

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
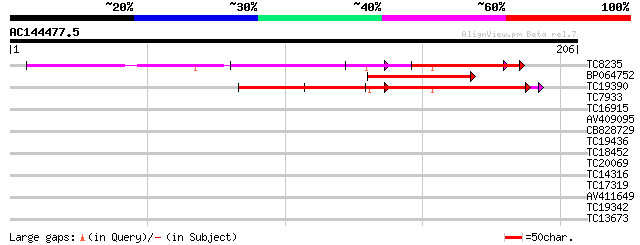
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8235 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (27%) 78 1e-15
BP064752 67 3e-12
TC19390 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (17%) 54 3e-08
TC7933 weakly similar to UP|AAQ56728 (AAQ56728) Polygalacturonas... 38 0.001
TC16915 weakly similar to UP|Q9SDA8 (Q9SDA8) AT2G17020 protein (... 35 0.009
AV409095 34 0.016
CB828729 31 0.13
TC19436 similar to UP|Q93Y06 (Q93Y06) Receptor protein kinase-li... 31 0.17
TC18452 weakly similar to GB|AAO64875.1|29028992|BT005940 At5g27... 30 0.39
TC20069 similar to UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kina... 29 0.66
TC14316 weakly similar to GB|AAL47412.1|17978881|AY066045 At1g75... 27 2.5
TC17319 weakly similar to UP|Q8H5I9 (Q8H5I9) OJ1119_A04.29 prote... 26 4.3
AV411649 25 7.3
TC19342 25 9.6
TC13673 similar to UP|Q9AXK0 (Q9AXK0) Cellulose synthase CesA-1,... 25 9.6
>TC8235 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (27%)
Length = 847
Score = 77.8 bits (190), Expect = 1e-15
Identities = 39/41 (95%), Positives = 40/41 (97%)
Frame = +2
Query: 147 LNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
LNLDARQITDAGLANLTSLSGLITLDLFGARI+DSGT YLR
Sbjct: 2 LNLDARQITDAGLANLTSLSGLITLDLFGARISDSGTAYLR 124
Score = 59.7 bits (143), Expect = 4e-10
Identities = 34/108 (31%), Positives = 59/108 (54%), Gaps = 1/108 (0%)
Frame = +2
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
+I D GL NLT L+ L +L L + +SG Y+ KL+ L + ++D G+K +
Sbjct: 20 QITDAGLANLTSLSGLITLDLFGARISDSGTAYLRSFKKLQSLEICGGGISDAGVKNIRE 199
Query: 141 LTNLKSLNLDAR-QITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
+ +L LNL +TD L ++ ++ L +L++ RIT+ G +L+
Sbjct: 200 IVSLTQLNLSQNGNLTDRTLELISGMTALRSLNVSNCRITNEGLRHLK 343
Score = 53.5 bits (127), Expect = 3e-08
Identities = 34/102 (33%), Positives = 56/102 (54%), Gaps = 1/102 (0%)
Frame = +2
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLL 139
+I D G L L+SL + + ++G++ I + L LNLS ++TD L+ +
Sbjct: 92 RISDSGTAYLRSFKKLQSLEICGGGISDAGVKNIREIVSLTQLNLSQNGNLTDRTLELIS 271
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDS 181
G+T L+SLN+ +IT+ GL +L L L +L L ++T S
Sbjct: 272 GMTALRSLNVSNCRITNEGLRHLKPLKNLRSLTLESCKVTAS 397
Score = 45.4 bits (106), Expect = 7e-06
Identities = 25/65 (38%), Positives = 38/65 (58%)
Frame = +2
Query: 123 LNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
LNL +TD GL L L+ L +L+L +I+D+G A L S L +L++ G I+D+G
Sbjct: 2 LNLDARQITDAGLANLTSLSGLITLDLFGARISDSGTAYLRSFKKLQSLEICGGGISDAG 181
Query: 183 TTYLR 187
+R
Sbjct: 182 VKNIR 196
Score = 42.4 bits (98), Expect = 6e-05
Identities = 39/143 (27%), Positives = 66/143 (45%), Gaps = 11/143 (7%)
Frame = +2
Query: 7 LNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQA*RG*V 66
LN++ IT A +++L+ L L+L +SD G + F LQ L+ G +
Sbjct: 2 LNLDARQITDAGLANLTSLSGLITLDLFGARISDSG----TAYLRSFKKLQSLEICGGGI 169
Query: 67 -----------LHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYIS 115
+ LT+L + D L ++G+T L+SL +S+ + N G+R++
Sbjct: 170 SDAGVKNIREIVSLTQLNLSQNG--NLTDRTLELISGMTALRSLNVSNCRITNEGLRHLK 343
Query: 116 GLNKLEDLNLSFTSVTDNGLKRL 138
L L L L VT + +K+L
Sbjct: 344 PLKNLRSLTLESCKVTASEIKKL 412
Score = 38.1 bits (87), Expect = 0.001
Identities = 28/100 (28%), Positives = 50/100 (50%), Gaps = 7/100 (7%)
Frame = +2
Query: 99 LVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAG 158
L L ++ ++G+ ++ L+ L L+L ++D+G L L+SL + I+DAG
Sbjct: 2 LNLDARQITDAGLANLTSLSGLITLDLFGARISDSGTAYLRSFKKLQSLEICGGGISDAG 181
Query: 159 LANLTSLSGLITLDLF-GARITD------SGTTYLRCMYI 191
+ N+ + L L+L +TD SG T LR + +
Sbjct: 182 VKNIREIVSLTQLNLSQNGNLTDRTLELISGMTALRSLNV 301
>BP064752
Length = 493
Score = 66.6 bits (161), Expect = 3e-12
Identities = 34/39 (87%), Positives = 35/39 (89%)
Frame = +3
Query: 131 TDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLI 169
TDNGLKRL GLTNLKSLNLDARQITDAGLANLTS L+
Sbjct: 3 TDNGLKRLAGLTNLKSLNLDARQITDAGLANLTSKHFLV 119
Score = 27.7 bits (60), Expect = 1.5
Identities = 18/46 (39%), Positives = 25/46 (54%)
Frame = +3
Query: 84 DEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS 129
D GL L GLT LKSL L ++ ++G ++ L L S+TS
Sbjct: 6 DNGLKRLAGLTNLKSLNLDARQITDAG---LANLTSKHFLVYSYTS 134
>TC19390 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (17%)
Length = 579
Score = 53.5 bits (127), Expect = 3e-08
Identities = 34/88 (38%), Positives = 52/88 (58%), Gaps = 1/88 (1%)
Frame = +2
Query: 108 NSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLS 166
+SG++ I L+ L LNLS S +TD L+ + GLT L SLN+ +IT+AGL +L +L
Sbjct: 11 DSGVKNIKDLSSLTCLNLSQNSNLTDKTLELISGLTGLISLNVSNSRITNAGLRHLKTLK 190
Query: 167 GLITLDLFGARITDSGTTYLRCMYIYQL 194
L +L L ++T + L+ Y+ L
Sbjct: 191 NLRSLTLESCKVTANDIKKLKSTYLPNL 274
Score = 43.1 bits (100), Expect = 3e-05
Identities = 22/61 (36%), Positives = 41/61 (67%), Gaps = 1/61 (1%)
Frame = +2
Query: 130 VTDNGLKRLLGLTNLKSLNLDAR-QITDAGLANLTSLSGLITLDLFGARITDSGTTYLRC 188
+TD+G+K + L++L LNL +TD L ++ L+GLI+L++ +RIT++G +L+
Sbjct: 5 LTDSGVKNIKDLSSLTCLNLSQNSNLTDKTLELISGLTGLISLNVSNSRITNAGLRHLKT 184
Query: 189 M 189
+
Sbjct: 185 L 187
Score = 40.0 bits (92), Expect = 3e-04
Identities = 21/55 (38%), Positives = 34/55 (61%)
Frame = +2
Query: 84 DEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRL 138
D+ L ++GLT L SL +S++ + N+G+R++ L L L L VT N +K+L
Sbjct: 86 DKTLELISGLTGLISLNVSNSRITNAGLRHLKTLKNLRSLTLESCKVTANDIKKL 250
>TC7933 weakly similar to UP|AAQ56728 (AAQ56728) Polygalacturonase
inhibiting protein, partial (78%)
Length = 1320
Score = 37.7 bits (86), Expect = 0.001
Identities = 24/63 (38%), Positives = 36/63 (57%)
Frame = +3
Query: 93 LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDAR 152
LT LKSL ++ T V +++ L L+ L LSF +++ + L L NL SLNLD
Sbjct: 363 LTKLKSLSITWTNVSGPIPDFLAQLKNLDSLYLSFNNLSGHIPASLSRLPNLLSLNLDRN 542
Query: 153 QIT 155
++T
Sbjct: 543 KLT 551
>TC16915 weakly similar to UP|Q9SDA8 (Q9SDA8) AT2G17020 protein
(At2g17020/At2g17020), partial (29%)
Length = 1011
Score = 35.0 bits (79), Expect = 0.009
Identities = 29/74 (39%), Positives = 45/74 (60%), Gaps = 4/74 (5%)
Frame = +3
Query: 111 IRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSG 167
+R +S + +L+ L+L + S+ D+ L+ + LT LK L LD ITDAGL+ L + ++
Sbjct: 30 LRLVSNV-ELKILDLRYCRSLGDDALQAIGTLTKLKVLLLDGSDITDAGLSCLRPSVINS 206
Query: 168 LITLDLFGA-RITD 180
L L L G R+TD
Sbjct: 207 LYALSLRGCKRLTD 248
>AV409095
Length = 414
Score = 34.3 bits (77), Expect = 0.016
Identities = 31/89 (34%), Positives = 43/89 (47%), Gaps = 6/89 (6%)
Frame = +2
Query: 111 IRY----ISGLNKLEDLNLSFTSVTD--NGLKRLLGLTNLKSLNLDARQITDAGLANLTS 164
IRY I L LE L+LSF + + L+GL ++K N ++ A +TS
Sbjct: 2 IRYLPPEIGCLKSLEYLDLSFNKIKTLPTEITYLIGLISMKVANNKLVELPSA----MTS 169
Query: 165 LSGLITLDLFGARITDSGTTYLRCMYIYQ 193
LS L LDL R+T G+ L M+ Q
Sbjct: 170 LSRLECLDLSNNRLTSLGSLELASMHRLQ 256
>CB828729
Length = 478
Score = 31.2 bits (69), Expect = 0.13
Identities = 27/84 (32%), Positives = 41/84 (48%)
Frame = +2
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L L+ L +L LS+ +GN+ I Y N L+ LNL+ + + +T+L LNL
Sbjct: 47 LQSLSSLTNLDLSNNNLGNT-IPYSLPQN-LKHLNLANNNFNGGIPYSISDMTSLVDLNL 220
Query: 150 DARQITDAGLANLTSLSGLITLDL 173
Q+ + LS L TLD+
Sbjct: 221 GHNQLQQGLTVDFEKLSSLSTLDV 292
>TC19436 similar to UP|Q93Y06 (Q93Y06) Receptor protein kinase-like protein,
partial (19%)
Length = 570
Score = 30.8 bits (68), Expect = 0.17
Identities = 23/60 (38%), Positives = 33/60 (54%)
Frame = +1
Query: 114 ISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
++ L++L L L S+T + L LTNLKSL LD + A ++ SL L+TL L
Sbjct: 307 LTRLDQLRVLTLRNNSLT-GPIPDLSPLTNLKSLFLDRNHFSGAFPPSILSLHRLLTLSL 483
>TC18452 weakly similar to GB|AAO64875.1|29028992|BT005940 At5g27920
{Arabidopsis thaliana;}, partial (11%)
Length = 698
Score = 29.6 bits (65), Expect = 0.39
Identities = 20/57 (35%), Positives = 34/57 (59%), Gaps = 2/57 (3%)
Frame = +2
Query: 106 VGNSGIRYISGLNK-LEDLNLSFTSVTDNGLKRLLGLTNLKSLN-LDARQITDAGLA 160
+ +SG+ ++ +K L +NLS++SVTD GL L ++ L+S L + + GLA
Sbjct: 488 IDDSGMIPLAYFSKSLRQINLSYSSVTDVGLLSLASISCLQSFTILHLQGLAPGGLA 658
Score = 29.3 bits (64), Expect = 0.51
Identities = 29/83 (34%), Positives = 43/83 (50%), Gaps = 4/83 (4%)
Frame = +2
Query: 104 TEVGNSGIRYISGLNKLEDLNLSF-TSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLAN 161
T++G S I G LE +N S+ T++TD L L NLK+L + +T GLA+
Sbjct: 260 TDLGISAIA--CGCPGLEIINTSYCTNITDGSLISLSKCRNLKTLEIRGCLLVTSIGLAS 433
Query: 162 LT-SLSGLITLDLFGA-RITDSG 182
+ + LI +D+ I DSG
Sbjct: 434 IAMNCKQLICVDIKKCYNIDDSG 502
>TC20069 similar to UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3,
partial (14%)
Length = 517
Score = 28.9 bits (63), Expect = 0.66
Identities = 27/97 (27%), Positives = 43/97 (43%), Gaps = 5/97 (5%)
Frame = +3
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLK-----RLLGLT 142
+NLTGL+L +L ++ + ++ L+ L S+ DNGL L +T
Sbjct: 261 LNLTGLSLTGTL--------SADVAHLPFLSNL--------SLADNGLSGPIPPSLSAVT 392
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
L+ LNL + L+ L L LDL+ +T
Sbjct: 393 GLRFLNLSNNGFNGTFPSELSVLKNLEVLDLYNNNLT 503
>TC14316 weakly similar to GB|AAL47412.1|17978881|AY066045
At1g75380/F1B16_15 {Arabidopsis thaliana;}, partial
(22%)
Length = 578
Score = 26.9 bits (58), Expect = 2.5
Identities = 15/27 (55%), Positives = 19/27 (69%), Gaps = 1/27 (3%)
Frame = +3
Query: 133 NGLKRLL-GLTNLKSLNLDARQITDAG 158
N +K LL G TNL +L L+ R+ITD G
Sbjct: 162 NAIKMLLFGGTNLINLGLERRRITDPG 242
>TC17319 weakly similar to UP|Q8H5I9 (Q8H5I9) OJ1119_A04.29 protein, partial
(74%)
Length = 920
Score = 26.2 bits (56), Expect = 4.3
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = +1
Query: 14 ITAACFEYISALAALACLNLN 34
+T CF I++LAA+ C+ +N
Sbjct: 139 VTCKCFSVITSLAAILCIAVN 201
>AV411649
Length = 358
Score = 25.4 bits (54), Expect = 7.3
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 1/48 (2%)
Frame = +2
Query: 143 NLKSLNLDARQITDAGLANLTS-LSGLITLDLFGARITDSGTTYLRCM 189
NLKSLNL ++ AGL L + L L L + D+ ++ M
Sbjct: 50 NLKSLNLSCTAVSSAGLGVLAGHVPNLEILSLSQTPVDDTAIPFIGMM 193
>TC19342
Length = 548
Score = 25.0 bits (53), Expect = 9.6
Identities = 12/34 (35%), Positives = 16/34 (46%)
Frame = -1
Query: 14 ITAACFEYISALAALACLNLNRCGLSDDGFEKFS 47
+ CF Y+SAL + L L CG G + S
Sbjct: 452 VPLGCFFYVSALVIGSILRLYYCGFKIAGAKSLS 351
>TC13673 similar to UP|Q9AXK0 (Q9AXK0) Cellulose synthase CesA-1, partial
(10%)
Length = 574
Score = 25.0 bits (53), Expect = 9.6
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 38 LSDDGFEKFSVLQHMFPFLQVL 59
L+ GF F V+ H++PFLQ L
Sbjct: 125 LTGKGFFAFWVIFHLYPFLQGL 190
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,070,759
Number of Sequences: 28460
Number of extensions: 34730
Number of successful extensions: 272
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 270
length of query: 206
length of database: 4,897,600
effective HSP length: 86
effective length of query: 120
effective length of database: 2,450,040
effective search space: 294004800
effective search space used: 294004800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144477.5


