

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
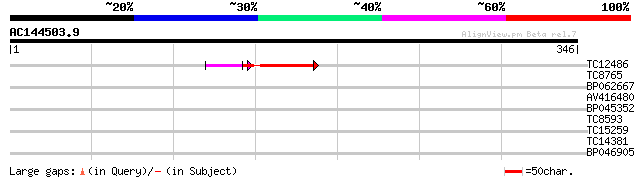
Score E
Sequences producing significant alignments: (bits) Value
TC12486 45 8e-08
TC8765 31 0.35
BP062667 27 3.9
AV416480 27 3.9
BP045352 27 3.9
TC8593 homologue to UP|AAR83877 (AAR83877) 60S ribosomal protein... 27 6.6
TC15259 homologue to UP|AAR83877 (AAR83877) 60S ribosomal protei... 26 8.6
TC14381 similar to UP|RNS2_ARATH (P42814) Ribonuclease 2 precurs... 26 8.6
BP046905 26 8.6
>TC12486
Length = 742
Score = 45.4 bits (106), Expect(2) = 8e-08
Identities = 22/46 (47%), Positives = 30/46 (64%)
Frame = +2
Query: 143 CMLFADVVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECN 188
C L+ +V E RE++N RL RQ L+ YGF L++S EYMEC+
Sbjct: 119 CELYVEV---RELREDINARLAICRQTLQTYGF*LNKSKAEYMECS 247
Score = 26.9 bits (58), Expect(2) = 8e-08
Identities = 14/29 (48%), Positives = 17/29 (58%)
Frame = +3
Query: 120 LSPYLFTLVLDVLTEHIQELAPRCMLFAD 148
LS Y F +VLD+L E I CMLF +
Sbjct: 39 LSHYHFIVVLDMLIEQI*ATMQLCMLFVN 125
>TC8765
Length = 748
Score = 30.8 bits (68), Expect = 0.35
Identities = 16/31 (51%), Positives = 21/31 (67%)
Frame = +3
Query: 316 VGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
+ V P V KL E RLR +GHV R ++A+VR
Sbjct: 531 IWVVPNV*KLTEIRLRWYGHVPSRLMEALVR 623
>BP062667
Length = 528
Score = 27.3 bits (59), Expect = 3.9
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -2
Query: 81 IAYIRAIKDMYEGASTSVRTQDGTTEDFP 109
+ ++ I D +GAS++V T TT FP
Sbjct: 407 LGFVETIADALDGASSAVLTSASTTGSFP 321
>AV416480
Length = 410
Score = 27.3 bits (59), Expect = 3.9
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +1
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHII 207
R L AYG ++ S++EY++ N S S +V DH +
Sbjct: 208 RGGLGAYGGKILGSSSEYVQSNISRYFSDPQYYFQVNDHYV 330
>BP045352
Length = 434
Score = 27.3 bits (59), Expect = 3.9
Identities = 8/24 (33%), Positives = 17/24 (70%)
Frame = -1
Query: 277 WAVKSQHENQVSVAEMRMLRWMSR 300
W++KSQ ++++ +LRW+S+
Sbjct: 407 WSIKSQRSKHMTISLNELLRWLSQ 336
>TC8593 homologue to UP|AAR83877 (AAR83877) 60S ribosomal protein L19,
partial (80%)
Length = 874
Score = 26.6 bits (57), Expect = 6.6
Identities = 18/67 (26%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Frame = +1
Query: 29 LTMEAIYLLRRVIERYRTDKK-DLHLVFIDLEKTYDRV--PREILWKALEKKGVRIAYIR 85
L M + +LRR++ +YR KK D H+ K V + +L +++ K A +
Sbjct: 307 LWMRRMRVLRRLLRKYRESKKIDKHMYHDMYMKVKGNVFKNKRVLMESIHKSKAEKAREK 486
Query: 86 AIKDMYE 92
+ D +E
Sbjct: 487 TLSDQFE 507
>TC15259 homologue to UP|AAR83877 (AAR83877) 60S ribosomal protein L19,
partial (80%)
Length = 956
Score = 26.2 bits (56), Expect = 8.6
Identities = 18/67 (26%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Frame = +3
Query: 29 LTMEAIYLLRRVIERYRTDKK-DLHLVFIDLEKTYDRV--PREILWKALEKKGVRIAYIR 85
L M + +LRR++ +YR KK D H+ K V + +L +++ K A +
Sbjct: 306 LWMRRMRVLRRLLRKYRESKKIDKHMYHDMYMKVKGNVFKNKRVLMESIHKTKAEKAREK 485
Query: 86 AIKDMYE 92
+ D +E
Sbjct: 486 TLSDQFE 506
>TC14381 similar to UP|RNS2_ARATH (P42814) Ribonuclease 2 precursor ,
partial (75%)
Length = 858
Score = 26.2 bits (56), Expect = 8.6
Identities = 12/41 (29%), Positives = 18/41 (43%)
Frame = -2
Query: 266 VRPALLYGTECWAVKSQHENQVSVAEMRMLRWMSRKTRHDR 306
VRP +YG CW++ + + A L SR + R
Sbjct: 314 VRPESMYGEYCWSIGPSAASVIGTAMASALAVSSRPLQRQR 192
>BP046905
Length = 538
Score = 26.2 bits (56), Expect = 8.6
Identities = 19/74 (25%), Positives = 33/74 (43%), Gaps = 5/74 (6%)
Frame = -3
Query: 152 VGESREEVNGRLETWRQALEAYGF-----RLSRSTTEYMECNFSGRRSRSTLEVKVGDHI 206
V ++ NG ET + E Y F + EYM SG+ + +L+ I
Sbjct: 530 VFKTESPANGVAETLQIKEERYDFSEFLPEMIEERPEYMSRQLSGKEWKPSLDTIKEKKI 351
Query: 207 IPQVTRFKYLGSFV 220
+++R+ +L SF+
Sbjct: 350 KKKLSRWFFLKSFI 309
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,394,536
Number of Sequences: 28460
Number of extensions: 66154
Number of successful extensions: 318
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 318
length of query: 346
length of database: 4,897,600
effective HSP length: 91
effective length of query: 255
effective length of database: 2,307,740
effective search space: 588473700
effective search space used: 588473700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144503.9


