

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
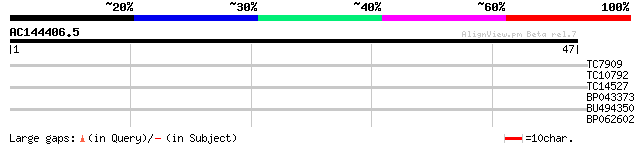
Query= AC144406.5 - phase: 0
(47 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7909 similar to UP|CB4C_ARATH (Q9S7W1) Chlorophyll a-b binding... 28 0.51
TC10792 27 1.1
TC14527 similar to PIR|F86340|F86340 protein F2D10.34 [imported]... 26 1.9
BP043373 25 2.5
BU494350 24 7.4
BP062602 23 9.7
>TC7909 similar to UP|CB4C_ARATH (Q9S7W1) Chlorophyll a-b binding protein
CP29.3, chloroplast precursor (LHCII protein 4.3)
(LHCB4.3), partial (80%)
Length = 1155
Score = 27.7 bits (60), Expect = 0.51
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 5 SRCPGRTRLLLTLEDHWKRPGTRACRER-STPG 36
SR P RT ++ HW +P R CR R S+PG
Sbjct: 522 SRKPPRTAETASVGLHWSQPWFR*CRRRDSSPG 424
>TC10792
Length = 480
Score = 26.6 bits (57), Expect = 1.1
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -1
Query: 20 HWKRPGTRACRERSTPGGRASTVSPG 45
H K+P R C+E+ +PG + PG
Sbjct: 216 HQKQPTQRPCKEKRSPGTTQTLGKPG 139
>TC14527 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(42%)
Length = 1095
Score = 25.8 bits (55), Expect = 1.9
Identities = 12/36 (33%), Positives = 15/36 (41%)
Frame = +3
Query: 5 SRCPGRTRLLLTLEDHWKRPGTRACRERSTPGGRAS 40
SRCP E W+R R STP R++
Sbjct: 447 SRCPNSPTQTAARESGWRRRSARFAWRSSTPATRSA 554
>BP043373
Length = 460
Score = 25.4 bits (54), Expect = 2.5
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = +2
Query: 2 GGRSRCPGRTRLLLTLEDHWKRPGTRACRER 32
GGR +L+L +ED W+R R R++
Sbjct: 188 GGRRHNTQVLKLMLKIEDEWQRSRRRCRRQK 280
>BU494350
Length = 537
Score = 23.9 bits (50), Expect = 7.4
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +1
Query: 21 WKRPGTRACRERSTP 35
W RPGT+ RER +P
Sbjct: 124 WFRPGTKYYRERLSP 168
>BP062602
Length = 536
Score = 23.5 bits (49), Expect = 9.7
Identities = 7/17 (41%), Positives = 12/17 (70%)
Frame = +1
Query: 19 DHWKRPGTRACRERSTP 35
+HWK G+R ++ +TP
Sbjct: 427 NHWKESGSRTTKKLATP 477
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,044,317
Number of Sequences: 28460
Number of extensions: 11406
Number of successful extensions: 78
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 78
length of query: 47
length of database: 4,897,600
effective HSP length: 23
effective length of query: 24
effective length of database: 4,243,020
effective search space: 101832480
effective search space used: 101832480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 49 (23.5 bits)
Medicago: description of AC144406.5


