

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
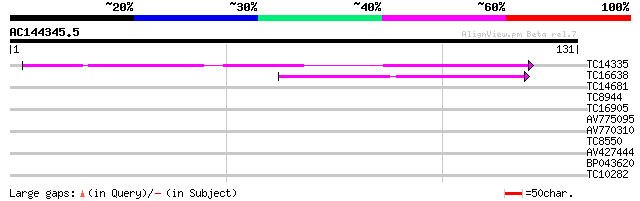
Query= AC144345.5 - phase: 0
(131 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14335 similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase ... 64 1e-11
TC16638 weakly similar to UP|MEP1_SOYBN (P29136) Metalloendoprot... 44 9e-06
TC14681 similar to UP|Q9SLZ4 (Q9SLZ4) Retinoblastoma-related pro... 28 0.67
TC8944 similar to UP|Q9SIR2 (Q9SIR2) At2g25310 protein, partial ... 27 1.5
TC16905 similar to UP|Q8GUE9 (Q8GUE9) Tyrosine aminotransferase ... 24 7.4
AV775095 24 7.4
AV770310 24 7.4
TC8550 similar to UP|Q94IP5 (Q94IP5) Mono-lipoyl E2, partial (21%) 24 9.7
AV427444 24 9.7
BP043620 24 9.7
TC10282 similar to GB|AAP31938.1|30387545|BT006594 At2g32480 {Ar... 24 9.7
>TC14335 similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor
(SMEP1) , partial (9%)
Length = 1574
Score = 63.5 bits (153), Expect = 1e-11
Identities = 40/118 (33%), Positives = 58/118 (48%)
Frame = +3
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNS 63
VFR AF RWS T LNF+ET ++D ++IKI F + G VG +
Sbjct: 591 VFREAFDRWSNETS-LNFTETATFDASDIKIAFVKWDGKGGR----VGAAYTNYSMHAGG 755
Query: 64 GFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAIL 121
F+ + + ++ L MH+IGHLLGL HS++KE +MYP ++
Sbjct: 756 MFLDI------------------DEQYVLLNVVMHEIGHLLGLGHSYEKESIMYPIVM 875
>TC16638 weakly similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1
precursor (SMEP1) , partial (29%)
Length = 621
Score = 43.9 bits (102), Expect = 9e-06
Identities = 24/58 (41%), Positives = 35/58 (59%)
Frame = +2
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
+G L A++ WV+ D + DLE+ A+H+IGHLLGL HS +E +M+P I
Sbjct: 23 NGRFHLDAAEDWVVSGDVSKSALATA-VDLESVAVHEIGHLLGLGHSSVEEAIMFPTI 193
>TC14681 similar to UP|Q9SLZ4 (Q9SLZ4) Retinoblastoma-related protein,
partial (22%)
Length = 911
Score = 27.7 bits (60), Expect = 0.67
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = +1
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVV 50
VFR+ F WS R + T D +I I FYN +I V ++V
Sbjct: 103 VFRSVFVDWSSARRNGASKQRTGQDHVDI-ITFYNQIFIPSVKPLLV 240
>TC8944 similar to UP|Q9SIR2 (Q9SIR2) At2g25310 protein, partial (69%)
Length = 1150
Score = 26.6 bits (57), Expect = 1.5
Identities = 13/63 (20%), Positives = 31/63 (48%)
Frame = +3
Query: 31 NIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQNGEF 90
N++ GF+ I + D ++ + +++ S S YWV +++WS ++ +F
Sbjct: 78 NLRYGFHQITHSFSFDPIMFLTSTLRIRS----------ISCYWVYRRFHHLWSSEDPQF 227
Query: 91 DLE 93
++
Sbjct: 228 GIK 236
>TC16905 similar to UP|Q8GUE9 (Q8GUE9) Tyrosine aminotransferase , partial
(37%)
Length = 610
Score = 24.3 bits (51), Expect = 7.4
Identities = 12/23 (52%), Positives = 15/23 (65%), Gaps = 2/23 (8%)
Frame = -2
Query: 108 HSFDKEYVMYPAILPLQ--QQRK 128
H+FDK V YP L +Q QQ+K
Sbjct: 474 HAFDKNIVSYPCPLQVQKGQQQK 406
>AV775095
Length = 358
Score = 24.3 bits (51), Expect = 7.4
Identities = 10/34 (29%), Positives = 16/34 (46%)
Frame = +3
Query: 77 PTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSF 110
P Y SW+ G E+ + + GH+ G S+
Sbjct: 222 PNLRYHQSWERGRVKAESIRIKRYGHISGSSASW 323
>AV770310
Length = 466
Score = 24.3 bits (51), Expect = 7.4
Identities = 6/13 (46%), Positives = 12/13 (92%)
Frame = +1
Query: 72 KYWVLPTDNYMWS 84
K+WV+P++N +W+
Sbjct: 43 KWWVVPSNNLLWT 81
>TC8550 similar to UP|Q94IP5 (Q94IP5) Mono-lipoyl E2, partial (21%)
Length = 852
Score = 23.9 bits (50), Expect = 9.7
Identities = 7/15 (46%), Positives = 11/15 (72%)
Frame = +3
Query: 69 IASKYWVLPTDNYMW 83
+ S YW++ T +YMW
Sbjct: 771 VISGYWLIQTYDYMW 815
>AV427444
Length = 428
Score = 23.9 bits (50), Expect = 9.7
Identities = 8/25 (32%), Positives = 13/25 (52%)
Frame = -2
Query: 98 HQIGHLLGLDHSFDKEYVMYPAILP 122
H+ H G DH ++ +P +LP
Sbjct: 268 HKFHHGAGADHQMYSSFLEHPQVLP 194
>BP043620
Length = 507
Score = 23.9 bits (50), Expect = 9.7
Identities = 15/56 (26%), Positives = 24/56 (42%)
Frame = +1
Query: 50 VGDTVIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLG 105
V T IKL ++ K W P+ +Y WS++ F ++ +G LG
Sbjct: 328 VWSTTIKLAFRPCLFRWTILLRKIWFSPSGSY*WSFRLAHFWFLSSRSFLVGCWLG 495
>TC10282 similar to GB|AAP31938.1|30387545|BT006594 At2g32480 {Arabidopsis
thaliana;}, partial (21%)
Length = 644
Score = 23.9 bits (50), Expect = 9.7
Identities = 8/26 (30%), Positives = 13/26 (49%)
Frame = +3
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLL 104
++Y+W W NG E + G+ L
Sbjct: 165 ESYLWKWSNGSCRQELCLFYFWGYTL 242
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,194,735
Number of Sequences: 28460
Number of extensions: 26175
Number of successful extensions: 124
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 123
length of query: 131
length of database: 4,897,600
effective HSP length: 80
effective length of query: 51
effective length of database: 2,620,800
effective search space: 133660800
effective search space used: 133660800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 50 (23.9 bits)
Medicago: description of AC144345.5


