

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
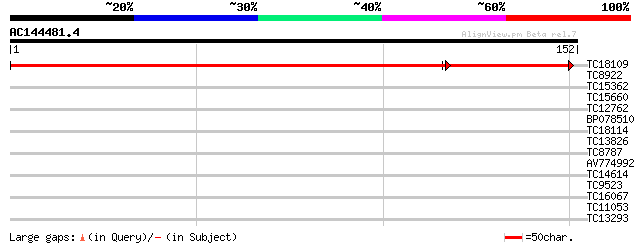
Query= AC144481.4 + phase: 0
(152 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18109 202 2e-62
TC8922 28 0.90
TC15362 similar to UP|Q8DLK1 (Q8DLK1) ABC transporter subunit, p... 28 0.90
TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, part... 27 1.5
TC12762 26 2.6
BP078510 26 3.4
TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D... 26 3.4
TC13826 26 3.4
TC8787 similar to UP|Q90YX0 (Q90YX0) Ribosomal protein P1, parti... 25 4.5
AV774992 25 7.6
TC14614 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammoni... 25 7.6
TC9523 similar to UP|O48721 (O48721) Expressed protein, partial ... 24 10.0
TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete 24 10.0
TC11053 weakly similar to UP|Q9FHK7 (Q9FHK7) Receptor-like prote... 24 10.0
TC13293 24 10.0
>TC18109
Length = 948
Score = 202 bits (513), Expect(2) = 2e-62
Identities = 97/118 (82%), Positives = 108/118 (91%)
Frame = +3
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+VK+DIGGNISRLESKY SN +FN LYSLVQVEVETKTAK+SSSCT GLLWLTRAM
Sbjct: 186 MALVKADIGGNISRLESKYTSNSTRFNYLYSLVQVEVETKTAKSSSSCTNGLLWLTRAMH 365
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+F+NL++H DWSMSQACTD+Y KTLKKWHGWLASS+ TVAM LAPDRKKFMEV+
Sbjct: 366 FLVALFQNLLDHEDWSMSQACTDAYTKTLKKWHGWLASSSFTVAMKLAPDRKKFMEVL 539
Score = 52.4 bits (124), Expect(2) = 2e-62
Identities = 28/35 (80%), Positives = 28/35 (80%)
Frame = +1
Query: 117 VVISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*R 151
VVIS LT SNFV FLLSLKRITSFWL L WM *R
Sbjct: 547 VVISKLTFSNFVPPFLLSLKRITSFWLVLAWMI*R 651
>TC8922
Length = 638
Score = 27.7 bits (60), Expect = 0.90
Identities = 25/92 (27%), Positives = 36/92 (38%), Gaps = 7/92 (7%)
Frame = +2
Query: 49 TTGLLWLT-------RAMDFLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTV 101
T LLWL R + L RN D+ + T + K L++ G + STV
Sbjct: 116 TRHLLWLGSTFSSPGRKIQALYLHKRNCQMVLDFLAKECMTHQFQKMLQRVRG*MLISTV 295
Query: 102 TVAMMLAPDRKKFMEVVISMLTLSNFVLAFLL 133
+ + R FM +M+ S FLL
Sbjct: 296 PTLLWTSQSRTSFMWTWRTMMKFSPASFRFLL 391
>TC15362 similar to UP|Q8DLK1 (Q8DLK1) ABC transporter subunit, partial
(41%)
Length = 848
Score = 27.7 bits (60), Expect = 0.90
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +2
Query: 13 SRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLL 53
SR+ SK +S NC LVQV+ + + A+ S C + L+
Sbjct: 221 SRIISKGISVGNSRNCYRGLVQVQPKAENAQNKSQCDSMLI 343
>TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, partial (34%)
Length = 618
Score = 26.9 bits (58), Expect = 1.5
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Frame = +3
Query: 42 AKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACTD-SYYKTLKKWHGWLA 97
A++ SS +T L W F+ +FR W +++C+D S+ + KW LA
Sbjct: 120 AQSCSSSSTCLCWY*CRRFFVPTLFRRSFLWLLWEGTKSCSDGSWVRKRSKWSKVLA 290
>TC12762
Length = 865
Score = 26.2 bits (56), Expect = 2.6
Identities = 11/24 (45%), Positives = 18/24 (74%)
Frame = -1
Query: 117 VVISMLTLSNFVLAFLLSLKRITS 140
+ IS++ S + L FLLSLK+++S
Sbjct: 787 IKISVINFSKYTL*FLLSLKKVSS 716
>BP078510
Length = 391
Score = 25.8 bits (55), Expect = 3.4
Identities = 14/49 (28%), Positives = 19/49 (38%)
Frame = +3
Query: 91 KWHGWLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRIT 139
KWH WL S T+ + L N +L FL+ K+ T
Sbjct: 15 KWHSWLKSQTINFKIKL------------------NVILQFLIHFKQTT 107
>TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;}, partial (4%)
Length = 576
Score = 25.8 bits (55), Expect = 3.4
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +1
Query: 130 AFLLSLKRITSFWLDLDWMN 149
+ LL+LK T W+D W+N
Sbjct: 478 SILLTLKTCTGLWIDRFWLN 537
>TC13826
Length = 533
Score = 25.8 bits (55), Expect = 3.4
Identities = 10/31 (32%), Positives = 19/31 (61%), Gaps = 1/31 (3%)
Frame = +2
Query: 64 AVFRNLIE-HADWSMSQACTDSYYKTLKKWH 93
+ +RN+ WS+S++C D ++K +K H
Sbjct: 2 SAYRNIFPIWVPWSISKSCIDLWWKRIKSKH 94
>TC8787 similar to UP|Q90YX0 (Q90YX0) Ribosomal protein P1, partial (15%)
Length = 925
Score = 25.4 bits (54), Expect = 4.5
Identities = 10/34 (29%), Positives = 17/34 (49%), Gaps = 2/34 (5%)
Frame = -2
Query: 64 AVFRNLIEHADWSMSQACTDSYY--KTLKKWHGW 95
+++RN + + DW + SYY + W GW
Sbjct: 243 SIYRNSLPNIDWRSNSYRRSSYYLMHCRRWWKGW 142
>AV774992
Length = 491
Score = 24.6 bits (52), Expect = 7.6
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +1
Query: 89 LKKWHGWLASSTVTVAMMLAPDRKK 113
LK W GWL S VA + A D K
Sbjct: 298 LKNWVGWL*FSMAAVADISAADANK 372
>TC14614 homologue to UP|PAL1_SOYBN (P27991) Phenylalanine ammonia-lyase 1
, partial (26%)
Length = 550
Score = 24.6 bits (52), Expect = 7.6
Identities = 15/36 (41%), Positives = 20/36 (54%)
Frame = +3
Query: 13 SRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSC 48
S+L SK LSN K L+ +E TK ++ SSC
Sbjct: 342 SQLMSKVLSNTTKT*TLWV*FLLEKPTKPLRSLSSC 449
>TC9523 similar to UP|O48721 (O48721) Expressed protein, partial (32%)
Length = 634
Score = 24.3 bits (51), Expect = 10.0
Identities = 22/84 (26%), Positives = 33/84 (39%), Gaps = 15/84 (17%)
Frame = +1
Query: 30 YSLVQVE-VETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQAC------- 81
YS V V+ V+ K + S +T A + ++NLI ++WS AC
Sbjct: 64 YSTVPVQHVDLMVLKEALSAPV----VTVASPSAIRAWKNLISDSEWSNYVACIGETTAA 231
Query: 82 -------TDSYYKTLKKWHGWLAS 98
+ YY T GW+ S
Sbjct: 232 KARGLGFRNVYYPTQPGLEGWVES 303
>TC16067 UP|Q9ZRY4 (Q9ZRY4) NDX1 homeobox protein, complete
Length = 3271
Score = 24.3 bits (51), Expect = 10.0
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = +3
Query: 131 FLLSLKRITSFWLDLDWMN*RL 152
FL+S K + + L+L W+N*RL
Sbjct: 2580 FLVSQK*LLAITLNLHWLN*RL 2645
>TC11053 weakly similar to UP|Q9FHK7 (Q9FHK7) Receptor-like protein kinase,
partial (9%)
Length = 606
Score = 24.3 bits (51), Expect = 10.0
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 6/58 (10%)
Frame = +3
Query: 12 ISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGL------LWLTRAMDFLV 63
I++ + SNP +++CL L + T A +SSSC L WL + LV
Sbjct: 216 IAKFAFAFSSNP*RWSCLAVLAYI*Q*TVFACSSSSCIVLLQIIY*CFWLLILLSVLV 389
>TC13293
Length = 556
Score = 24.3 bits (51), Expect = 10.0
Identities = 16/66 (24%), Positives = 31/66 (46%), Gaps = 10/66 (15%)
Frame = +1
Query: 26 FNCLYSLVQVEVETKTAKASSSCTTGLL----------WLTRAMDFLVAVFRNLIEHADW 75
++C Y VQ ++ +K + T LL L+ + + + +FR +++HA W
Sbjct: 211 WSCCYPQVQA*DPSRMSKLGTRSTLHLLN*GLYFVQQVELSFCVVYWM*IFRFILDHAFW 390
Query: 76 SMSQAC 81
+S C
Sbjct: 391 RLSCNC 408
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,886,627
Number of Sequences: 28460
Number of extensions: 35621
Number of successful extensions: 257
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 257
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 257
length of query: 152
length of database: 4,897,600
effective HSP length: 82
effective length of query: 70
effective length of database: 2,563,880
effective search space: 179471600
effective search space used: 179471600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144481.4


