

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144431.6 - phase: 0
(377 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
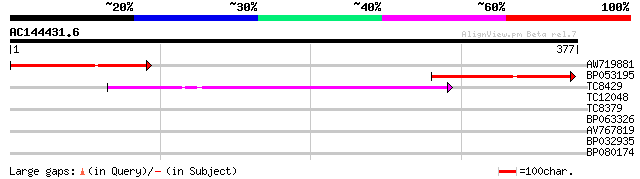
Score E
Sequences producing significant alignments: (bits) Value
AW719881 92 1e-19
BP053195 73 7e-14
TC8429 weakly similar to UP|BAD03306 (BAD03306) MAP3K-like prote... 46 1e-05
TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749),... 33 0.078
TC8379 similar to UP|Q9SNJ3 (Q9SNJ3) ESTs AU031435(E61570), part... 31 0.39
BP063326 30 0.66
AV767819 28 3.3
BP032935 26 9.5
BP080174 26 9.5
>AW719881
Length = 495
Score = 92.4 bits (228), Expect = 1e-19
Identities = 59/95 (62%), Positives = 64/95 (67%), Gaps = 1/95 (1%)
Frame = +2
Query: 1 MPVLHVKHVLQMEIHDPDVAGYYLQIPLTK*RVYRLIGIHG*RHIIHLLWLVQDQPQVQL 60
M VL V+H LQM I DPDV G+ LQ PL K*R + L GIHG R LL +D+P L
Sbjct: 212 MVVLPVRHALQMGIPDPDVHGFNLQTPLPK*RGFHLTGIHGSRLTTRLLRPERDRPP-GL 388
Query: 61 SLHL-*TRMTLSLINSKMVFEDLC*ICMIFKTMYG 94
SL L *TRMTL L NSKM FED C*ICMIFKT G
Sbjct: 389 SL*LR*TRMTLLLSNSKMAFEDSC*ICMIFKTTSG 493
>BP053195
Length = 417
Score = 73.2 bits (178), Expect = 7e-14
Identities = 50/96 (52%), Positives = 59/96 (61%)
Frame = -3
Query: 281 LIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHLME 340
LII GVM EE+ KLLM +V+SL VT LLIAR+ VRL VM L F H LQK +
Sbjct: 415 LIITRGVMAEEHPKLLM*PVVISLAGVTTLLIARETVRLGVVMFLLFHHPPPLQK--PLP 242
Query: 341 TRNLLIIVKQAMHIVVEQL**CNWLC*LWLPQLSWH 376
T II++ AM I+ E *C+ LC* WL SWH
Sbjct: 241 TLTRQIIIQPAMLILGEPPS*CHPLC*FWLLPCSWH 134
>TC8429 weakly similar to UP|BAD03306 (BAD03306) MAP3K-like protein kinase,
partial (55%)
Length = 1298
Score = 45.8 bits (107), Expect = 1e-05
Identities = 73/230 (31%), Positives = 95/230 (40%), Gaps = 1/230 (0%)
Frame = +1
Query: 66 TRMTLSLINSKMVFEDLC*ICMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCVHFWTQ 125
TR I+S+MV ED C C I K +GCV+ L N+ + * R C TQ
Sbjct: 379 TRKIPLQISSRMV*ED*CWTCGIMKIPFGCVVGLAPNIQPFNLPLMF*KR-SECSS*HTQ 555
Query: 126 IHLK*SPSLLRIMLELLQP*LRSSKLQGLTSTCSRLVGCPRTVAIGLLLMT*S*TINDLS 185
+ SP L IM * R ++ + + C R IG * I
Sbjct: 556 --QRSSPFSLMIM*HQEMV*TRCLIKLD*GNSGFQCLKCLRMEVIGQQSKQ**GRIIG*L 729
Query: 186 HSPRDLPRKQLKAFLSRGNTS*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F**TFFIL 245
S + L + LK G *K N+EM VLV+I++N Q*T H* **T +
Sbjct: 730 CSLQMLLGRLLKELPMSGIML*KANLEMLGLREVLVKIDLNHFQ*TMLQSH*S**TTSEM 909
Query: 246 HRIVLRPVATI-QLHYSAC*RLAMKRPATDGQISLLLIII*GVMVEEYHK 294
I++ A I LH A++ TDG LLL EE K
Sbjct: 910 SEIMMMKHAGITHLH*LQ*CMSALELRVTDGLTLLLLTFTREATEEEPQK 1059
>TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749), partial
(20%)
Length = 697
Score = 33.1 bits (74), Expect = 0.078
Identities = 15/35 (42%), Positives = 23/35 (64%)
Frame = -1
Query: 314 RQMVRLVHVMSLQFLHLHLLQKLHLMETRNLLIIV 348
R+ +R H+ L+ LHLHLL LHL ++LI++
Sbjct: 466 RRRLRHSHLRLLRLLHLHLLHLLHLPVLSHILIVI 362
>TC8379 similar to UP|Q9SNJ3 (Q9SNJ3) ESTs AU031435(E61570), partial (5%)
Length = 809
Score = 30.8 bits (68), Expect = 0.39
Identities = 16/55 (29%), Positives = 31/55 (56%)
Frame = +2
Query: 301 VVSLVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHLMETRNLLIIVKQAMHIV 355
+ LV L I + + ++S+ +HL LLQ+++L + +NL+I ++H V
Sbjct: 230 LTQLVAARTLAIMLHSLSMTRILSVPLVHLRLLQRINL-QRKNLIIGGHSSLHAV 391
>BP063326
Length = 492
Score = 30.0 bits (66), Expect = 0.66
Identities = 28/75 (37%), Positives = 37/75 (49%), Gaps = 1/75 (1%)
Frame = -1
Query: 208 KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F**TFFILHRIVLRPVATI-QLHYSAC*RL 266
K +EM VLV+I++N Q*T H* **T + I++ A I LH
Sbjct: 405 KXPLEMLGLREVLVKIDLNHFQ*TMLQSH*S**TTSEMSEIMMMKHAGITHLH*LQ*CMS 226
Query: 267 AMKRPATDGQISLLL 281
A++ TDG LLL
Sbjct: 225 ALELRVTDGLTLLLL 181
>AV767819
Length = 528
Score = 27.7 bits (60), Expect = 3.3
Identities = 19/55 (34%), Positives = 31/55 (55%), Gaps = 2/55 (3%)
Frame = +1
Query: 292 YHKLLMQ--QMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHLMETRNL 344
YH L + MVV+++ V A+ +AR + L+ LHL L +KLH ++ + L
Sbjct: 244 YHSLSVTIVAMVVAVI-VVAVGVARVAAFFPRKLELRNLHLSLHEKLHFLQLQVL 405
>BP032935
Length = 534
Score = 26.2 bits (56), Expect = 9.5
Identities = 16/61 (26%), Positives = 28/61 (45%), Gaps = 2/61 (3%)
Frame = +2
Query: 82 LC*ICMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCVH--FWTQIHLK*SPSLLRIML 139
LC I M+G + +E N++ +H YL R+ H W+ + + S+ I L
Sbjct: 59 LCSFWQIMGLMFGLLTHVEPNIACDIHGYLVTARIIGIGHGMNWSHMIFRQHSSMCTIKL 238
Query: 140 E 140
+
Sbjct: 239 D 241
>BP080174
Length = 380
Score = 26.2 bits (56), Expect = 9.5
Identities = 7/18 (38%), Positives = 14/18 (76%)
Frame = +1
Query: 3 VLHVKHVLQMEIHDPDVA 20
+LH+KH++ + H+P +A
Sbjct: 82 LLHIKHIISLNYHNPSLA 135
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.158 0.540
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,513,533
Number of Sequences: 28460
Number of extensions: 89994
Number of successful extensions: 987
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 969
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 982
length of query: 377
length of database: 4,897,600
effective HSP length: 92
effective length of query: 285
effective length of database: 2,279,280
effective search space: 649594800
effective search space used: 649594800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144431.6


