

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144515.4 + phase: 0
(292 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
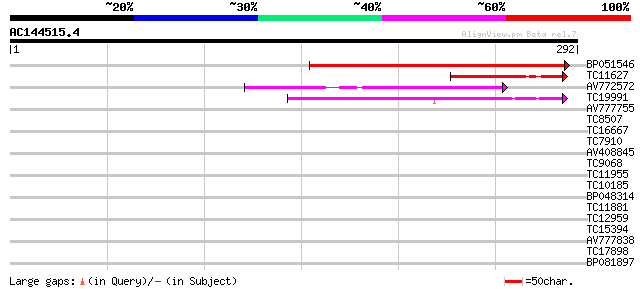
Score E
Sequences producing significant alignments: (bits) Value
BP051546 233 3e-62
TC11627 homologue to UP|Q9SVY1 (Q9SVY1) Zinc finger-like protein... 86 7e-18
AV772572 84 2e-17
TC19991 78 2e-15
AV777755 36 0.009
TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 31 0.21
TC16667 similar to UP|O64594 (O64594) F17O7.4 (AT1G70420/F17O7_4... 31 0.28
TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%) 30 0.48
AV408845 29 1.1
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 28 1.4
TC11955 similar to UP|Q9LHH9 (Q9LHH9) Protein arginine N-methylt... 28 1.4
TC10185 weakly similar to UP|Q9LLK5 (Q9LLK5) PSIEP1L protein, pa... 28 1.4
BP048314 28 1.8
TC11881 similar to PIR|T48303|T48303 meiosis specific-like prote... 28 2.4
TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%) 28 2.4
TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%) 28 2.4
AV777838 27 3.1
TC17898 similar to AAR27999 (AAR27999) TFIIA-L2 (Fragment), part... 27 4.0
BP081897 26 6.9
>BP051546
Length = 574
Score = 233 bits (594), Expect = 3e-62
Identities = 102/135 (75%), Positives = 115/135 (84%), Gaps = 1/135 (0%)
Frame = -2
Query: 155 RKGPDSLRGTQPAAMLRLPCYCCVQGCKNNINHPRAKPLKDFRTLQTHYKRKHGTKPFMC 214
RKGP+SLRGTQP AMLRLPCYCC GCKNNI+HPRAKPLKDFRTLQTHYKRKHG KPFMC
Sbjct: 573 RKGPESLRGTQPTAMLRLPCYCCAPGCKNNIDHPRAKPLKDFRTLQTHYKRKHGIKPFMC 394
Query: 215 RKCGKTFAVKGDWRTHEKNCGKLWYCTCGSDFKHKRSLKDHIRSFGKGHRR-LSSIDDRV 273
RKC K FAV+GDWRTHEKNCGKLWYC+CGSDFKHKRSLKDHI++FG GH ++ IDD
Sbjct: 393 RKCCKAFAVRGDWRTHEKNCGKLWYCSCGSDFKHKRSLKDHIKAFGNGHXAYINGIDDNC 214
Query: 274 FEEEKECVTGSEEDE 288
++ + GSE ++
Sbjct: 213 LLDQDDIEGGSEVEQ 169
>TC11627 homologue to UP|Q9SVY1 (Q9SVY1) Zinc finger-like protein (WIP2
protein), partial (15%)
Length = 602
Score = 85.9 bits (211), Expect = 7e-18
Identities = 41/60 (68%), Positives = 45/60 (74%)
Frame = +3
Query: 228 RTHEKNCGKLWYCTCGSDFKHKRSLKDHIRSFGKGHRRLSSIDDRVFEEEKECVTGSEED 287
RTHEKNCGKLWYC CGSDFKHKRSLKDHI++FG GH ID FEEE E + E+D
Sbjct: 3 RTHEKNCGKLWYCICGSDFKHKRSLKDHIKAFGSGHAAY-GIDG--FEEEDEPASEVEQD 173
>AV772572
Length = 491
Score = 84.3 bits (207), Expect = 2e-17
Identities = 46/137 (33%), Positives = 69/137 (49%), Gaps = 2/137 (1%)
Frame = -3
Query: 122 LVGPMQFACSICNKTFNRYNNMQMHMWGHGSEFRKGPDSLRGTQPAAMLRLPCYCCVQGC 181
L+ +F C ICN F R N+Q+H GH ++ S + +R Y C +
Sbjct: 468 LMATNRFVCVICN*GFQRDQNLQLHRRGHNLPWKLRQRSNKE------VRKKVYICPE-- 313
Query: 182 KNNINHPRAKPLKDFRTLQTHYKRKHGTKPFMCRKCGKTFAVKGDWRTHEKNCGKLWY-C 240
K ++H ++ L D ++ H+ RKHG K + C KC K +AV DW+ H + CG Y C
Sbjct: 312 KTCVHHDASRALGDLTGIKKHFSRKHGEKKWKCEKCSKKYAVPSDWKAHTQTCGTREYKC 133
Query: 241 TCGSDF-KHKRSLKDHI 256
CG+ F +H + HI
Sbjct: 132 DCGTLFSRHVINTHTHI 82
>TC19991
Length = 540
Score = 78.2 bits (191), Expect = 2e-15
Identities = 48/149 (32%), Positives = 78/149 (52%), Gaps = 5/149 (3%)
Frame = +1
Query: 144 QMHMWGHGSEFRKGPDSLRG-TQPAAMLRLPCYCC-VQGCKNNINHPRAKPLKDFRTLQT 201
+MHM HG++F+ + + A R + C +GC N H R + LK ++
Sbjct: 1 RMHMRAHGNQFKTAEALAKPLVEDAGGQRERRFSCPFKGCNRNKRHRRFRALKSVVCVRN 180
Query: 202 HYKRKHGTKPFMCRKC--GKTFAVKGDWRTHEKNCGK-LWYCTCGSDFKHKRSLKDHIRS 258
H+KR H K + C +C K+F+V D ++H ++C + W C+CG+ F K L HI
Sbjct: 181 HFKRSHCPKMYACDRCHKRKSFSVLSDLKSHMRHCSESRWRCSCGTTFSRKDKLFGHIAR 360
Query: 259 FGKGHRRLSSIDDRVFEEEKECVTGSEED 287
F +GH ++D+ + E+ K V +EED
Sbjct: 361 F-EGHVPAIALDE-IGEKGKHVVEENEED 441
>AV777755
Length = 373
Score = 35.8 bits (81), Expect = 0.009
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +2
Query: 122 LVGPMQFACSICNKTFNRYNNMQMHMWGH 150
L+ +F C ICNK F R N+Q+H GH
Sbjct: 194 LMATNRFVCEICNKGFQRDQNLQLHRRGH 280
>TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (40%)
Length = 620
Score = 31.2 bits (69), Expect = 0.21
Identities = 29/129 (22%), Positives = 47/129 (35%)
Frame = +3
Query: 18 NSSKSYLTSSSSSSSSTTQNETIQFFPILSANFSKDEEREVPQIEGFEFKEEKEIVALHI 77
+ S + + S S + ++T + P+ A+ K + K+ K I
Sbjct: 42 DDSDEEMDDADSDSDESDSDDTDEETPVKKADQGKKRANDSALKTPVSTKKAKNIT---- 209
Query: 78 GLPHDTKKYLDDEKKFFHFKEEEEEEKASKKTFQRFWIPTPAQILVGPMQFACSICNKTF 137
P T D KK H +K K + + P QF+CS C+K+F
Sbjct: 210 --PEKT-----DGKKTGHTATPHPGKKGGKNSESKAQTPKSGG------QFSCSPCSKSF 350
Query: 138 NRYNNMQMH 146
N + Q H
Sbjct: 351 NSESGFQQH 377
>TC16667 similar to UP|O64594 (O64594) F17O7.4 (AT1G70420/F17O7_4), partial
(23%)
Length = 953
Score = 30.8 bits (68), Expect = 0.28
Identities = 17/33 (51%), Positives = 23/33 (69%)
Frame = -3
Query: 15 KPSNSSKSYLTSSSSSSSSTTQNETIQFFPILS 47
+PS +K+ L SSSSSSSS+T + + F ILS
Sbjct: 279 EPSGFTKTKLNSSSSSSSSSTCSSSSTSFSILS 181
>TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%)
Length = 1035
Score = 30.0 bits (66), Expect = 0.48
Identities = 12/47 (25%), Positives = 20/47 (42%)
Frame = +3
Query: 104 KASKKTFQRFWIPTPAQILVGPMQFACSICNKTFNRYNNMQMHMWGH 150
+ S + + PA + + + CS+C KTF Y + H H
Sbjct: 291 RGSTAVTPKLTLSRPAPVTAEKLSYKCSVCEKTFPSYQALGGHKASH 431
>AV408845
Length = 429
Score = 28.9 bits (63), Expect = 1.1
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 3/75 (4%)
Frame = +3
Query: 12 EWLKPSNSSKSYLTSSSSSSSSTTQNETIQFFPILSANFSKDEEREVPQIEGFEFKEEKE 71
++ KP+NS S +SSSSSSSS++ F P+ +F + + + F F +
Sbjct: 69 DFSKPTNSPSSSSSSSSSSSSSSS------FQPLTLFSFFTPQPSSLNSL-CFIFSTMAQ 227
Query: 72 IVA---LHIGLPHDT 83
+VA +H + H T
Sbjct: 228 VVATRSIHTSISHPT 272
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 28.5 bits (62), Expect = 1.4
Identities = 14/27 (51%), Positives = 21/27 (76%)
Frame = -1
Query: 13 WLKPSNSSKSYLTSSSSSSSSTTQNET 39
+L PS+SS S +SSSSSSSS++ + +
Sbjct: 165 FLSPSSSSSSSSSSSSSSSSSSSSSSS 85
>TC11955 similar to UP|Q9LHH9 (Q9LHH9) Protein arginine N-methyltransferase
3-like protein, partial (24%)
Length = 572
Score = 28.5 bits (62), Expect = 1.4
Identities = 20/56 (35%), Positives = 28/56 (49%), Gaps = 4/56 (7%)
Frame = -3
Query: 137 FNRYNNMQMHMWGHGSE-FRKGPDS-LRGTQPAAML--RLPCYCCVQGCKNNINHP 188
FN + N+ H H + F + DS L TQP ++ L C+CC G N N+P
Sbjct: 426 FNGFVNLFNHAINHFYDSFLEAIDSTLAFTQPETIVFCNLRCHCCHLGTSLNCNNP 259
>TC10185 weakly similar to UP|Q9LLK5 (Q9LLK5) PSIEP1L protein, partial (57%)
Length = 504
Score = 28.5 bits (62), Expect = 1.4
Identities = 20/66 (30%), Positives = 28/66 (42%), Gaps = 2/66 (3%)
Frame = +1
Query: 186 NHPRAKPLKDFRTLQTHYKRKHGTKPFMCRKCGKTFAVKGDWRTHEKNCGKLWY--CTCG 243
NH AK + + + HY + G +P+M + GD R E NCG C C
Sbjct: 64 NHCSAKAVTSCKASEFHYYQLEGVEPYMSK------YTTGD-RVSETNCGNKCTKDCKCV 222
Query: 244 SDFKHK 249
F +K
Sbjct: 223 GYFYNK 240
>BP048314
Length = 394
Score = 28.1 bits (61), Expect = 1.8
Identities = 24/79 (30%), Positives = 38/79 (47%), Gaps = 15/79 (18%)
Frame = -3
Query: 17 SNSSKSYLTSSSSSSSSTT-------QNETIQFFPILSANF--------SKDEEREVPQI 61
S+SS S +SSSSSSSS++ + + F PI + +F S +E++E P+
Sbjct: 239 SSSSSSSSSSSSSSSSSSSCLSPAPPETSPLIFLPITNTSFKSIQTPPSSSEEKQESPKH 60
Query: 62 EGFEFKEEKEIVALHIGLP 80
+ + K A H P
Sbjct: 59 PATKPSKCKASTASHNPFP 3
>TC11881 similar to PIR|T48303|T48303 meiosis specific-like protein -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(34%)
Length = 631
Score = 27.7 bits (60), Expect = 2.4
Identities = 17/54 (31%), Positives = 27/54 (49%), Gaps = 1/54 (1%)
Frame = -3
Query: 215 RKCGKTFAVKGDWRTHEKNCGKLWY-CTCGSDFKHKRSLKDHIRSFGKGHRRLS 267
R CG+ + + G WR ++ G+++ CG H+ R G+GHRR S
Sbjct: 494 RGCGR*WWICGRWRCYQTGGGRIYP*GRCGRVQGHRAGW----RRSGRGHRRRS 345
>TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%)
Length = 517
Score = 27.7 bits (60), Expect = 2.4
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = -3
Query: 5 VEDNNFIEWLKPSNSSKSYLTSSSSSSSSTTQNETIQFFP 44
V+ NNF +SS S L+SSSSSS S++ + + P
Sbjct: 236 VDLNNFESHQDHHSSSSSSLSSSSSSSESSSSSSSSLLLP 117
>TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%)
Length = 1093
Score = 27.7 bits (60), Expect = 2.4
Identities = 18/51 (35%), Positives = 28/51 (54%)
Frame = +2
Query: 62 EGFEFKEEKEIVALHIGLPHDTKKYLDDEKKFFHFKEEEEEEKASKKTFQR 112
E E + K+ ++ PH K+ DDE K +EEE++ KA KKT ++
Sbjct: 5 EKAELSDAKDESQENVAAPH--KESDDDESKS---EEEEDKSKAQKKTSKK 142
>AV777838
Length = 565
Score = 27.3 bits (59), Expect = 3.1
Identities = 14/24 (58%), Positives = 18/24 (74%)
Frame = +1
Query: 16 PSNSSKSYLTSSSSSSSSTTQNET 39
PS+SS S +SSSSSSSS++ T
Sbjct: 214 PSSSSSSSSSSSSSSSSSSSMRAT 285
>TC17898 similar to AAR27999 (AAR27999) TFIIA-L2 (Fragment), partial (17%)
Length = 568
Score = 26.9 bits (58), Expect = 4.0
Identities = 12/22 (54%), Positives = 16/22 (72%)
Frame = -2
Query: 13 WLKPSNSSKSYLTSSSSSSSST 34
W + SNSS S+ + SSSSSS+
Sbjct: 90 WARSSNSSSSFSFNGSSSSSSS 25
>BP081897
Length = 305
Score = 26.2 bits (56), Expect = 6.9
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -3
Query: 170 LRLPCYCCVQGCKNNIN 186
LR C+ CV C NN+N
Sbjct: 177 LRXXCFLCVYPCHNNVN 127
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,564,900
Number of Sequences: 28460
Number of extensions: 113134
Number of successful extensions: 1424
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 1031
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1232
length of query: 292
length of database: 4,897,600
effective HSP length: 90
effective length of query: 202
effective length of database: 2,336,200
effective search space: 471912400
effective search space used: 471912400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144515.4


