

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144515.13 + phase: 0
(381 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
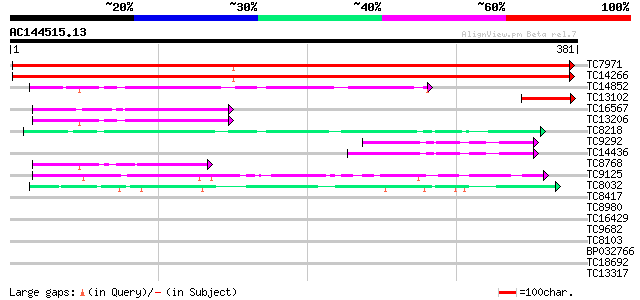
Sequences producing significant alignments: (bits) Value
TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported... 603 e-173
TC14266 homologue to PIR|C86216|C86216 protein T23G18.6 [importe... 596 e-171
TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydrata... 80 7e-16
TC13102 67 4e-12
TC16567 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carbox... 56 1e-08
TC13206 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carbox... 55 1e-08
TC8218 homologue to UP|Q93VR3 (Q93VR3) AT5g28840/F7P1_20, complete 52 2e-07
TC9292 homologue to UP|AAR07600 (AAR07600) Fiber dTDP-glucose 4-... 51 4e-07
TC14436 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carbox... 50 8e-07
TC8768 homologue to UP|Q9SMJ5 (Q9SMJ5) DTDP-glucose 4-6-dehydrat... 49 1e-06
TC9125 similar to UP|Q40316 (Q40316) Vestitone reductase, complete 44 6e-05
TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-... 41 4e-04
TC8417 similar to UP|GAE1_PEA (Q43070) UDP-glucose 4-epimerase ... 39 0.001
TC8980 similar to UP|Q7X998 (Q7X998) mRNA-binding protein (Fragm... 37 0.007
TC16429 homologue to UP|Q93VR3 (Q93VR3) AT5g28840/F7P1_20, parti... 32 0.23
TC9682 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratas... 32 0.23
TC8103 homologue to UP|AAR07600 (AAR07600) Fiber dTDP-glucose 4-... 31 0.30
BP032766 31 0.39
TC18692 similar to PIR|T00419|T00419 dTDP-glucose 4-6-dehydratas... 30 0.51
TC13317 similar to UP|Q9SPJ5 (Q9SPJ5) Dihydroflavonol-4-reductas... 30 0.87
>TC7971 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(98%)
Length = 1662
Score = 603 bits (1554), Expect = e-173
Identities = 284/380 (74%), Positives = 335/380 (87%), Gaps = 3/380 (0%)
Frame = +3
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-PW 61
RV+LDG PI P++IC+IG GGFIGSHL EKLMSET HK + +DV S+K+ HLL+ + PW
Sbjct: 60 RVDLDGNPIAPLTICMIGAGGFIGSHLCEKLMSETPHKVLALDVYSDKIKHLLEPDNLPW 239
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
RI FH++NIK+DSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K+
Sbjct: 240 HGRIHFHRLNIKHDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVKY 419
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEE--YRKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+ENNKRLIHFSTCEV+GKTIGSFLP++ R++P YY LKED SPCIFG + KQRWSYA
Sbjct: 420 CSENNKRLIHFSTCEVYGKTIGSFLPKDSPLRQDPAYYVLKEDESPCIFGSIEKQRWSYA 599
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RLIYAE AEN ++FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLLR
Sbjct: 600 CAKQLIERLIYAEGAENNMEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLLR 779
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
GEPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP RANGHIFNVGNP NEV+V+QLAE+MI
Sbjct: 780 GEPLKLVDGGESQRTFVYIKDAIEAVLLMIENPGRANGHIFNVGNPHNEVTVRQLAEMMI 959
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDS 359
+VY+KV+G TLDVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+S
Sbjct: 960 QVYSKVSGEQAPETPTLDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLES 1139
Query: 360 TLQYQHQTYSHAIKKELSKP 379
TL YQH+TY+ A+KK ++KP
Sbjct: 1140TLTYQHRTYAEAVKKAIAKP 1199
>TC14266 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, complete
Length = 1607
Score = 596 bits (1537), Expect = e-171
Identities = 285/380 (75%), Positives = 331/380 (87%), Gaps = 3/380 (0%)
Frame = +1
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-PW 61
RV+LDG PI P++IC+IG GGFIGSHL EKLMSET HK + +DV S+K+ HLL+ PW
Sbjct: 43 RVDLDGNPIKPLTICMIGAGGFIGSHLCEKLMSETPHKVLALDVYSDKIKHLLEPDTLPW 222
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
A RI FH++NIK+DSRLE L+K SDLTINLAAICTPADYNTRPLDTIYSNFIDA+PV K+
Sbjct: 223 AGRIHFHRLNIKHDSRLEGLIKMSDLTINLAAICTPADYNTRPLDTIYSNFIDALPVAKY 402
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEE--YRKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+ENNKRLIHFSTCEV+GKTIGSFLP++ R++P YY LKED SPCIFG + KQRWSYA
Sbjct: 403 CSENNKRLIHFSTCEVYGKTIGSFLPKDSPLRQDPAYYVLKEDESPCIFGSIEKQRWSYA 582
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RLIYAE AENG++FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLLR
Sbjct: 583 CAKQLIERLIYAEGAENGMEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLLR 762
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP RANGHIFNVGNP NEV+V+QLAE+M
Sbjct: 763 REPLKLVDGGESQRTFIYIKDAIEAVLLMIENPARANGHIFNVGNPHNEVTVRQLAEMMT 942
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDS 359
KVY+KV+ +DVSS+ FYG+GYDDSD+RIPDMTII KQLGW PKTSL DLL+S
Sbjct: 943 KVYSKVSEEAPLENPAIDVSSKEFYGEGYDDSDKRIPDMTIINKQLGWNPKTSLWDLLES 1122
Query: 360 TLQYQHQTYSHAIKKELSKP 379
TL YQH+TY+ AIKK ++KP
Sbjct: 1123TLTYQHRTYAEAIKKAIAKP 1182
>TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydratase homolog
D18 (At3g62830/F26K9_260), partial (70%)
Length = 1470
Score = 79.7 bits (195), Expect = 7e-16
Identities = 73/275 (26%), Positives = 118/275 (42%), Gaps = 4/275 (1%)
Frame = +3
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMN 71
+ I + GG GF+GSHL ++LM+ IV+D + K N + +P R E + +
Sbjct: 759 LRIVVTGGAGFVGSHLVDRLMAR-GDSVIVVDNFFTGRKENVMHHFGNP---RFELIRHD 926
Query: 72 IKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIH 131
+ +E L+ D +LA +P Y P+ TI +N + + ++ R +
Sbjct: 927 V-----VEPLLLEVDQIYHLACPASPVHYKFNPVKTIKTNVVGTLNMLGLAKRVGARFLL 1091
Query: 132 FSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYA 191
ST EV+G + + PQ +V+P R Y K+ + L
Sbjct: 1092TSTSEVYGDPL---------QHPQKETYWGNVNPI------GVRSCYDEGKRTAETLTMD 1226
Query: 192 EHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHS 251
H G++ I R +N GPRM G RV + F L EPL + G
Sbjct: 1227YHRGAGVEVRIARIFNTYGPRMCLDDG---------RVASNFDAQALTKEPLTVYGDGKQ 1379
Query: 252 QRTFLYIKDAIEAVMLMIDNPDRANGHI--FNVGN 284
R+F Y+ D +E +M +++ H+ FN+GN
Sbjct: 1380TRSFQYVSDLVEGLMRLMEGE-----HVGPFNLGN 1469
>TC13102
Length = 461
Score = 67.4 bits (163), Expect = 4e-12
Identities = 30/36 (83%), Positives = 33/36 (91%)
Frame = +2
Query: 345 LGWKPKTSLDDLLDSTLQYQHQTYSHAIKKELSKPS 380
LGWKPKT LD+LL TL+YQH+TYSHAIKKELSKPS
Sbjct: 2 LGWKPKTPLDELLQVTLKYQHETYSHAIKKELSKPS 109
>TC16567 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carboxy-lyase ,
partial (46%)
Length = 610
Score = 55.8 bits (133), Expect = 1e-08
Identities = 42/135 (31%), Positives = 60/135 (44%)
Frame = +3
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
I + GG GFIGSHL ++LM ++ IV D D W F I++D
Sbjct: 192 ILITGGAGFIGSHLVDRLMENEKNEVIVAD---NYFTGSRDNLKKWIGHPRFEL--IRHD 356
Query: 76 SRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFSTC 135
ETL+ D +LA +P Y P+ TI +N I + ++ R++ ST
Sbjct: 357 V-TETLMIEVDQIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTSTS 533
Query: 136 EVFGKTIGSFLPEEY 150
EV G + PE Y
Sbjct: 534 EVDGDPLIHPQPESY 578
>TC13206 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carboxy-lyase ,
partial (51%)
Length = 650
Score = 55.5 bits (132), Expect = 1e-08
Identities = 42/137 (30%), Positives = 63/137 (45%), Gaps = 2/137 (1%)
Frame = +2
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + GG GFIGSHL ++LM ++ IV D + K N HP R E + ++
Sbjct: 221 ILVTGGAGFIGSHLVDRLMENEKNEVIVADNYFTGSKDNLKKWIGHP---RFELIRHDV- 388
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
E L+ D +LA +P Y P+ TI +N I + ++ R++ S
Sbjct: 389 ----TEPLLIEVDQIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTS 556
Query: 134 TCEVFGKTIGSFLPEEY 150
T EV+G + PE Y
Sbjct: 557 TSEVYGDPLLHPQPESY 607
>TC8218 homologue to UP|Q93VR3 (Q93VR3) AT5g28840/F7P1_20, complete
Length = 1590
Score = 52.0 bits (123), Expect = 2e-07
Identities = 77/354 (21%), Positives = 143/354 (39%), Gaps = 3/354 (0%)
Frame = +3
Query: 10 PIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQ 69
P + I + G GGFI SH+ +L +E + +I +K H+ + EFH
Sbjct: 135 PSEKLKISITGAGGFIASHIARRLKTEGHY---IIASDWKKNEHMSEDMF----CDEFHL 293
Query: 70 MNIKNDSRLETLVKASDLTINLAAICTPADY-NTRPLDTIYSNFIDAIPVIKFCTENN-K 127
++++ + K D NLAA + + +Y+N + + +I+ N K
Sbjct: 294 VDLRVMDNCLKVTKGVDHVFNLAADMGGMGFIQSNHSVIMYNNTMISFNMIEAARINGIK 473
Query: 128 RLIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDR 187
R + S+ ++ PE + E + D P + + +Y K T+
Sbjct: 474 RFFYASSACIY--------PEFKQLETNVSLKEADAWPA------EPQDAYGLEKLATEE 611
Query: 188 LIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRG-EPLKLV 246
L + + G++ I R +N GP + G + A F + + ++
Sbjct: 612 LCKHYNKDFGIECRIGRFHNIYGPFGTW-------KGGREKAPAAFCRKTITSTDRFEMW 770
Query: 247 DGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVA 306
G R+F +I + +E V+ + + R N+G+ D VS+ ++AE+++ K
Sbjct: 771 GDGLQTRSFTFIDECVEGVLRLTKSDFR---EPVNIGS-DEMVSMNEMAEIVLSFENK-- 932
Query: 307 GVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDST 360
+P + + G + R D T+I ++LGW P L D L T
Sbjct: 933 NIP------------IHHIPGPEGVRGRNSDNTLIKEKLGWAPTMKLKDGLRIT 1058
>TC9292 homologue to UP|AAR07600 (AAR07600) Fiber dTDP-glucose
4-6-dehydratase (Fragment), partial (66%)
Length = 641
Score = 50.8 bits (120), Expect = 4e-07
Identities = 34/118 (28%), Positives = 59/118 (49%)
Frame = +2
Query: 238 LRGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAEL 297
LRGEPL + G R+F Y+ D ++ ++ +++ D N+GNP E ++ +LAE
Sbjct: 14 LRGEPLTVQSPGTQTRSFCYVSDLVDGLIRLMEGSDTGP---INLGNP-GEFTMTELAET 181
Query: 298 MIKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDD 355
+ ++ P+ + ++ + DD R PD+T + LGW+PK L D
Sbjct: 182 VKELIN-----PKVEIKMVENTP--------DDPRPRKPDITKAKELLGWEPKVKLRD 316
>TC14436 homologue to UP|Q9AV98 (Q9AV98) UDP-D-glucuronate carboxy-lyase ,
partial (40%)
Length = 719
Score = 49.7 bits (117), Expect = 8e-07
Identities = 35/128 (27%), Positives = 62/128 (48%)
Frame = +2
Query: 228 RVLACFSNNLLRGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDN 287
RV++ F +R EPL + G R+F Y+ D ++ ++ ++D D N+GNP
Sbjct: 26 RVVSNFIAQAIRNEPLTVQSPGTQTRSFCYVSDLVDGLIRLMDGSDTGP---INLGNP-G 193
Query: 288 EVSVKQLAELMIKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGW 347
E ++ +LAE + ++ P + ++ + DD +R P +T + LGW
Sbjct: 194 EFTMLELAETVKELIN-----PNVEIKIVENTP--------DDPRQRKPIITKAKELLGW 334
Query: 348 KPKTSLDD 355
+PK L D
Sbjct: 335 EPKVKLRD 358
>TC8768 homologue to UP|Q9SMJ5 (Q9SMJ5) DTDP-glucose 4-6-dehydratase ,
partial (43%)
Length = 562
Score = 48.9 bits (115), Expect = 1e-06
Identities = 39/123 (31%), Positives = 58/123 (46%), Gaps = 2/123 (1%)
Frame = +2
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVID--VSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + GG GFIGSHL ++LM ++ IV D + K N HP R E I+
Sbjct: 218 ILVTGGAGFIGSHLVDRLMENEKNEVIVADNFFTGSKDNLKKWIGHP---RFEL----IR 376
Query: 74 NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFS 133
+D + LV+ D +LA +P Y P+ TI +N I + ++ R++ S
Sbjct: 377 HDVTEQLLVEV-DQIYHLACPASPIFYKYNPVKTIKTNVIGTLNMLGLAKRVGARILLTS 553
Query: 134 TCE 136
T E
Sbjct: 554 TSE 562
>TC9125 similar to UP|Q40316 (Q40316) Vestitone reductase, complete
Length = 1200
Score = 43.5 bits (101), Expect = 6e-05
Identities = 76/360 (21%), Positives = 153/360 (42%), Gaps = 13/360 (3%)
Frame = +3
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSS---EKVNHLLDKSHPWANRIEFHQMNI 72
+C+ GG GF+ S + ++L+ + I +S V+ L + + +++ ++
Sbjct: 60 VCVTGGTGFLASWIIKRLLEDGYSVNTTIRSNSGGKRDVSFLTNLPGATSEKLQIFNADL 239
Query: 73 KNDSRLETLVKASDLTINLAAICTPADYNTR-PLDTIYSNFID-AIPVIKFCTENN--KR 128
E+ A + + + TP D+ P + + +D A+ ++K C + KR
Sbjct: 240 CIP---ESFGPAVEGCVGIFHTATPVDFAVNEPEEVVTKRTVDGALGILKACVNSKTVKR 410
Query: 129 LIHFST---CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMT 185
+++ S+ GK G L E Y + L V P FG WSYA +K +
Sbjct: 411 VVYTSSGSAASFSGKDGGDALDESYWSDVD---LLRKVKP--FG------WSYAVSKTLA 557
Query: 186 DRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKL 245
++ + ++GL+ + P +GP + P + D V R L L+ G+ ++
Sbjct: 558 EKAVLEFGEKHGLEVVTLIPTFVVGPWV--CPKL---PDSVERALV-----LVLGKKEQI 707
Query: 246 VDGGHSQRTFLYIKDAIEAVMLMIDNPD---RANGHIFNVGNPDNEVSVKQLAELMIKVY 302
G + +++ D A + ++++P+ R N F V ++++++L+ Y
Sbjct: 708 ---GVIRFHMVHVDDVARAHIFLLEHPNPKGRYNCSPF-------IVPIEEMSKLLSAKY 857
Query: 303 AKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQ 362
PE + +++ + K D +I D G++ K SL+DL D +Q
Sbjct: 858 ------PEYQIPSVEELKGIEGFKQPDLISNKIRD-------AGFEFKYSLEDLFDDAIQ 998
>TC8032 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-like
protein, partial (97%)
Length = 2230
Score = 40.8 bits (94), Expect = 4e-04
Identities = 80/387 (20%), Positives = 148/387 (37%), Gaps = 30/387 (7%)
Frame = +3
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNI- 72
+S+ + G GF+GSH++ L + +D ++ + L K+ + H + I
Sbjct: 426 MSVLVTGAAGFVGSHVSLAL-KRRGDGVVGLDNFNDYYDPSLKKAR--KTLLTTHGVFIV 596
Query: 73 ---KNDSRLETLVKASDL-----TINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTE 124
ND++L L K D+ ++LAA P ++SN + +++ C
Sbjct: 597 EGDVNDAKL--LAKLFDVVAFTHVMHLAAQAGVRYAMENPHSYVHSNIAALVTLMEACKS 770
Query: 125 NNKR--LIHFSTCEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAK 182
N + ++ S+ V+G L E V + YA K
Sbjct: 771 ANPQPAIVWASSSSVYG-------------------LNEKVPFSESDRTDQPASLYAATK 893
Query: 183 QMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEP 242
+ + + + + GL T +R + V GP F+ N+L+G+P
Sbjct: 894 KAGEEITHTYNHIYGLSITGLRFFT-----------VYGPWGRPDMAYFSFTRNILQGKP 1040
Query: 243 LKLVDGGHS---QRTFLYIKDAIEAVMLMIDNPDRANG-----------HIFNVGNPDNE 288
+ + G + R F YI D ++ + +D ++ G IFN+GN +
Sbjct: 1041ITVYRGKNRVDLARDFTYIDDIVKGCVGSLDTAGKSTGSGGKKKGAAPYRIFNLGN-TSP 1217
Query: 289 VSVKQLAELM---IKVYAK--VAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITK 343
V+V L ++ +KV AK V +P G D +++ +
Sbjct: 1218VTVPTLVSILEGHLKVKAKRNVVDMP-----------------GNGDVPFTHANISSARR 1346
Query: 344 QLGWKPKTSLDDLLDSTLQYQHQTYSH 370
+LG+KP T L L +++ Y +
Sbjct: 1347ELGYKPTTDLQTGLKKFVKWYLSYYGY 1427
>TC8417 similar to UP|GAE1_PEA (Q43070) UDP-glucose 4-epimerase
(Galactowaldenase) (UDP-galactose 4-epimerase) , partial
(65%)
Length = 789
Score = 39.3 bits (90), Expect = 0.001
Identities = 30/132 (22%), Positives = 64/132 (47%), Gaps = 7/132 (5%)
Frame = +3
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSS---EKVNHLLDKSHPWANR-IEFHQMN 71
I + GG GFIG+H +L+ + H +I+ + + E V+ + + P ++ ++F Q +
Sbjct: 111 ILVTGGAGFIGTHTVVQLLHDGFHVSIIDNFDNSCMEAVDRVREVVGPHLSKNLQFTQGD 290
Query: 72 IKNDSRLETLVKAS--DLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENN-KR 128
++N LE + + D I+ A + + P N + I + + + N K+
Sbjct: 291 LRNKDDLEEIFSKTKFDAVIHFAGLKAVGESVANPRRYFDFNLVGTINLYQAMAKYNVKK 470
Query: 129 LIHFSTCEVFGK 140
++ S+ V+G+
Sbjct: 471 MVFSSSATVYGQ 506
>TC8980 similar to UP|Q7X998 (Q7X998) mRNA-binding protein (Fragment),
partial (40%)
Length = 598
Score = 36.6 bits (83), Expect = 0.007
Identities = 22/77 (28%), Positives = 40/77 (51%)
Frame = +1
Query: 233 FSNNLLRGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVK 292
F + ++R P+ + + G +++D + L ++NPD AN IFN + D V++
Sbjct: 25 FFDRIVRDRPVPIPNSGLQLTNIAHVRDLSSMLTLAVENPDAANHTIFNCVS-DRAVTLD 201
Query: 293 QLAELMIKVYAKVAGVP 309
+A K+ A+ AG P
Sbjct: 202 GMA----KLCAQAAGRP 240
>TC16429 homologue to UP|Q93VR3 (Q93VR3) AT5g28840/F7P1_20, partial (49%)
Length = 632
Score = 31.6 bits (70), Expect = 0.23
Identities = 24/84 (28%), Positives = 37/84 (43%)
Frame = +1
Query: 10 PIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQ 69
P + I + G GGFI SH+ +L SE H I D + + H+ + EFH
Sbjct: 121 PSEKLRISITGAGGFIASHIARRLKSE-GHYIIASDWKTNE--HMTEDMFCH----EFHL 279
Query: 70 MNIKNDSRLETLVKASDLTINLAA 93
++++ + D NLAA
Sbjct: 280 IDLRVMDNCLKVTNGVDHVFNLAA 351
>TC9682 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratase-like
protein, partial (29%)
Length = 731
Score = 31.6 bits (70), Expect = 0.23
Identities = 14/32 (43%), Positives = 21/32 (64%)
Frame = +3
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVID 45
+ + + GG GF+GSHL +KL+ + IVID
Sbjct: 519 LRVVVTGGAGFVGSHLVDKLIGR-GNDVIVID 611
>TC8103 homologue to UP|AAR07600 (AAR07600) Fiber dTDP-glucose
4-6-dehydratase (Fragment), partial (51%)
Length = 630
Score = 31.2 bits (69), Expect = 0.30
Identities = 24/75 (32%), Positives = 35/75 (46%)
Frame = +2
Query: 281 NVGNPDNEVSVKQLAELMIKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTI 340
N+GNP E ++ +LAE V E ST+++ DD +R PD+
Sbjct: 38 NIGNP-GEFTMIELAE----------NVKELINSTVEIK---MIENTPDDPRQRKPDIAK 175
Query: 341 ITKQLGWKPKTSLDD 355
+ LGW+PK L D
Sbjct: 176 AKELLGWEPKVKLRD 220
>BP032766
Length = 535
Score = 30.8 bits (68), Expect = 0.39
Identities = 26/126 (20%), Positives = 59/126 (46%), Gaps = 6/126 (4%)
Frame = +1
Query: 15 SICLIGGGGFIGSHLTEKLMSE--TSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNI 72
++C+ G GFIGS L +L+ T + + +KV HLL+ ++ + ++
Sbjct: 73 TVCVTGAAGFIGSWLVMRLIERGYTVRATVRDPANMKKVKHLLELPDA-KTKLSLWKADL 249
Query: 73 KNDSRLETLVKASDLTINLAAICTPADYNTR-PLDTIYSNFIDA-IPVIKFC--TENNKR 128
+ + ++ ++A TP D+ ++ P + + I+ + ++K C + +R
Sbjct: 250 AEEGSFDEAIRGCTGVFHVA---TPMDFESKDPENEVIKPTINGLLDILKACEKAKTVRR 420
Query: 129 LIHFST 134
L+ S+
Sbjct: 421 LVFTSS 438
>TC18692 similar to PIR|T00419|T00419 dTDP-glucose 4-6-dehydratase homolog
At2g47650 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (14%)
Length = 487
Score = 30.4 bits (67), Expect = 0.51
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 1/40 (2%)
Frame = +3
Query: 315 TLDVSSEV-FYGKGYDDSDRRIPDMTIITKQLGWKPKTSL 353
T+D S+++ F DD R PD++ + LGWKP SL
Sbjct: 12 TIDPSAKIEFRPNTEDDPHMRKPDISKAKELLGWKPSVSL 131
>TC13317 similar to UP|Q9SPJ5 (Q9SPJ5) Dihydroflavonol-4-reductase DFR1,
partial (36%)
Length = 426
Score = 29.6 bits (65), Expect = 0.87
Identities = 25/113 (22%), Positives = 51/113 (45%), Gaps = 5/113 (4%)
Frame = +2
Query: 15 SICLIGGGGFIGSHLTEKLMSETSHKAIVID---VSSEKVNHLLDKSHPWANRIEFHQMN 71
++C+ G GFIGS L +LM + + +KV HLL+ N + +
Sbjct: 65 TVCVTGSTGFIGSWLVMRLMERGYMVRATVQRDPDNMKKVKHLLELPGAKTN-LTIWNAD 241
Query: 72 IKNDSRLETLVKASDLTINLAAICTPADYNTR-PLDTIYSNFIDAI-PVIKFC 122
+ + + +K ++A +P D+N++ P + + I+ + ++K C
Sbjct: 242 LTEEGSFDEAIKGCSGVFHVA---SPMDFNSKDPENEVIKPAINGVLDIMKAC 391
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,456,144
Number of Sequences: 28460
Number of extensions: 85257
Number of successful extensions: 426
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 420
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 420
length of query: 381
length of database: 4,897,600
effective HSP length: 92
effective length of query: 289
effective length of database: 2,279,280
effective search space: 658711920
effective search space used: 658711920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144515.13


