

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.2 - phase: 0
(215 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
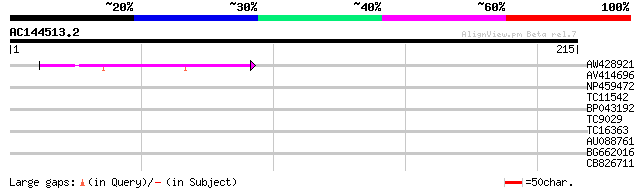
Score E
Sequences producing significant alignments: (bits) Value
AW428921 41 2e-04
AV414696 37 0.002
NP459472 RAB7A [Lotus japonicus] 29 0.71
TC11542 similar to UP|O04335 (O04335) At2g30600 protein, partial... 29 0.71
BP043192 28 1.2
TC9029 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosidase... 28 1.6
TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete 28 1.6
AU088761 27 2.7
BG662016 26 4.6
CB826711 26 6.0
>AW428921
Length = 359
Score = 40.8 bits (94), Expect = 2e-04
Identities = 23/84 (27%), Positives = 42/84 (49%), Gaps = 2/84 (2%)
Frame = +2
Query: 12 VILLLFLLITGLGSVINASPCSN-CGDFEVPYPLSTNDDCGDKRYKIYCNNDSLE-FLSA 69
+++ L LL+ + + + SPC N CG + YP D CG +++ N ++ E F
Sbjct: 83 LVITLTLLLQAIPT-LTLSPCKNSCGTIPINYPFGLEDGCGAPQFRHMLNWNTTELFFET 259
Query: 70 TGTYYKILKIDTSANKLVIKPPNI 93
YK+ I+ + +VI P++
Sbjct: 260 PSGSYKVQSINYEKSSMVIYDPSM 331
>AV414696
Length = 270
Score = 37.4 bits (85), Expect = 0.002
Identities = 24/69 (34%), Positives = 37/69 (52%), Gaps = 3/69 (4%)
Frame = +1
Query: 8 SLKMVILLLFLLITGLGSVINASPCS-NCGDFEVPYPLSTNDDCGDKRYK--IYCNNDSL 64
S+ ++ + LFLL S+I A C NCG + YP + CGD R++ + C+ L
Sbjct: 52 SVPLLSIFLFLL----PSLILAQICQRNCGQ-TLKYPFGSGPGCGDPRFQPHVTCSEQRL 216
Query: 65 EFLSATGTY 73
F + TG+Y
Sbjct: 217 TFTTHTGSY 243
>NP459472 RAB7A [Lotus japonicus]
Length = 618
Score = 28.9 bits (63), Expect = 0.71
Identities = 12/33 (36%), Positives = 21/33 (63%)
Frame = -1
Query: 1 MYPCIWNSLKMVILLLFLLITGLGSVINASPCS 33
++ C+WNSLK+ +L++ + G S+ N CS
Sbjct: 588 LWSCLWNSLKIYVLVMLI---G*SSLGNTKECS 499
>TC11542 similar to UP|O04335 (O04335) At2g30600 protein, partial (21%)
Length = 756
Score = 28.9 bits (63), Expect = 0.71
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +2
Query: 4 CIWNSLKMVILLLFLLITGLGSVINASPCS 33
CIW L LL+FL +++ASPCS
Sbjct: 326 CIWKLLFFFFLLIFLCHHTTLGILSASPCS 415
>BP043192
Length = 453
Score = 28.1 bits (61), Expect = 1.2
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -2
Query: 150 EEKVEEGNGCMNTLCC 165
EEKV++G GC T CC
Sbjct: 320 EEKVKQGTGCCCTCCC 273
>TC9029 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosidase, partial
(73%)
Length = 1636
Score = 27.7 bits (60), Expect = 1.6
Identities = 17/67 (25%), Positives = 36/67 (53%), Gaps = 3/67 (4%)
Frame = +3
Query: 38 FEVPYPLSTNDDCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTS---ANKLVIKPPNIF 94
F + Y N+DCG K Y+ + L+F SA + Y ++I ++ +++ + P +I+
Sbjct: 1152 FNLKYVAVGNEDCGKKNYR----GNYLKFYSAIRSAYPDIQIISNCDGSSRPLDHPADIY 1319
Query: 95 KHTCYSS 101
+ Y++
Sbjct: 1320 DYHIYTN 1340
>TC16363 UP|Q84RQ3 (Q84RQ3) Sucrose transporter 4 protein, complete
Length = 2164
Score = 27.7 bits (60), Expect = 1.6
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = -3
Query: 140 CSSNSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHK 177
C++NS CR K E N T+C + +V N HK
Sbjct: 833 CTTNSACRG---KCESKNPVPTTVCSSRIAKNVTNGHK 729
>AU088761
Length = 254
Score = 26.9 bits (58), Expect = 2.7
Identities = 20/59 (33%), Positives = 25/59 (41%), Gaps = 3/59 (5%)
Frame = -2
Query: 113 SLPFNISTLNT---VMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYL 168
SLPF L + +ML + LQ+ L S S NGC N+ CHYL
Sbjct: 244 SLPFPTGLLKSFVVMMLKTLLPSFLQNLLAIFSASF----------SNGCSNSFLCHYL 98
>BG662016
Length = 459
Score = 26.2 bits (56), Expect = 4.6
Identities = 23/82 (28%), Positives = 38/82 (46%), Gaps = 3/82 (3%)
Frame = +1
Query: 113 SLPFNISTLNTVMLLNCSDNILQS-PLNCSSNSICRQFEEKVEEGNGCMNTLCCHY--LK 169
SLP +I L+ + LLN S N ++S P + S + + + +T+ LK
Sbjct: 31 SLPNSIGCLSKLKLLNVSGNFIESLPKTIENCSALEELNANFNKLSKLPDTIGFELINLK 210
Query: 170 DSVMNSHKIRLRVGSCTAYTCL 191
+NS+K+ L S + T L
Sbjct: 211 KLAVNSNKLVLLPSSTSHLTAL 276
>CB826711
Length = 444
Score = 25.8 bits (55), Expect = 6.0
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -1
Query: 3 PCIWNSLKMVILLLFLLITGLGSVINASPCS 33
P ++K++ILLLFLL+T S+ P S
Sbjct: 366 PATATAVKIIILLLFLLLTFAHSLCRGFPHS 274
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,214,294
Number of Sequences: 28460
Number of extensions: 80524
Number of successful extensions: 415
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 414
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 415
length of query: 215
length of database: 4,897,600
effective HSP length: 86
effective length of query: 129
effective length of database: 2,450,040
effective search space: 316055160
effective search space used: 316055160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144513.2


