

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144516.5 - phase: 0
(332 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
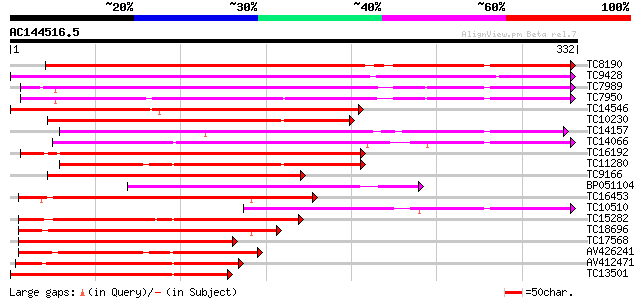
Score E
Sequences producing significant alignments: (bits) Value
TC8190 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1 p... 276 3e-75
TC9428 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), pa... 262 7e-71
TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , par... 236 5e-63
TC7950 similar to UP|Q9XFI8 (Q9XFI8) Peroxidase (Fragment), part... 197 3e-51
TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxida... 191 1e-49
TC10230 similar to UP|PE10_ARATH (Q9FX85) Peroxidase 10 precurso... 183 3e-47
TC14157 similar to UP|O24080 (O24080) Peroxidase2 precursor , p... 183 4e-47
TC14066 homologue to UP|O64970 (O64970) Cationic peroxidase 2 ,... 181 2e-46
TC16192 similar to UP|Q9SSZ9 (Q9SSZ9) Peroxidase 1, partial (49%) 174 1e-44
TC11280 similar to UP|Q9XFL5 (Q9XFL5) Peroxidase 4 (Fragment), p... 165 9e-42
TC9166 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, partial (49%) 154 3e-38
BP051104 151 1e-37
TC16453 similar to UP|Q40372 (Q40372) Peroxidase precursor, part... 144 3e-35
TC10510 similar to PIR|B56555|B56555 peroxidase , anionic, precu... 140 2e-34
TC15282 similar to UP|Q9ZNZ6 (Q9ZNZ6) Peroxidase precursor (Fra... 136 6e-33
TC18696 similar to UP|Q40372 (Q40372) Peroxidase precursor, part... 132 1e-31
TC17568 similar to UP|Q8RVP6 (Q8RVP6) Gaiacol peroxidase , part... 126 6e-30
AV426241 125 8e-30
AV412471 125 1e-29
TC13501 similar to UP|PE47_ARATH (Q9SZB9) Peroxidase 47 precurso... 124 2e-29
>TC8190 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1 precursor
(PNPC1) , partial (96%)
Length = 1202
Score = 276 bits (706), Expect = 3e-75
Identities = 150/310 (48%), Positives = 189/310 (60%)
Frame = +1
Query: 22 GSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDS 81
GS L +Y + CPL ++ V +AV K+ R+ ASLLRLHFHDCFV GCDAS+LLD
Sbjct: 130 GSAQLSSTFYAKTCPLVLATIKTQVNLAVAKEARMGASLLRLHFHDCFVQGCDASILLDD 309
Query: 82 VEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRW 141
T EK AGPN NS+RG++VID IK +E CP VSCADI+A+ ARD+V GG W
Sbjct: 310 TSSFTGEKTAGPNANSVRGYDVIDTIKSKVESLCPGVVSCADIVAVAARDSVVALGGFSW 489
Query: 142 EVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCL 201
V LGR+DS +S S AN +P P+S+L+ L F +G ++V LSGSHTIG+ARCL
Sbjct: 490 AVPLGRRDSTTASLSSANSELPGPSSNLDGLNTAFSNKGFTTREMVALSGSHTIGQARCL 669
Query: 202 SFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFI 261
FR RIY Y D +TF + LQ CP G D +PLD +P FDN Y+
Sbjct: 670 FFRTRIY-----YETHID-----STFAKNLQGNCPFNGGDSNLSPLDTTSPTTFDNGYYR 819
Query: 262 NIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEG 321
N+ KGL SD VL + G QV Y +N F FA +M+KMGN++ LTGS G
Sbjct: 820 NLQSQKGLFHSDQVLFN---GGSTDSQVNSYVTNPASFKTDFANAMVKMGNLSPLTGSSG 990
Query: 322 EIRRNCRFVN 331
+IR +CR N
Sbjct: 991 QIRTHCRKTN 1020
>TC9428 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), partial
(97%)
Length = 1354
Score = 262 bits (669), Expect = 7e-71
Identities = 148/331 (44%), Positives = 195/331 (58%)
Frame = +2
Query: 1 MEPLRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASL 60
M P+ L ++ ++ + L + +YK+ CP IVR V DPR+ SL
Sbjct: 41 MSPIHLAVVFCVVIGALVPYSSDAQLDNSFYKDTCPNVHSIVREVVRNVSKSDPRMLGSL 220
Query: 61 LRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVS 120
+RLHFHDCFV GCDAS+LL+ + SE+ A PN NS+RG +V+++IK +E CP TVS
Sbjct: 221 IRLHFHDCFVQGCDASILLNDTSTIVSEQGAFPNNNSIRGLDVVNQIKTAVENACPNTVS 400
Query: 121 CADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQG 180
CADILA+ A + L G W+V LGR+DSL ++ S AN +P PN +L L +F Q
Sbjct: 401 CADILALAAEISSVLANGVDWKVPLGRRDSLTANRSLANQNLPGPNFNLSQLNASFAVQN 580
Query: 181 LDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGR 240
L+I DLV LSG+HTIGR +C F R+Y + TT+ + L+SICP G
Sbjct: 581 LNITDLVALSGAHTIGRGQCQFFVNRLYNFS---NTGNPDPTLNTTYLQTLRSICPHGGP 751
Query: 241 DDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFF 300
A LD TP D+QYF N+ GKGL SD VL S + V ++SN LFF
Sbjct: 752 GTTLAHLDPTTPDTLDSQYFSNLQGGKGLFQSDPVLFSTSGAATV-AIVNSFSSNPTLFF 928
Query: 301 DSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
++F SMIKM I VLTGS+GEIR+ C FVN
Sbjct: 929 ENFKASMIKMSRIGVLTGSQGEIRKPCNFVN 1021
>TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (85%)
Length = 1352
Score = 236 bits (601), Expect = 5e-63
Identities = 140/335 (41%), Positives = 193/335 (56%), Gaps = 10/335 (2%)
Frame = +3
Query: 7 LLILITILSNIHTLRGSEL--------LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAA 58
LL++ + LS H +R SE L +Y CP E I+R + KD AA
Sbjct: 60 LLLISSFLSFSH-IRVSEAATAPIVKGLSWTFYDSSCPKLESIIRKELKKVFDKDIAQAA 236
Query: 59 SLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVN-SLRGFEVIDKIKYLLEKECPL 117
LLRLHFHDCFV GCDASVLLD SEK A PN+ + F++I+ ++ +EK+C
Sbjct: 237 GLLRLHFHDCFVQGCDASVLLDGSASGPSEKDAPPNLTLRAQAFKIIEDLRSQIEKKCGR 416
Query: 118 TVSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANL-FIPAPNSSLETLINNF 176
VSCADI A+ ARDAV L GGP +E+ LGR+D L + L +PAP S+ T++N+
Sbjct: 417 VVSCADITAIAARDAVFLSGGPDYELPLGRRDGLNFATRNVTLDNLPAPQSNTTTILNSL 596
Query: 177 KQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICP 236
+ LD D+V LSG HTIG + C SF R+Y +K TF + L+ CP
Sbjct: 597 ATKNLDPTDVVSLSGGHTIGISHCSSFTDRLYPSKDPVMD--------QTFEKNLKLTCP 752
Query: 237 VTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNE 296
+ D+ LD ++P FDN+Y+++++ +GL SD L + D R + V +A N+
Sbjct: 753 ASNTDNT-TVLDLRSPNTFDNKYYVDLMNRQGLFFSDQDLYT---DKRTKDIVTSFAVNQ 920
Query: 297 KLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
LFF+ F +M+KMG +NVLTGS+GEIR NC N
Sbjct: 921 SLFFEKFVVAMLKMGQLNVLTGSQGEIRANCSVRN 1025
>TC7950 similar to UP|Q9XFI8 (Q9XFI8) Peroxidase (Fragment), partial (86%)
Length = 1286
Score = 197 bits (500), Expect = 3e-51
Identities = 120/335 (35%), Positives = 177/335 (52%), Gaps = 10/335 (2%)
Frame = +1
Query: 7 LLILITILSNIHTLRGSEL---------LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLA 57
L ++ +IL H GSE L +Y + CP E +VR+++ + KD A
Sbjct: 22 LFLIFSILFTSHFFLGSEAQTKPPVVEGLSFSFYSKTCPKLETVVRNHLKKVLKKDNGQA 201
Query: 58 ASLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVN-SLRGFEVIDKIKYLLEKECP 116
LLR+ FHDCFV GCD SVLLD G E+ N+ + I+ I+ L+ K+C
Sbjct: 202 PGLLRIFFHDCFVQGCDGSVLLD---GSPGERDQPANIGIRPEALQTIEDIRALVHKQCG 372
Query: 117 LTVSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNF 176
VSCADI + +RDAV L GGP + V LGR+D + S G +P+P ++ + F
Sbjct: 373 KIVSCADITILASRDAVFLTGGPDYAVPLGRRDGVSFSTVGTQK-LPSPINNTTATLKAF 549
Query: 177 KQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICP 236
+ D D+V LSG+HT GRA C +F R+ T + L + CP
Sbjct: 550 ADRNFDATDVVALSGAHTFGRAHCGTFFNRLSPLDPNMD---------KTLAKNLTATCP 702
Query: 237 VTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNE 296
+ A LD +TP FDN+Y+++++ +G+ SD L+S D R + V +A N+
Sbjct: 703 AQNSTNT-ANLDIRTPNVFDNKYYLDLMNRQGVFTSDQDLLS---DKRTKGLVNAFAVNQ 870
Query: 297 KLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
LFF+ F ++IK+ ++VLTG++GEIR C VN
Sbjct: 871 TLFFEKFVDAVIKLSQLDVLTGNQGEIRGRCNVVN 975
>TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (66%)
Length = 738
Score = 191 bits (486), Expect = 1e-49
Identities = 102/209 (48%), Positives = 130/209 (61%), Gaps = 2/209 (0%)
Frame = +1
Query: 1 MEPLRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASL 60
M P ++IL ++ + L +Y CP + IVR+ + + K+ R+ AS+
Sbjct: 25 MAPHSHFFAALSILLSLLACSANAQLTPTFYDRSCPKLQTIVRNAMVQTIKKEARMGASI 204
Query: 61 LRLHFHDCFVMGCDASVLLDSVEGMT--SEKQAGPNVNSLRGFEVIDKIKYLLEKECPLT 118
LRL FHDCFV GCD S+LLD + G T EK A PN NS RGFEVID IK +E C T
Sbjct: 205 LRLFFHDCFVNGCDGSILLDDI-GTTFVGEKNAAPNKNSARGFEVIDTIKTNVEASCNNT 381
Query: 119 VSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQ 178
VSCADILA+ RD + L GGP W+V LGR+D+ +S S AN IP+P+S L TLI+ F
Sbjct: 382 VSCADILALATRDGINLLGGPTWQVPLGRRDARTASQSKANTEIPSPSSDLSTLISMFSA 561
Query: 179 QGLDIEDLVVLSGSHTIGRARCLSFRQRI 207
+GL DL VLSG HTIG+A C FR R+
Sbjct: 562 KGLSARDLTVLSGGHTIGQAECQFFRSRV 648
>TC10230 similar to UP|PE10_ARATH (Q9FX85) Peroxidase 10 precursor (Atperox
P10) (ATP5a) , partial (42%)
Length = 753
Score = 183 bits (465), Expect = 3e-47
Identities = 97/180 (53%), Positives = 118/180 (64%)
Frame = +3
Query: 23 SELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSV 82
S L + +Y CP IVR+N+ A+ D R+AASLLRLHFHDCFV GCD SVLLD
Sbjct: 216 SSQLYYNFYDTTCPNLTRIVRYNLLSAMSNDTRIAASLLRLHFHDCFVNGCDGSVLLDDT 395
Query: 83 EGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWE 142
EK A PN NS+RGFEVID IK LEK CP TVSCADIL + AR+AV L GP W
Sbjct: 396 STQKGEKNALPNKNSIRGFEVIDTIKSALEKACPSTVSCADILTLAAREAVYLSRGPFWS 575
Query: 143 VWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLS 202
V LGR+D +S S AN +P+P LE + F +GL+ +D+ VLSG+HT G A+C +
Sbjct: 576 VPLGRRDGTTASESEAN-NLPSPFEPLENITAKFISKGLEKKDVAVLSGAHTFGFAQCFT 752
>TC14157 similar to UP|O24080 (O24080) Peroxidase2 precursor , partial
(96%)
Length = 1390
Score = 183 bits (464), Expect = 4e-47
Identities = 116/301 (38%), Positives = 159/301 (52%), Gaps = 3/301 (0%)
Frame = +1
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y CP + IV+ V + + LRL FHDCFV GCDASV++ S +EK
Sbjct: 139 HYASTCPNLQSIVKGVVQKKFQQTFVTVPATLRLFFHDCFVQGCDASVMVASSGNNKAEK 318
Query: 90 QAGPNVNSLR-GFEVIDKIKYLLEK--ECPLTVSCADILAMVARDAVELRGGPRWEVWLG 146
N++ GF+ + K K ++ +C VSCADILA+ RD V L GGP + V LG
Sbjct: 319 DHPDNLSLAGDGFDTVIKAKAAVDAVPQCRNKVSCADILALATRDVVVLAGGPSYTVELG 498
Query: 147 RKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQR 206
R D L S S N +P PN +L L + F QGL D++ LSG+HT+G + C F R
Sbjct: 499 RFDGLVSRASDVNGRLPEPNFNLNQLNSLFASQGLTQTDMIALSGAHTLGFSHCNRFSNR 678
Query: 207 IYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEG 266
IY T + + Y T LQ +CP +D TP+ FDN Y+ N+ +G
Sbjct: 679 IYSTPVD----PTLNRNYAT---QLQQMCPKNVNPQIAINMDPTTPRTFDNIYYKNLQQG 837
Query: 267 KGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRN 326
KGL SD +L + D R + V +ASN F +FA +MIK+G + V T G+IR +
Sbjct: 838 KGLFTSDQILFT---DQRSKATVNSFASNSNTFNANFAAAMIKLGRVGVKTARNGKIRTD 1008
Query: 327 C 327
C
Sbjct: 1009C 1011
>TC14066 homologue to UP|O64970 (O64970) Cationic peroxidase 2 , partial
(92%)
Length = 1436
Score = 181 bits (458), Expect = 2e-46
Identities = 111/312 (35%), Positives = 156/312 (49%), Gaps = 6/312 (1%)
Frame = +2
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L +YKE CP AEDI+ V + + A S LR FHDC V CDAS+LLDS
Sbjct: 152 LAMNFYKETCPQAEDIITEQVKLLYKRHKNTAFSWLRNIFHDCAVQSCDASLLLDSTRKT 331
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
SEK+ + LR F I+ IK LE+ECP VSC+DIL + ARD + GGP +
Sbjct: 332 LSEKETDRSFG-LRNFRYIETIKEALERECPGVVSCSDILVLSARDGIAALGGPHIPLRT 508
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GR+D S F+P N S+ +++ F G+D +V L G+H++GR C+
Sbjct: 509 GRRDGRRSRADVVEQFLPDHNESISAVLDKFGAMGIDTPGVVALLGAHSVGRTHCVKLVH 688
Query: 206 RIY---ETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDK---FAPLDFQTPKRFDNQY 259
R+Y + H K+ CP D K + D TP DN Y
Sbjct: 689 RLYPEVDPALNPDHIPHMLKK-----------CPDAIPDPKAVQYVRNDRGTPMILDNNY 835
Query: 260 FINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGS 319
+ NI++ KGLL D+ L + D R + V A ++ FF F++++ + N LTG+
Sbjct: 836 YRNILDNKGLLLVDHQLAN---DKRTKPYVKKMAKSQDYFFKEFSRAITLLSENNPLTGT 1006
Query: 320 EGEIRRNCRFVN 331
+GEIR+ C N
Sbjct: 1007KGEIRKQCNVAN 1042
>TC16192 similar to UP|Q9SSZ9 (Q9SSZ9) Peroxidase 1, partial (49%)
Length = 645
Score = 174 bits (442), Expect = 1e-44
Identities = 96/202 (47%), Positives = 127/202 (62%)
Frame = +2
Query: 7 LLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFH 66
+++L+ L N + SEL V YY C +AE IV+ V +V K+P +AA L+R+HFH
Sbjct: 35 IIVLVIFLLNQNAY--SELEVG-YYSYSCGMAEFIVKDEVRRSVTKNPGIAAGLVRMHFH 205
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCF+ GCDASVLLDS T+EK + N SLRGFEVID K LE C VSCADI+A
Sbjct: 206 DCFIRGCDASVLLDSTPSNTAEKDSPANKPSLRGFEVIDSAKAKLEAVCKGVVSCADIIA 385
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
ARD+VEL GG ++V GR+D S S +P P ++ L F ++GL E++
Sbjct: 386 FAARDSVELAGGVGYDVPAGRRDGRISLASDTRTDLPPPTFNVNQLTQLFAKKGLTQEEM 565
Query: 187 VVLSGSHTIGRARCLSFRQRIY 208
V LSG+HTIGR+ C +F R+Y
Sbjct: 566 VTLSGAHTIGRSHCAAFSSRLY 631
>TC11280 similar to UP|Q9XFL5 (Q9XFL5) Peroxidase 4 (Fragment), partial
(68%)
Length = 614
Score = 165 bits (418), Expect = 9e-42
Identities = 93/179 (51%), Positives = 111/179 (61%)
Frame = +3
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y CP AE IVR V V DP LAA LLR+HFHDCFV GCDASVL + G +E+
Sbjct: 36 FYLGTCPRAESIVRSTVESHVNSDPTLAAGLLRMHFHDCFVQGCDASVL---IAGAGTER 206
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
A PN+ SLRGFEVID K +E CP VSCADILA+ ARD+V L GG W+V GR+D
Sbjct: 207 TAIPNL-SLRGFEVIDDAKAKVEAACPGVVSCADILALAARDSVVLSGGLSWQVPTGRRD 383
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIY 208
S S N +PAP S++ F +GL+ +DLV L G HTIG C F R+Y
Sbjct: 384 GRVSQASDVN-NLPAPFDSVDVQKQKFTAKGLNTQDLVTLVGGHTIGTTACQFFSNRLY 557
>TC9166 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, partial (49%)
Length = 559
Score = 154 bits (388), Expect = 3e-38
Identities = 82/152 (53%), Positives = 100/152 (64%), Gaps = 1/152 (0%)
Frame = +3
Query: 23 SELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSV 82
S L +Y +KCP + V+ V AV K+ R+ SLLRL FHDCFV GCD SVLLD
Sbjct: 102 SAQLSENFYVKKCPSVFNAVKSVVHSAVAKEARMGGSLLRLFFHDCFVNGCDGSVLLDDT 281
Query: 83 EGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWE 142
EK A PN NSLRGF+VID IK +E CP VSCAD++A+ ARD+V + GGP W+
Sbjct: 282 SSFKGEKTAPPNSNSLRGFDVIDAIKSKVEAVCPGVVSCADVVAIAARDSVAILGGPYWK 461
Query: 143 VWLGRKDSLESSFSGANL-FIPAPNSSLETLI 173
V LGR+DS +SF+ AN IP+P SSL LI
Sbjct: 462 VKLGRRDSKTASFNAANSGVIPSPFSSLSDLI 557
>BP051104
Length = 566
Score = 151 bits (382), Expect = 1e-37
Identities = 83/173 (47%), Positives = 102/173 (57%)
Frame = +2
Query: 70 VMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVA 129
V GCDAS+L D E+ A N S RGF VID IK LEK+CP VSCAD+LA+ A
Sbjct: 71 VNGCDASILXDDTSNFIGEQTAAANNRSARGFNVIDGIKANLEKQCPGVVSCADVLALAA 250
Query: 130 RDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVL 189
RD+V GGP WEV LGR+DS +S AN IP P SL LI NF QGL + DLV L
Sbjct: 251 RDSVVQLGGPSWEVGLGRRDSTTASRGTANNTIPGPFLSLSGLITNFANQGLSVTDLVAL 430
Query: 190 SGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDD 242
SG+HTIG A+C +FR H Y+ ++ + L+ CP +G D+
Sbjct: 431 SGAHTIGLAQCKNFRA----------HIYNDSNIDASYAKFLKX*CPRSGNDN 559
>TC16453 similar to UP|Q40372 (Q40372) Peroxidase precursor, partial (49%)
Length = 581
Score = 144 bits (362), Expect = 3e-35
Identities = 81/180 (45%), Positives = 116/180 (64%), Gaps = 5/180 (2%)
Frame = +2
Query: 6 LLLILITILSNI--HTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRL 63
L+L+L+T+ + + TL G L YY + CP A I++ V A+ ++ R+ ASLLRL
Sbjct: 50 LVLVLVTLATFLIPSTLAG---LTPGYYDKVCPQALPIIKSVVQKAINREQRIGASLLRL 220
Query: 64 HFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPL-TVSCA 122
HFHDCFV GCD S+LLD EK A PN+NS+RGFEV+D+IK +++ C VSCA
Sbjct: 221 HFHDCFVNGCDGSLLLDDTSSFVGEKTALPNLNSVRGFEVVDEIKAAVDRACKRPVVSCA 400
Query: 123 DILAMVARDAVELRGGPR--WEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQG 180
DILA+ ARD+V + GG + ++V LGR+D+ +S + AN +P P + L+ F+ QG
Sbjct: 401 DILAIAARDSVAILGGKQYWYQVRLGRRDARSASQAAANSNLPPPFLNFTQLLPAFQAQG 580
>TC10510 similar to PIR|B56555|B56555 peroxidase , anionic, precursor - wood
tobacco {Nicotiana sylvestris;} , partial (53%)
Length = 764
Score = 140 bits (354), Expect = 2e-34
Identities = 85/197 (43%), Positives = 108/197 (54%), Gaps = 3/197 (1%)
Frame = +3
Query: 138 GPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGR 197
GP W V LGR+DS +S S AN +P+ L+TLI+ F +GL D+V+LSG+HTIG+
Sbjct: 3 GPSWTVKLGRRDSTTASRSLANTELPSFTDDLQTLISRFDNKGLTARDMVILSGAHTIGQ 182
Query: 198 ARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVT---GRDDKFAPLDFQTPKR 254
A+C +FR RIY + + +R CP T D K A LD TP
Sbjct: 183 AQCSTFRGRIYNNASDIDAGFGSTRRRG---------CPSTISVENDKKLAALDLVTPNS 335
Query: 255 FDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNIN 314
FDN YF N+I+ KGLL SD VL G V Y+ N F FA +MIKMG+I
Sbjct: 336 FDNNYFKNLIQKKGLLRSDQVLFR---GGSTDTIVSEYSKNPTTFKSDFAAAMIKMGDIQ 506
Query: 315 VLTGSEGEIRRNCRFVN 331
LTGS G IR+ C +N
Sbjct: 507 PLTGSAGIIRKICSAIN 557
>TC15282 similar to UP|Q9ZNZ6 (Q9ZNZ6) Peroxidase precursor (Fragment) ,
partial (47%)
Length = 604
Score = 136 bits (342), Expect = 6e-33
Identities = 73/167 (43%), Positives = 108/167 (63%)
Frame = +1
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
L++ L+ ++++ H L +Y + CP AE I+ V + P LAA+L+R++F
Sbjct: 118 LIVCLLALIASNHAQ-----LEQGFYTKTCPKAEKIILDFVHEHIHNAPSLAAALIRMNF 282
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
HDCFV GCDAS+LL+S +E+ A PN+ ++RGF+ ID+IK L+E +CP VSCADI+
Sbjct: 283 HDCFVRGCDASILLNSTSKQ-AERDAPPNL-TVRGFDFIDRIKSLVEAQCPGVVSCADII 456
Query: 126 AMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETL 172
A+ ARD++ GGP W+V GR+D + S+ A IPAP S+ TL
Sbjct: 457 ALAARDSIVATGGPFWKVPTGRRDGVISNLVEARSQIPAPTSNFTTL 597
>TC18696 similar to UP|Q40372 (Q40372) Peroxidase precursor, partial (47%)
Length = 500
Score = 132 bits (331), Expect = 1e-31
Identities = 71/157 (45%), Positives = 102/157 (64%), Gaps = 3/157 (1%)
Frame = +2
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
++L+++T+ I + L +Y + CP A ++ V A+L++ R+ ASLLRLHF
Sbjct: 38 IVLVMVTLTLVIPS---KAQLSPSFYNKVCPQALPVINSVVRRAILRERRIGASLLRLHF 208
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPL-TVSCADI 124
HDCFV GCD SVLLD EK A PN NS++GF+V+D+IK ++K C VSCADI
Sbjct: 209 HDCFVNGCDGSVLLDDTRNFIGEKTAFPNNNSIKGFDVVDEIKKAVDKACKRPVVSCADI 388
Query: 125 LAMVARDAVELRGGPR--WEVWLGRKDSLESSFSGAN 159
LA+ ARD+V + GGP ++V LGR+D+ +S + AN
Sbjct: 389 LAIAARDSVAILGGPSLVYKVLLGRRDARTASRAAAN 499
>TC17568 similar to UP|Q8RVP6 (Q8RVP6) Gaiacol peroxidase , partial (33%)
Length = 672
Score = 126 bits (316), Expect = 6e-30
Identities = 66/128 (51%), Positives = 83/128 (64%)
Frame = +2
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
LLL LI +L + S L +Y CP AE IV+ + A+ ++PR AS++R F
Sbjct: 284 LLLFLIFLLHTTSLVTSSSDLRPGFYSNTCPEAEYIVQDVMKKALFREPRSVASVMRFQF 463
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
HDCFV GCDAS+LLD M EK A N+NSLR FEV+D+IK LEK+CP VSCADI+
Sbjct: 464 HDCFVNGCDASMLLDDTPDMLGEKLALSNINSLRSFEVVDEIKEALEKKCPGVVSCADII 643
Query: 126 AMVARDAV 133
M +RDAV
Sbjct: 644 IMASRDAV 667
>AV426241
Length = 429
Score = 125 bits (315), Expect = 8e-30
Identities = 71/143 (49%), Positives = 90/143 (62%)
Frame = +3
Query: 6 LLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHF 65
LL+ LI + + G+ + +Y + CP E IV+ VA V D AA LLRLHF
Sbjct: 21 LLVFLIVLTLQAFAVHGTSV---GFYSKSCPSIESIVKSTVASHVKTDFEYAAGLLRLHF 191
Query: 66 HDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADIL 125
DCFV GCDAS+L + G +EKQA PN SL+G+EVID+ K LE +CP VSCADIL
Sbjct: 192 RDCFVRGCDASIL---IAGNGTEKQAPPN-RSLKGYEVIDEAKAKLEAQCPGVVSCADIL 359
Query: 126 AMVARDAVELRGGPRWEVWLGRK 148
A+ ARD+V L GG W+V GR+
Sbjct: 360 ALAARDSVVLSGGLSWQVPTGRR 428
>AV412471
Length = 430
Score = 125 bits (314), Expect = 1e-29
Identities = 67/134 (50%), Positives = 87/134 (64%)
Frame = +2
Query: 4 LRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRL 63
L L ++ I+S G + L YY CP + IV++ V+ A+ DP LAA L+R+
Sbjct: 38 LTLFFVMEVIVSGFTF--GVDDLNMNYYLFSCPFVDPIVKNKVSTALQNDPTLAAGLIRM 211
Query: 64 HFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCAD 123
HFHDCF+ GCD SVLLDS + T+EK + N+ SLRG+E+ID IK LEK CP VSCAD
Sbjct: 212 HFHDCFIEGCDGSVLLDSTKDNTAEKDSPANL-SLRGYEIIDDIKEELEKRCPGVVSCAD 388
Query: 124 ILAMVARDAVELRG 137
I+AM +RDAV G
Sbjct: 389 IVAMASRDAVFFAG 430
>TC13501 similar to UP|PE47_ARATH (Q9SZB9) Peroxidase 47 precursor (Atperox
P47) (ATP32) , partial (33%)
Length = 519
Score = 124 bits (311), Expect = 2e-29
Identities = 63/130 (48%), Positives = 88/130 (67%)
Frame = +1
Query: 1 MEPLRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASL 60
M L + +LI ++S G+ L YY +CP AE +V++ V A+ DP LAA L
Sbjct: 133 MASLLTVFLLIEVISCGFGFGGNNGLNMNYYLMRCPFAESVVKNIVNRALQNDPTLAAGL 312
Query: 61 LRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVS 120
+R+HFHDCFV GCD S+L+DS + T+EK + N+ SL+G+E+ID+IK LE++CP VS
Sbjct: 313 IRMHFHDCFVEGCDGSILIDSTKDNTAEKDSPANL-SLKGYEIIDEIKEELERQCPGVVS 489
Query: 121 CADILAMVAR 130
CAD+LAM AR
Sbjct: 490 CADVLAMAAR 519
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,584,104
Number of Sequences: 28460
Number of extensions: 70856
Number of successful extensions: 489
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 413
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 414
length of query: 332
length of database: 4,897,600
effective HSP length: 91
effective length of query: 241
effective length of database: 2,307,740
effective search space: 556165340
effective search space used: 556165340
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144516.5


