

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
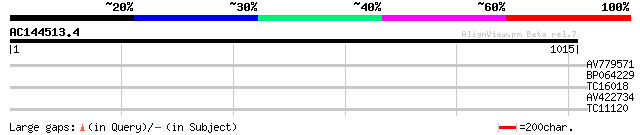
Query= AC144513.4 - phase: 0 /pseudo
(1015 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV779571 29 4.3
BP064229 28 7.3
TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%) 28 9.6
AV422734 28 9.6
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 28 9.6
>AV779571
Length = 538
Score = 28.9 bits (63), Expect = 4.3
Identities = 12/25 (48%), Positives = 21/25 (84%)
Frame = -2
Query: 18 IKNLSHVFSMVLSLILLLLIVLLVI 42
I NL ++ ++LSL+LLLL++LL++
Sbjct: 333 IANLLIIYHLLLSLLLLLLLLLLIV 259
>BP064229
Length = 484
Score = 28.1 bits (61), Expect = 7.3
Identities = 12/25 (48%), Positives = 18/25 (72%)
Frame = -3
Query: 382 LVLDLLFPLMLWRWLVTELIMICLV 406
L+LDLLF L + RW++ E I C++
Sbjct: 242 LLLDLLFTLPVSRWMMKEPIHYCIL 168
>TC16018 UP|Q8MZB8 (Q8MZB8) AT15479p (CG31542-PA), partial (7%)
Length = 425
Score = 27.7 bits (60), Expect = 9.6
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = +2
Query: 962 HTRDLPTYGHLSNCPASHH 980
HT TYGH+S+ P SHH
Sbjct: 311 HTSHDVTYGHVSHPPQSHH 367
>AV422734
Length = 448
Score = 27.7 bits (60), Expect = 9.6
Identities = 15/41 (36%), Positives = 26/41 (62%)
Frame = -2
Query: 1 LPILMWFFSQKPNLIKLIKNLSHVFSMVLSLILLLLIVLLV 41
L +L+ Q+ L+ L+ L H +VL ++LLL+++LLV
Sbjct: 213 LLLLLQLL*QQEMLLLLLLLLKHQLLLVLLMLLLLMLMLLV 91
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 27.7 bits (60), Expect = 9.6
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -1
Query: 369 FMRSLLPILRRCLLVLDLLFPLMLWR 394
F+ SL + RC+ ++D LFP M WR
Sbjct: 384 FLISLSVLPGRCVAIVDHLFPRMEWR 307
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.368 0.167 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,868,810
Number of Sequences: 28460
Number of extensions: 315873
Number of successful extensions: 5211
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 2920
Number of HSP's successfully gapped in prelim test: 213
Number of HSP's that attempted gapping in prelim test: 2141
Number of HSP's gapped (non-prelim): 3412
length of query: 1015
length of database: 4,897,600
effective HSP length: 99
effective length of query: 916
effective length of database: 2,080,060
effective search space: 1905334960
effective search space used: 1905334960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144513.4


