

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
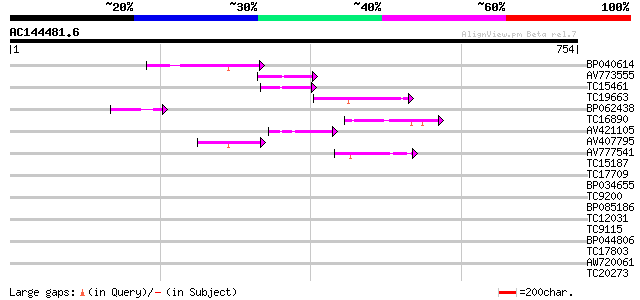
Query= AC144481.6 - phase: 0
(754 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP040614 54 9e-08
AV773555 53 2e-07
TC15461 similar to UP|O24147 (O24147) Kinesin-like protein, part... 50 1e-06
TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation fac... 49 2e-06
BP062438 49 3e-06
TC16890 43 2e-04
AV421105 43 2e-04
AV407795 42 4e-04
AV777541 42 5e-04
TC15187 similar to UP|O24147 (O24147) Kinesin-like protein, part... 39 0.004
TC17709 weakly similar to UP|Q9MA09 (Q9MA09) F20B17.9 (At1g79660... 36 0.020
BP034655 36 0.026
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 35 0.034
BP085186 34 0.075
TC12031 homologue to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-l... 34 0.075
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 34 0.098
BP044806 33 0.13
TC17803 33 0.22
AW720061 33 0.22
TC20273 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partia... 33 0.22
>BP040614
Length = 504
Score = 53.9 bits (128), Expect = 9e-08
Identities = 46/166 (27%), Positives = 79/166 (46%), Gaps = 8/166 (4%)
Frame = +1
Query: 182 SKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKK 241
SK ++ V V ++EIYNE++ DLL +NG ++ + ++ G
Sbjct: 43 SKDRADSIKYEVFVQMIEIYNEQVRDLLVSNGS-----------NRRLDIRNNSQLNGLN 189
Query: 242 AKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVG 293
+A + + ++ +K R V +T N+RSSRSH ++ + V +
Sbjct: 190 VPDAYLVPVTCTQDVLYLMEVGQKNRAVGATALNERSSRSHSVLTVHVRGRELVSGSILR 369
Query: 294 GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIA 339
G L LVD+AGSE +E++ G E + IN+ AL V+ ++A
Sbjct: 370 GCLHLVDLAGSERVEKSEAVG-ERLKEAQHINRSLSALGDVISALA 504
>AV773555
Length = 460
Score = 53.1 bits (126), Expect = 2e-07
Identities = 30/83 (36%), Positives = 49/83 (58%), Gaps = 3/83 (3%)
Frame = -1
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
+L V+ ++A D H+PFR SKLT LLQ + D SK +M + SPDP +++ +L +
Sbjct: 451 SLSDVIFALAQQDDHIPFRHSKLTYLLQPALGGD-SKTVMFVNISPDPASAGESLCSLRF 275
Query: 390 GAKAK-CIVRGP--HTPVKEEDS 409
++ C++ P HT V+ +S
Sbjct: 274 ASRVNACVIGTPRRHTHVRPTES 206
>TC15461 similar to UP|O24147 (O24147) Kinesin-like protein, partial (9%)
Length = 527
Score = 50.4 bits (119), Expect = 1e-06
Identities = 24/75 (32%), Positives = 44/75 (58%)
Frame = +3
Query: 334 VVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKA 393
V+ ++++G H+P+R+ KLTML+ DS +K LM + SP + +T ++L Y ++
Sbjct: 9 VISALSSGGQHIPYRNHKLTMLMSDSL-GGNAKTLMFVNVSPIESSLDETHNSLMYASRV 185
Query: 394 KCIVRGPHTPVKEED 408
+ IV P V ++
Sbjct: 186 RSIVNDPSKNVSSKE 230
>TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation factor,
partial (10%)
Length = 485
Score = 49.3 bits (116), Expect = 2e-06
Identities = 39/138 (28%), Positives = 70/138 (50%), Gaps = 6/138 (4%)
Frame = +3
Query: 405 KEEDSSSTVILGSRIAAMDEFIMKLQM-ENKLREKERNEAHKKLMK---KEEEIAALRTK 460
KE++ + A + +K ++ E+K E + A KKL K + +E A R +
Sbjct: 60 KEKEKKAAAAAAGSAPANETVEVKAEVVESKKNESKTKAADKKLPKHVREMQEALARRKE 239
Query: 461 VETAPASEEEINLKVNERTRHLRQELEKKLEECQR--MTNEFVELERKRMEERILQQQEE 518
E EEE L+ E R ++ELE++ EE +R E +L +K+ E ++L +++
Sbjct: 240 AEERKKREEEEKLRKEEEERRRQEELERQAEEARRRKKEREKEKLLKKKQEGKLLTGKQK 419
Query: 519 VEILRKRLEEIELQLCSS 536
E ++RLE + Q+ S+
Sbjct: 420 EE--QRRLEAMRRQILST 467
>BP062438
Length = 498
Score = 48.9 bits (115), Expect = 3e-06
Identities = 29/79 (36%), Positives = 42/79 (52%), Gaps = 2/79 (2%)
Frame = +3
Query: 134 TIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDI--LGDGDTDSEGSDGDSSKGFCLGVRT 191
T+ YG T SGK+HTM G G++ RA+RD+ + D D E
Sbjct: 234 TVFAYGQTNSGKTHTMRGSKADPGVIPRAVRDLFQIIQQDVDRE---------------F 368
Query: 192 FVQVTVLEIYNEEIYDLLS 210
++++ +EIYNEEI DLL+
Sbjct: 369 LLRMSYMEIYNEEINDLLA 425
>TC16890
Length = 965
Score = 42.7 bits (99), Expect = 2e-04
Identities = 36/141 (25%), Positives = 69/141 (48%), Gaps = 9/141 (6%)
Frame = +2
Query: 446 KLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELER 505
++ K++EE++ R+ + A EEEI K E ++ +L + EE +R+ R
Sbjct: 221 QMSKEDEEMS--RSALSNFKAKEEEIERKKMEVREKVQLQLGRVEEETKRLAT-----IR 379
Query: 506 KRMEERILQQQEEVEILRKRLEEIELQL------CSSSKQERKDENES---KEMEPNGFM 556
+ +E ++EV I+RKR++ + +L C ++E KD E+ K E +
Sbjct: 380 EELESLADPMRKEVAIVRKRIDSVNKELKPLGHTCQKKEKEYKDALEAFNEKNREKVQLI 559
Query: 557 RKLLSVYKSTDDLGMVKSMDL 577
+L+ + ++ L M K +L
Sbjct: 560 TRLMELVSESERLRMKKLEEL 622
>AV421105
Length = 415
Score = 42.7 bits (99), Expect = 2e-04
Identities = 29/91 (31%), Positives = 48/91 (51%)
Frame = +2
Query: 345 VPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPV 404
VP+R+SKLT +LQDS S+ LM+ C +P E +++ T+ A+++ I +
Sbjct: 5 VPYRESKLTRILQDSL-GGTSRALMVACLNPG--EYQESVHTVSLAARSRHITNVVPSAH 175
Query: 405 KEEDSSSTVILGSRIAAMDEFIMKLQMENKL 435
KEE V + +++ A E K + KL
Sbjct: 176 KEETPKVMVDMEAKLRAWLESKGKAKSSQKL 268
>AV407795
Length = 424
Score = 42.0 bits (97), Expect = 4e-04
Identities = 26/101 (25%), Positives = 49/101 (47%), Gaps = 11/101 (10%)
Frame = +2
Query: 251 NEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV-----------PTVGGRLMLV 299
++ I + I++ R +T N+ SSRSH ++ L + P G+L +
Sbjct: 119 SDVESIKELIERGNATRSTGTTGANEESSRSHAILQLAIKRSVDGNESKPPRPVGKLSFI 298
Query: 300 DMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIAN 340
D+AGSE + +++ A+IN+ +ALK + ++ N
Sbjct: 299 DLAGSERGADTTDNDKQTRIEGAEINKSLLALKECIRALDN 421
>AV777541
Length = 466
Score = 41.6 bits (96), Expect = 5e-04
Identities = 33/113 (29%), Positives = 53/113 (46%), Gaps = 3/113 (2%)
Frame = +1
Query: 433 NKLREKERNEAHKKLMKKE---EEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKK 489
++LR E+ + K K++ EE R K+ A + ++ + +R+E E+K
Sbjct: 34 SELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERK 213
Query: 490 LEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERK 542
R +L RK EER QQEE E+L ++L E + S+Q RK
Sbjct: 214 ASSAAREARAIEQLRRK--EERAKAQQEEAELLAQKLAE----RLNESEQRRK 354
>TC15187 similar to UP|O24147 (O24147) Kinesin-like protein, partial (8%)
Length = 611
Score = 38.5 bits (88), Expect = 0.004
Identities = 19/64 (29%), Positives = 36/64 (55%)
Frame = +2
Query: 345 VPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPV 404
+P+R+ KLTML+ DS ++ LM + SP + +T ++L Y ++ + I+ P V
Sbjct: 2 IPYRNHKLTMLMSDSL-GGNAQTLMFVNVSPVESSLDETHNSLMYASRVRSIINDPSQNV 178
Query: 405 KEED 408
++
Sbjct: 179 ASKE 190
>TC17709 weakly similar to UP|Q9MA09 (Q9MA09) F20B17.9 (At1g79660/F20B17_9),
partial (39%)
Length = 576
Score = 36.2 bits (82), Expect = 0.020
Identities = 23/54 (42%), Positives = 27/54 (49%), Gaps = 6/54 (11%)
Frame = -1
Query: 178 DGDSSKGFCLGVRTFVQVTVLEIYNEEIY---DLL---STNGGGGGGGGFGFGW 225
DGD ++G V + T E Y DL+ NGGGGGGGG GFGW
Sbjct: 198 DGDLNRGEDRMVSRTMASTTTFFTPPEAYAACDLVPGREANGGGGGGGGGGFGW 37
>BP034655
Length = 517
Score = 35.8 bits (81), Expect = 0.026
Identities = 21/46 (45%), Positives = 30/46 (64%)
Frame = +1
Query: 615 AYVCAPNFGQKACLSTVYEEEGEEEAEQDHDKVEEDEEVEKEVIEE 660
A++ A +FG LS EEE EEE E++ ++ EE+EE E+E EE
Sbjct: 103 AHLTAADFG----LSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEE 228
Score = 31.2 bits (69), Expect = 0.64
Identities = 14/28 (50%), Positives = 21/28 (75%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEE 660
EEE EEE E++ ++ EEDEE ++E +E
Sbjct: 169 EEEEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 31.2 bits (69), Expect = 0.64
Identities = 14/29 (48%), Positives = 21/29 (72%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEEK 661
EEE EEE E++ ++ EE+EE E E +E+
Sbjct: 154 EEEEEEEEEEEEEEEEEEEEEEDEEDDEE 240
Score = 30.0 bits (66), Expect = 1.4
Identities = 13/29 (44%), Positives = 22/29 (75%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEEK 661
EEE EEE E++ ++ EE+EE E++ E++
Sbjct: 160 EEEEEEEEEEEEEEEEEEEEDEEDDEEDE 246
Score = 29.3 bits (64), Expect = 2.4
Identities = 19/43 (44%), Positives = 26/43 (60%), Gaps = 4/43 (9%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKE----VIEEKRVCSVVNKSP 671
EEE EEE E++ D+ E+DEE E E V RV V+++P
Sbjct: 184 EEEEEEEEEEEEDE-EDDEEDEDEDYIPVAPSHRVSPNVHQNP 309
Score = 28.9 bits (63), Expect = 3.2
Identities = 12/29 (41%), Positives = 21/29 (72%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEEK 661
EEE EEE E++ ++ EE++E + E E++
Sbjct: 166 EEEEEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 27.7 bits (60), Expect = 7.1
Identities = 11/28 (39%), Positives = 21/28 (74%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEE 660
EEE EEE E++ ++ E++E+ E++ E+
Sbjct: 172 EEEEEEEEEEEEEEEEDEEDDEEDEDED 255
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 35.4 bits (80), Expect = 0.034
Identities = 16/28 (57%), Positives = 21/28 (74%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEE 660
EEE EEE E++ ++ EE+EEVE E EE
Sbjct: 247 EEEEEEEGEEEEEEEEEEEEVEAEAEEE 330
Score = 33.1 bits (74), Expect = 0.17
Identities = 17/43 (39%), Positives = 28/43 (64%)
Frame = +1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEEKRVCSVVNKSPKIED 675
EEE EEE E++ ++ EE+EE E+E EE+ V ++ + +D
Sbjct: 208 EEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDD 336
Score = 32.7 bits (73), Expect = 0.22
Identities = 16/30 (53%), Positives = 22/30 (73%)
Frame = +1
Query: 631 VYEEEGEEEAEQDHDKVEEDEEVEKEVIEE 660
V EEE EEE E++ ++ EE+EE E+EV E
Sbjct: 229 VEEEEEEEEEEEEGEEEEEEEEEEEEVEAE 318
>BP085186
Length = 388
Score = 34.3 bits (77), Expect = 0.075
Identities = 25/73 (34%), Positives = 36/73 (49%)
Frame = -1
Query: 633 EEEGEEEAEQDHDKVEEDEEVEKEVIEEKRVCSVVNKSPKIEDYTGADKENNGSNRLLRI 692
EEE EEE E++ ++ EE+EE E E EEK VN +E G N L++
Sbjct: 259 EEEEEEEEEEEEEEEEEEEEAEPE--EEKNEEENVN-----------TQEGEGENP*LQV 119
Query: 693 HNIFTLCGNQREL 705
+F L ++L
Sbjct: 118 LLLFLLANYVKDL 80
>TC12031 homologue to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein
A), partial (7%)
Length = 740
Score = 34.3 bits (77), Expect = 0.075
Identities = 24/75 (32%), Positives = 41/75 (54%), Gaps = 1/75 (1%)
Frame = +2
Query: 348 RDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAK-CIVRGPHTPVKE 406
R+SKLT LLQ D SK LM + SPD + +++ +L + ++ C + P +
Sbjct: 2 RNSKLTYLLQPCLGGD-SKTLMFVNISPDQSSVGESLCSLRFASRVNACEIGIP----RR 166
Query: 407 EDSSSTVILGSRIAA 421
+ +ST L SR+++
Sbjct: 167 QTQTSTRSLDSRLSS 211
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 33.9 bits (76), Expect = 0.098
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = +1
Query: 5 PSSSSKQIHTTHLITPRSKHRLNFNGVKPAPTP-PHPHPSPHPNFNNKDSPPEHP 58
PSS S ++ HL+ + + + KP P+P H P PHP N S P P
Sbjct: 184 PSSPSSNPNSHHLLPHNNNPTRSKSQPKPPPSPTSHSPPPPHPRRPNPPSNPPSP 348
>BP044806
Length = 538
Score = 33.5 bits (75), Expect = 0.13
Identities = 19/73 (26%), Positives = 32/73 (43%)
Frame = +3
Query: 3 PTPSSSSKQIHTTHLITPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPIEVI 62
P P+ + K++ I K + +K + P P+PS N ++ +P H I V
Sbjct: 303 PNPTPNRKRLCQCRSIKSTQKRKRQCRSIKHQTSIPPPNPSDPSNTKHQTNPQIHQIRVT 482
Query: 63 ARIRDYPDRKDKP 75
+ R P K +P
Sbjct: 483 TQCRS-PPMKTRP 518
>TC17803
Length = 485
Score = 32.7 bits (73), Expect = 0.22
Identities = 18/47 (38%), Positives = 23/47 (48%)
Frame = -3
Query: 19 TPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHPIEVIARI 65
TPR FNG+ PP PHP+P P K +P P +V+ I
Sbjct: 123 TPRPN---KFNGI-----PPKPHPTPKPQPTPKPNPTPAPSKVVMTI 7
Score = 31.2 bits (69), Expect = 0.64
Identities = 14/27 (51%), Positives = 16/27 (58%), Gaps = 1/27 (3%)
Frame = -3
Query: 33 PAPTP-PHPHPSPHPNFNNKDSPPEHP 58
P+PTP P P P+P PN N P HP
Sbjct: 156 PSPTPTPQPPPTPRPNKFNGIPPKPHP 76
>AW720061
Length = 304
Score = 32.7 bits (73), Expect = 0.22
Identities = 16/48 (33%), Positives = 25/48 (51%), Gaps = 8/48 (16%)
Frame = -1
Query: 7 SSSKQIHTT-----HLITPRSKHRLNFNGVKPAPTPPHPH---PSPHP 46
+S + +HTT H + PR+ + P+PTPP P P+P+P
Sbjct: 259 TSFESLHTTCV*TPHYVPPRTSYSRGCRNPSPSPTPPPPRAAPPTPNP 116
>TC20273 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (64%)
Length = 569
Score = 32.7 bits (73), Expect = 0.22
Identities = 40/182 (21%), Positives = 68/182 (36%), Gaps = 4/182 (2%)
Frame = +1
Query: 350 SKLTML--LQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEE 407
SKL + LQ E D + + P + K + +V TPV E
Sbjct: 43 SKLASMAELQKKAESDTTPAPVAAVPPPSNDAVAKKAPEVAADDSKALVVVPEKTPVPEN 222
Query: 408 DSSSTVILGSRIAAMD-EFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPA 466
SS L +A + E +L E E+++ K K ++ A + A
Sbjct: 223 KPSSKGSLDRDVALAELEKEKRLSYVKAWEESEKSKTENKAQKNLSDVVAWENSKKAA-- 396
Query: 467 SEEEINLKVNERTRHLRQELEKKLEEC-QRMTNEFVELERKRMEERILQQQEEVEILRKR 525
+ + R + + LEKK E ++M N+ + ++ E R + + + E L K
Sbjct: 397 --------LEAQLRKIEERLEKKKAEYGEKMKNKIALVHKEAEERRAMIEAKRGEDLLKA 552
Query: 526 LE 527
E
Sbjct: 553 EE 558
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,348,161
Number of Sequences: 28460
Number of extensions: 184379
Number of successful extensions: 3962
Number of sequences better than 10.0: 199
Number of HSP's better than 10.0 without gapping: 2384
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3262
length of query: 754
length of database: 4,897,600
effective HSP length: 97
effective length of query: 657
effective length of database: 2,136,980
effective search space: 1403995860
effective search space used: 1403995860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144481.6


