), Expect = 6e-13
Identities = 54/167 (32%), Positives = 84/167 (49%), Gaps = 9/167 (5%)
Frame = +1
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAP----SPSPFAVNYPTSPPPASFGSPASPV 80
SS P +PA P S S+P P ++ S + ++ P+ PPPAS PA+
Sbjct: 253 SSEPTCLLAPASPPTPKSSHSSTPPPSRSSSTTPKSKNATSLTAPSCPPPAS---PAATA 423
Query: 81 AAPAVTPLSISTPPSQAPSPAIAPSTSANS-PVASSPVPVTSSPSPSPSAINSPPSP--- 136
AP+ TP PP+ + P+ S+ N+ P + S P PSP+ S I++P SP
Sbjct: 424 PAPSKTP---PNPPTNSSKPSSCKSSLPNTTPSSLSSAP----PSPTTSPISTPSSPENQ 582
Query: 137 -PPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
P ++P +AP + SS S+ P P S+ TKAP++ G+ ++
Sbjct: 583 EKPASTPPSAPPRSTSSSSSSPINTPRKPSS-ATKAPASSPPGSELS 720
Score = 68.9 bits (167), Expect = 6e-13
Identities = 60/182 (32%), Positives = 84/182 (45%), Gaps = 11/182 (6%)
Frame = +1
Query: 17 SSTFARSLSSSPVATPS--PAGSPLAISPAVSS--PVPLMNAPSPSPFAVNYPTSPPPAS 72
SST +S +++ + PS P SP A +PA S P P N+ PS + P + P +
Sbjct: 340 SSTTPKSKNATSLTAPSCPPPASPAATAPAPSKTPPNPPTNSSKPSSCKSSLPNTTPSSL 519
Query: 73 FGSPASPVAAPAVTPLS-------ISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSP 125
+P SP +P TP S STPPS P STS++S SSP+ PS
Sbjct: 520 SSAPPSPTTSPISTPSSPENQEKPASTPPSAPPR-----STSSSS---SSPINTPRKPS- 672
Query: 126 SPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFN 185
SA +P S PP + + +P S S + P S+ P+ + NGAT N
Sbjct: 673 --SATKAPASSPPGSELSLHH--SPPSASRKSSTTPKISSSEPSHSRRGQ*NGATRNSAR 840
Query: 186 VA 187
V+
Sbjct: 841 VS 846
Score = 64.3 bits (155), Expect = 1e-11
Identities = 59/173 (34%), Positives = 84/173 (48%), Gaps = 14/173 (8%)
Frame = +1
Query: 19 TFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS 78
T A + S++ AT S + +P S P L+ AP+ P + +S PP S S +
Sbjct: 175 TTATTTSTTKAATNSSPPESRSTTPPSSEPTCLL-APASPPTPKSSHSSTPPPSRSSSTT 351
Query: 79 PVAAPAVTPLSISTPPSQAPSP-AIAPS-TSANSPVASS-PVPVTSS-PSPSPSAINSPP 134
P + A + + S PP +P+ A APS T N P SS P SS P+ +PS+++S P
Sbjct: 352 PKSKNATSLTAPSCPPPASPAATAPAPSKTPPNPPTNSSKPSSCKSSLPNTTPSSLSSAP 531
Query: 135 SPPPHASPAAAPAITPSS-------ISTPPTKAP---SSISTPPTKAPSTPTN 177
SP +P TPSS STPP+ P SS S+ P P P++
Sbjct: 532 -----PSPTTSPISTPSSPENQEKPASTPPSAPPRSTSSSSSSPINTPRKPSS 675
Score = 50.4 bits (119), Expect = 2e-07
Identities = 45/150 (30%), Positives = 71/150 (47%), Gaps = 28/150 (18%)
Frame = +1
Query: 56 SPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSI--STPPSQAPSPAIAPST--SANSP 111
+PS A P+S P + + ++ AA +P +TPPS P+ +AP++ + S
Sbjct: 130 NPSLEATASPSSEPCTTATTTSTTKAATNSSPPESRSTTPPSSEPTCLLAPASPPTPKSS 309
Query: 112 VASSPVPVTSSP----SPSPSAINSPPSPPPHASPAAAPAIT------PSSISTP----- 156
+S+P P SS S + +++ +P PPP + A APA + P++ S P
Sbjct: 310 HSSTPPPSRSSSTTPKSKNATSLTAPSCPPPASPAATAPAPSKTPPNPPTNSSKPSSCKS 489
Query: 157 --PTKAPSSIS-------TPPTKAPSTPTN 177
P PSS+S T P PS+P N
Sbjct: 490 SLPNTTPSSLSSAPPSPTTSPISTPSSPEN 579
Score = 48.5 bits (114), Expect = 8e-07
Identities = 31/113 (27%), Positives = 57/113 (50%), Gaps = 5/113 (4%)
Frame = +1
Query: 72 SFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAIN 131
S + ASP + P T + ST ++A + + P + + +P +S P + + SP +
Sbjct: 136 SLEATASPSSEPCTTATTTST--TKAATNSSPPESRSTTPPSSEPTCLLAPASPPTPKSS 309
Query: 132 SPPSPPPHASPAAAP----AITPSSISTPPTKAPSSISTPPTKA-PSTPTNGA 179
+PPP S + P A + ++ S PP +P++ + P+K P+ PTN +
Sbjct: 310 HSSTPPPSRSSSTTPKSKNATSLTAPSCPPPASPAATAPAPSKTPPNPPTNSS 468
Score = 39.7 bits (91), Expect = 4e-04
Identities = 24/86 (27%), Positives = 40/86 (45%)
Frame = +1
Query: 91 STPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITP 150
+TP + A + PS A + +S P ++ S + +A NS P +P ++
Sbjct: 94 TTPKTAAATFL*NPSLEATASPSSEPCTTATTTSTTKAATNSSPPESRSTTPPSSEPTCL 273
Query: 151 SSISTPPTKAPSSISTPPTKAPSTPT 176
+ ++PPT S STPP S+ T
Sbjct: 274 LAPASPPTPKSSHSSTPPPSRSSSTT 351
>TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, partial
(24%)
Length = 702
Score = 68.6 bits (166), Expect = 7e-13
Identities = 59/179 (32%), Positives = 87/179 (47%), Gaps = 4/179 (2%)
Frame = +3
Query: 18 STFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASF---G 74
S+F+ S SS + P P IS S PL P+ SP +PPP S
Sbjct: 138 SSFSHSPSSPSSSLHKPPLPPTQISTH*SPSKPLPTPPTNSPHG-----TPPPTSAPGTA 302
Query: 75 SPASPVAAPA-VTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSP 133
SPAS A+PA + +S ST PS P+++ +SA+ AS+ TS SP+ + +SP
Sbjct: 303 SPASETASPASSSKISTSTAPSTP*PPSLSSVSSASKTTASTAPYQTSPTSPA*DSSSSP 482
Query: 134 PSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAA 192
+ P SP+ +P PS ST T +P++ S + P T + G++ + V AA
Sbjct: 483 TTTSPVTSPSRSP---PSPASTALT-SPTTASPARFRPP*TTSRGSSHSALMVTNSTAA 647
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 68.6 bits (166), Expect = 7e-13
Identities = 59/177 (33%), Positives = 82/177 (45%), Gaps = 8/177 (4%)
Frame = +3
Query: 12 ASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSP------FAVNYP 65
A+ L +S + +L S TP P P SP N P P P F+ + P
Sbjct: 12 AATLATSHGS*NLVSPIQTTPPPPPPP-------PSPNHRNNLPPPFPPSLTTSFSTSSP 170
Query: 66 TSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSP 125
S P S SP+S P + STPP+ S A A TS+ +P + SP P ++SP P
Sbjct: 171 ASHPLPSLPSPSSAAVGP-----TSSTPPTSPTSAAAA--TSSATPPSPSPPPTSASPPP 329
Query: 126 SPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPP--TKAPSTPTNGAT 180
S +P PP +SPA P +PS STP + +S+ + P + + TP G T
Sbjct: 330 PSS---TPNGTPPSSSPATTP--SPSRPSTPSSPTRASLPSAPGFSSSEETPRCGTT 485
Score = 62.0 bits (149), Expect = 7e-11
Identities = 38/138 (27%), Positives = 74/138 (53%), Gaps = 3/138 (2%)
Frame = +3
Query: 58 SPFAVNYPTSPPPAS--FGSPASPVAAPAVTPLSISTPPSQAPSPAI-APSTSANSPVAS 114
SP P PPP S + P P++T ++ P+ P P++ +PS++A P +S
Sbjct: 54 SPIQTTPPPPPPPPSPNHRNNLPPPFPPSLTTSFSTSSPASHPLPSLPSPSSAAVGPTSS 233
Query: 115 SPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPST 174
+P P + + + + ++ +PPSP P + A+ P + + TPP+ +P+ +TP PST
Sbjct: 234 TP-PTSPTSAAAATSSATPPSPSPPPTSASPPPPSSTPNGTPPSSSPA--TTPSPSRPST 404
Query: 175 PTNGATMNGFNVAGYAAA 192
P++ + + G++++
Sbjct: 405 PSSPTRASLPSAPGFSSS 458
Score = 43.9 bits (102), Expect = 2e-05
Identities = 38/107 (35%), Positives = 49/107 (45%), Gaps = 8/107 (7%)
Frame = +3
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAA 144
V+P+ + PP P P +P+ N P P TS + SP++ P P P +S A
Sbjct: 51 VSPIQTTPPP---PPPPPSPNHRNNLPPPFPPSLTTSFSTSSPASHPLPSLPSP-SSAAV 218
Query: 145 APAITPSSISTPPTKA--------PSSISTPPTKAPSTPTNGATMNG 183
P T S+ T PT A P S S PPT A S P +T NG
Sbjct: 219 GP--TSSTPPTSPTSAAAATSSATPPSPSPPPTSA-SPPPPSSTPNG 350
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 68.6 bits (166), Expect = 7e-13
Identities = 57/158 (36%), Positives = 74/158 (46%), Gaps = 7/158 (4%)
Frame = +2
Query: 25 SSSPVATP--SPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFG--SPASPV 80
SSSP P +P SP A SP+ S P SPSP TSPP ++ G SP++P
Sbjct: 194 SSSPWLLPPQTPTTSP-AFSPSTPSSPP-STTTSPSP------TSPPRSTNGPQSPSAPS 349
Query: 81 AAPAVTPLSIST---PPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
P T S +T PPS+ SP+ + ST TS+P S + +PPSPP
Sbjct: 350 TTPPWTTSSQNTHQSPPSRTSSPSTSSST-------------TSAPRSSTRSPTAPPSPP 490
Query: 138 PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
P A P P S ++P + A S S P T +P
Sbjct: 491 QCTKPPAPPRAPPDSSTSPTSAAGKSDSAPRTTTAPSP 604
Score = 58.9 bits (141), Expect = 6e-10
Identities = 55/194 (28%), Positives = 79/194 (40%), Gaps = 36/194 (18%)
Frame = +2
Query: 17 SSTFARSLSSSPVAT---PSPAGSPLAISPAVSSPVPLMNAP---------------SPS 58
S F+ S SSP +T PSP P + + S P P + S
Sbjct: 236 SPAFSPSTPSSPPSTTTSPSPTSPPRSTNGPQSPSAPSTTPPWTTSSQNTHQSPPSRTSS 415
Query: 59 PFAVNYPTSPPPASFGSPASPVAAPAVT--PLSISTPPSQAPSPAIAPSTSANSP-VASS 115
P + TS P +S SP +P + P T P PP + SP A S ++P ++
Sbjct: 416 PSTSSSTTSAPRSSTRSPTAPPSPPQCTKPPAPPRAPPDSSTSPTSAAGKSDSAPRTTTA 595
Query: 116 PVPVTSS------PSPSPSAINSPPSPPPHASP-------AAAPAITPSSI--STPPTKA 160
P P+ SS P+ SPS+ + PPP P +P + PS++ S+P
Sbjct: 596 PSPLRSSNP*RKSPTTSPSSRSVRFFPPPQQKPLHPHPLSRISPPLCPSTVARSSPTLSP 775
Query: 161 PSSISTPPTKAPST 174
P T P+ ST
Sbjct: 776 PRRTRTQPSPTTST 817
Score = 51.6 bits (122), Expect = 9e-08
Identities = 45/144 (31%), Positives = 60/144 (41%), Gaps = 19/144 (13%)
Frame = +2
Query: 22 RSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSP------SPFAVNYPTSPPPASFGS 75
RS + SP A PSP P P P ++ SP S A T+P P +
Sbjct: 449 RSSTRSPTAPPSP---PQCTKPPAPPRAPPDSSTSPTSAAGKSDSAPRTTTAPSPLRSSN 619
Query: 76 P--ASPVAAPAVTPLSISTPPSQAP---------SPAIAPSTSA-NSPVASSPVPVTSSP 123
P SP +P+ + PP Q P SP + PST A +SP S P + P
Sbjct: 620 P*RKSPTTSPSSRSVRFFPPPQQKPLHPHPLSRISPPLCPSTVARSSPTLSPPRRTRTQP 799
Query: 124 SPSPSAINSP-PSPPPHASPAAAP 146
SP+ S + P P+P A+ P
Sbjct: 800 SPTTSTVG*PFPAPSTTQFKASLP 871
>TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II , partial
(6%)
Length = 583
Score = 68.6 bits (166), Expect = 7e-13
Identities = 57/150 (38%), Positives = 70/150 (46%), Gaps = 7/150 (4%)
Frame = -1
Query: 18 STFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
S+ R SS P A P PA P P SPSP A + PTS P P
Sbjct: 427 SSPGRKKSSETAIRPHSASEP---DPAPPPPPPSSPPMSPSPTATSPPTSSPNTH---PP 266
Query: 78 SPVAAPAVTPL-SISTPPSQAPSPAIAPST----SANSPVASSP--VPVTSSPSPSPSAI 130
+P P +P S +T S +PS PST ++ SP +SSP P +S S SP I
Sbjct: 265 TPTPNP*SSPSPSTTTASSASPSTKTLPSTRPRPASESPTSSSPSSPPPSSGSSVSPPRI 86
Query: 131 NSPPSPPPHASPAAAPAITPSSISTPPTKA 160
SPP PPP SP A P PSS + P + +
Sbjct: 85 -SPPPPPPSNSPTATP---PSSATNPSSSS 8
Score = 65.1 bits (157), Expect = 8e-12
Identities = 57/164 (34%), Positives = 77/164 (46%), Gaps = 9/164 (5%)
Frame = -1
Query: 23 SLSSSPVATPSPAG----SPLAISP-AVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
S SSS A PS G S AI P + S P P AP P P P+SPP +
Sbjct: 457 SRSSSSSAPPSSPGRKKSSETAIRPHSASEPDP---APPPPP-----PSSPPMSP----- 317
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSP----VPVTSSPSPSPSAINSP 133
SP A T + PP+ P+P +PS S + ++SP +P T S S +S
Sbjct: 316 SPTATSPPTSSPNTHPPTPTPNP*SSPSPSTTTASSASPSTKTLPSTRPRPASESPTSSS 137
Query: 134 PSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
PS PP P++ +++P IS PP P ++P PS+ TN
Sbjct: 136 PSSPP---PSSGSSVSPPRISPPP---PPPSNSPTATPPSSATN 23
Score = 46.6 bits (109), Expect = 3e-06
Identities = 33/93 (35%), Positives = 47/93 (50%)
Frame = -1
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPA 84
S SP T + + SP + + P P +P+ S P+SPPP+S GS SP P
Sbjct: 241 SPSPSTTTASSASPSTKTLPSTRPRPASESPTSSS-----PSSPPPSS-GSSVSP---PR 89
Query: 85 VTPLSISTPPSQAPSPAIAPSTSANSPVASSPV 117
++P PP + SP P +SA +P +SS V
Sbjct: 88 ISP----PPPPPSNSPTATPPSSATNPSSSSSV 2
Score = 42.0 bits (97), Expect = 7e-05
Identities = 33/108 (30%), Positives = 48/108 (43%), Gaps = 1/108 (0%)
Frame = -1
Query: 70 PASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSA 129
P SF A +T I S++ S + PS+ + + + S+ P P+
Sbjct: 532 PLSFFHFTCVRIAMTLTR*GIKRCVSRSSSSSAPPSSPGRKKSSETAIRPHSASEPDPAP 353
Query: 130 INSPPSPPPHA-SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPT 176
PPS PP + SP A T S + PPT P+ S+P +PST T
Sbjct: 352 PPPPPSSPPMSPSPTATSPPTSSPNTHPPTPTPNP*SSP---SPSTTT 218
Score = 38.1 bits (87), Expect = 0.001
Identities = 29/87 (33%), Positives = 42/87 (47%), Gaps = 1/87 (1%)
Frame = -1
Query: 95 SQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSIS 154
S + +P +P +S A P S P P+P PP PP SP +P+ T +S
Sbjct: 448 SSSSAPPSSPGRKKSSETAIRPHSA-SEPDPAP-----PPPPPS--SPPMSPSPTATS-- 299
Query: 155 TPPTKAPSSISTPPTKAP-STPTNGAT 180
PPT +P++ PT P S+P+ T
Sbjct: 298 -PPTSSPNTHPPTPTPNP*SSPSPSTT 221
Score = 35.0 bits (79), Expect = 0.009
Identities = 20/66 (30%), Positives = 31/66 (46%), Gaps = 11/66 (16%)
Frame = -1
Query: 128 SAINSPPSPP-----------PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPT 176
S+ ++PPS P PH++ PA P S+PP + ++PPT +P+T
Sbjct: 448 SSSSAPPSSPGRKKSSETAIRPHSASEPDPAPPPPPPSSPPMSPSPTATSPPTSSPNTHP 269
Query: 177 NGATMN 182
T N
Sbjct: 268 PTPTPN 251
>TC9746
Length = 637
Score = 68.2 bits (165), Expect = 1e-12
Identities = 54/142 (38%), Positives = 75/142 (52%), Gaps = 6/142 (4%)
Frame = +2
Query: 31 TPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPP--PASFGS--PASPVAAPAVT 86
TPSP P P P P A S SP + + P S P PA +G+ P SP ++P++
Sbjct: 170 TPSPTSKPKCQPP----PHPSSTATS-SPASNSEPPSGPNSPARWGT*APTSPSSSPSL* 334
Query: 87 PLSISTPP-SQAPSPAIAP-STSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAA 144
++++PP S +P +P ++S+ SP SSP S PSPSP P SP P + P A
Sbjct: 335 STTLTSPPHSSSPHSTTSPLASSSASPCQSSP*S-PSPPSPSPK---HPHSPSPRSPPPA 502
Query: 145 APAITPSSISTPPTKAPSSIST 166
+P+ S S PP PSS +T
Sbjct: 503 SPSPPYFSSSAPPVSCPSSTAT 568
Score = 45.4 bits (106), Expect = 7e-06
Identities = 35/82 (42%), Positives = 41/82 (49%), Gaps = 6/82 (7%)
Frame = +2
Query: 25 SSSPVATPSPAGSPLAISPAVSSP-VPLMNAPSPSPFAVNYPTSPPPAS-----FGSPAS 78
SSSP +T SP S A SP SSP P +PSP P SPPPAS F S A
Sbjct: 362 SSSPHSTTSPLASSSA-SPCQSSP*SPSPPSPSPKHPHSPSPRSPPPASPSPPYFSSSAP 538
Query: 79 PVAAPAVTPLSISTPPSQAPSP 100
PV+ P+ T S+ + SP
Sbjct: 539 PVSCPSSTATSLCQSSAAFSSP 604
Score = 41.2 bits (95), Expect = 1e-04
Identities = 37/112 (33%), Positives = 52/112 (46%), Gaps = 1/112 (0%)
Frame = +2
Query: 65 PTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPS 124
PTS P P P + +P S S PPS SPA +P +SPS
Sbjct: 179 PTSKPKCQ--PPPHPSSTATSSPASNSEPPSGPNSPA---RWGT*AP---------TSPS 316
Query: 125 PSPSAINSPPSPPPHA-SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
SPS ++ + PPH+ SP + + SS ++P +P S S PP+ +P P
Sbjct: 317 SSPSL*STTLTSPPHSSSPHSTTSPLASSSASPCQSSP*SPS-PPSPSPKHP 469
Score = 26.9 bits (58), Expect = 2.4
Identities = 14/42 (33%), Positives = 17/42 (40%)
Frame = +3
Query: 133 PPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPST 174
PP PPHASPA + PT P + P + T
Sbjct: 396 PPLRPPHASPAHEVHRRRRHLRNTPTHHPPDLRRRPLRRRRT 521
>TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber hypocotyls
(Arabinogalactan protein), partial (26%)
Length = 795
Score = 67.8 bits (164), Expect = 1e-12
Identities = 58/164 (35%), Positives = 80/164 (48%), Gaps = 1/164 (0%)
Frame = +2
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSS-PVPLMNAPSPSPFAVNYPTSPPPASFGS 75
S+T S +PVA+P PA + A SP+ ++ P P P V TSPPPA
Sbjct: 149 SATTPVSTPVAPVASPKPAATTPAASPSTATAPAPATTTP------VTPVTSPPPA---- 298
Query: 76 PASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPS 135
A PVA+P + +S+PP+++P PA AP+ PVA+ PV P+P+PS
Sbjct: 299 -AVPVASPPPAAVPVSSPPAKSP-PAPAPTAV---PVAA-PVTTPEVPAPAPSKTKKD-- 454
Query: 136 PPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGA 179
AAPA +P +PP AP P+ A S +GA
Sbjct: 455 --------AAPAPSPVVPDSPPAGAPG-----PSDAVSPGPDGA 547
Score = 63.9 bits (154), Expect = 2e-11
Identities = 50/164 (30%), Positives = 75/164 (45%), Gaps = 1/164 (0%)
Frame = +2
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
S T A + S + + +P +P+A V+SP P P+ SP T+P PA+ +P
Sbjct: 110 SPTAAPTTSPATPSATTPVSTPVA---PVASPKPAATTPAASPSTA---TAPAPATT-TP 268
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSP 136
+PV +P + +++PP PA VPV+S P+ SPP+P
Sbjct: 269 VTPVTSPPPAAVPVASPP-----PA--------------AVPVSSPPA------KSPPAP 373
Query: 137 PPHASPAAAPAITPSSISTPPTKAPSSISTPPTK-APSTPTNGA 179
P A P AAP TP + P+K + P+ P +P GA
Sbjct: 374 APTAVPVAAPVTTPEVPAPAPSKTKKDAAPAPSPVVPDSPPAGA 505
>AV411518
Length = 360
Score = 67.4 bits (163), Expect = 2e-12
Identities = 44/114 (38%), Positives = 55/114 (47%), Gaps = 12/114 (10%)
Frame = +3
Query: 67 SPPPASFGSPASPVAAPAVTPLSISTPPSQ--APSPAIAPSTSAN----------SPVAS 114
SP PA+ P SP A P TP + PPS P+PA A S A+ P +
Sbjct: 18 SPSPATSWLPPSP*AHP--TPATSWLPPSP*AHPTPAWAGSALASWASPCAPTSSGPASP 191
Query: 115 SPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPP 168
SP SP P+PS+ + P SP PH P + S + PPT APSS + PP
Sbjct: 192 SPSSTAPSPRPNPSSTSGPTSPTPHTPSPPPPTSSSPSSAIPPTSAPSSSTPPP 353
Score = 42.7 bits (99), Expect = 4e-05
Identities = 33/90 (36%), Positives = 42/90 (46%), Gaps = 12/90 (13%)
Frame = +3
Query: 24 LSSSPVATPSPA--GSPLA--------ISPAVSSPVPLMNAPSP--SPFAVNYPTSPPPA 71
L SP A P+PA GS LA S +SP P APSP +P + + PTSP P
Sbjct: 87 LPPSP*AHPTPAWAGSALASWASPCAPTSSGPASPSPSSTAPSPRPNPSSTSGPTSPTPH 266
Query: 72 SFGSPASPVAAPAVTPLSISTPPSQAPSPA 101
+ P ++P+ S P S P PA
Sbjct: 267 TPSPPPPTSSSPSSAIPPTSAPSSSTPPPA 356
Score = 41.6 bits (96), Expect = 1e-04
Identities = 35/94 (37%), Positives = 48/94 (50%), Gaps = 15/94 (15%)
Frame = +3
Query: 98 PSPAIAPSTSAN----SPVASSPVPVTSSPSPSP----------SAINSPPSP-PPHASP 142
P+P+++PS + + SP A P P TS PSP SA+ S SP P +S
Sbjct: 3 PTPSLSPSPATSWLPPSP*AH-PTPATSWLPPSP*AHPTPAWAGSALASWASPCAPTSSG 179
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPTKAPSTPT 176
A+P +PSS + P PSS S P + P TP+
Sbjct: 180 PASP--SPSSTAPSPRPNPSSTSGPTSPTPHTPS 275
Score = 30.4 bits (67), Expect = 0.22
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 18/80 (22%)
Frame = +3
Query: 122 SPSPSPSAINS--PPSPPPHASPAAA-----------PAITPSSIST-----PPTKAPSS 163
+PS SPS S PPSP H +PA + PA S++++ PT + +
Sbjct: 6 TPSLSPSPATSWLPPSP*AHPTPATSWLPPSP*AHPTPAWAGSALASWASPCAPTSSGPA 185
Query: 164 ISTPPTKAPSTPTNGATMNG 183
+P + APS N ++ +G
Sbjct: 186 SPSPSSTAPSPRPNPSSTSG 245
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.306 0.120 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,363,438
Number of Sequences: 28460
Number of extensions: 134779
Number of successful extensions: 15389
Number of sequences better than 10.0: 2039
Number of HSP's better than 10.0 without gapping: 3921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8618
length of query: 202
length of database: 4,897,600
effective HSP length: 86
effective length of query: 116
effective length of database: 2,450,040
effective search space: 284204640
effective search space used: 284204640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144517.1


