

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.11 - phase: 0
(339 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
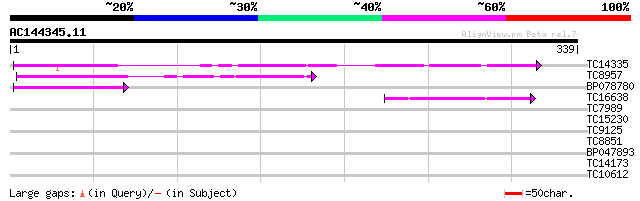
TC14335 similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase ... 111 2e-25
TC8957 weakly similar to UP|Q9ZUJ5 (Q9ZUJ5) T2K10.2 protein, par... 75 1e-14
BP078780 59 1e-09
TC16638 weakly similar to UP|MEP1_SOYBN (P29136) Metalloendoprot... 58 2e-09
TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , par... 33 0.089
TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Ar... 27 6.4
TC9125 similar to UP|Q40316 (Q40316) Vestitone reductase, complete 27 6.4
TC8851 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-phos... 27 6.4
BP047893 27 6.4
TC14173 homologue to UP|RL10_SOLME (P93847) 60S ribosomal protei... 26 8.4
TC10612 homologue to UP|RL10_VITRI (Q9SPB3) 60S ribosomal protei... 26 8.4
>TC14335 similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor
(SMEP1) , partial (9%)
Length = 1574
Score = 111 bits (277), Expect = 2e-25
Identities = 90/320 (28%), Positives = 134/320 (41%), Gaps = 4/320 (1%)
Frame = +3
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFD----DVLDEETISAIKTYQQFFNLQVTGNLNTETL 58
++GL IKK+L GY + +L + D E+ AI YQ+ FNL VTG ++
Sbjct: 258 VKGLENIKKYLDALGYIKDLVLALNSNDTDKFTEDLKPAIMEYQKNFNLPVTGVIDQNLY 437
Query: 59 QKISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPD 118
+ PR CG+PD
Sbjct: 438 NTLIQPR------------------------------------------------CGVPD 473
Query: 119 MRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRV 178
+VG+ +F W+ K+L+Y F P S D+ FR AF RWS T
Sbjct: 474 FTT---LNVGTTPAF---KPWWKSEKKELSYAFHPQNNVSDDIRTVFREAFDRWSNETS- 632
Query: 179 LNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVS 238
LNF+ET ++D +DIKI F + + K G + + + +
Sbjct: 633 LNFTETATFDASDIKIAF----------------------VKWDGKGGRVGAAYTNYSMH 746
Query: 239 PTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQ 298
+ E ++ L VVMH+IGHLLGL HS +KESIMYP++ +K+ TD D +
Sbjct: 747 AGGMFLDIDE--QYVLLNVVMHEIGHLLGLGHSYEKESIMYPIV---MREKIDFTDDDRK 911
Query: 299 AIQQLYTNTAKANPNSDHSG 318
IQ++ N+ ++ SG
Sbjct: 912 RIQKVTGNSNNGGTSASSSG 971
>TC8957 weakly similar to UP|Q9ZUJ5 (Q9ZUJ5) T2K10.2 protein, partial (20%)
Length = 724
Score = 75.5 bits (184), Expect = 1e-14
Identities = 58/180 (32%), Positives = 80/180 (44%), Gaps = 1/180 (0%)
Frame = +1
Query: 5 GLSQIKKHLSTFGYFRQFL-LKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISL 63
GLS +K + FGY F D D+ +AIKTYQ+ FNL +TG+L+ +TL++I L
Sbjct: 271 GLSNLKSYFKRFGYIPHAPPSNFSDDFDDALEAAIKTYQKNFNLNITGDLDDKTLRQIML 450
Query: 64 PRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEY 123
PRCG+ D+ GT + G + G ++ +
Sbjct: 451 PRCGVADI---------------------INGTTTMN---AAGNQDETASHGDSNLHFH- 555
Query: 124 GFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSE 183
V FP G P G ++LTY F PG S + F AF RWS+ T L F E
Sbjct: 556 --TVSHYALFP-GQPRGPAGKQELTYAFYPGNSLSDAVKSVFTGAFARWSEVT-TLTFRE 723
>BP078780
Length = 516
Score = 58.9 bits (141), Expect = 1e-09
Identities = 29/70 (41%), Positives = 42/70 (59%), Gaps = 1/70 (1%)
Frame = +3
Query: 3 IQGLSQIKKHLSTFGYFRQFL-LKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
+ GLSQ++ +L FGY L F + E SAI+ +Q+ +NLQVTG L+ +
Sbjct: 255 VDGLSQVQDYLDLFGYLNSTLHSNFTNEYTENAKSAIEEFQKNYNLQVTGQLDNNLYHVL 434
Query: 62 SLPRCGIPDM 71
S PRCG+PD+
Sbjct: 435 SQPRCGVPDI 464
>TC16638 weakly similar to UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1
precursor (SMEP1) , partial (29%)
Length = 621
Score = 58.2 bits (139), Expect = 2e-09
Identities = 35/90 (38%), Positives = 51/90 (55%)
Frame = +2
Query: 225 SGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDP 284
+G L+ ++ WV + K DLE+V +H+IGHLLGL HSS +E+IM+P I
Sbjct: 23 NGRFHLDAAEDWVV-SGDVSKSALATAVDLESVAVHEIGHLLGLGHSSVEEAIMFPTISS 199
Query: 285 LQEKKVQITDSDNQAIQQLYTNTAKANPNS 314
+ KKV + + D + IQ LY N +S
Sbjct: 200 -RTKKVVLAEDDIKGIQYLYGTNPSFNGSS 286
>TC7989 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (85%)
Length = 1352
Score = 32.7 bits (73), Expect = 0.089
Identities = 22/74 (29%), Positives = 33/74 (43%), Gaps = 8/74 (10%)
Frame = +3
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQ----EKKVQIT 293
S TTT + D VV GH +G+ H S +YP DP+ EK +++T
Sbjct: 567 SNTTTILNSLATKNLDPTDVVSLSGGHTIGISHCSSFTDRLYPSKDPVMDQTFEKNLKLT 746
Query: 294 ----DSDNQAIQQL 303
++DN + L
Sbjct: 747 CPASNTDNTTVLDL 788
>TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Arabidopsis
thaliana;}, partial (11%)
Length = 611
Score = 26.6 bits (57), Expect = 6.4
Identities = 11/34 (32%), Positives = 20/34 (58%)
Frame = +3
Query: 89 NKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
+KW +K L + ++ GKKF L C +P++ +
Sbjct: 186 SKWEKMTSKSLEFLWVRGKKFYSLSCLVPNVHID 287
>TC9125 similar to UP|Q40316 (Q40316) Vestitone reductase, complete
Length = 1200
Score = 26.6 bits (57), Expect = 6.4
Identities = 20/60 (33%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Frame = +1
Query: 246 KWELQEFDLETVVMHQIGHLL-GLDHSSDKESIM-YPLIDPLQEKKVQITDSDNQAIQQL 303
KW+ ++ E+V + G LL GL S K I+ PL+DP+Q + + S +Q L
Sbjct: 37 KWQREKE--ESVSLEVQGFLLHGLSRDSLKMDILSIPLLDPIQVVREMLASSQIYLVQHL 210
>TC8851 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-phosphate
1-dehydrogenase {Arabidopsis thaliana;} , partial (54%)
Length = 1291
Score = 26.6 bits (57), Expect = 6.4
Identities = 30/135 (22%), Positives = 54/135 (39%), Gaps = 15/135 (11%)
Frame = +1
Query: 171 RWSQTTRVLNFSETTSYDDADIKIGFYHI----YNNDIVDDV----------VVGDSFIS 216
RW ++ + A+I++ F H+ YN + D+ V D I
Sbjct: 481 RWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNVYNRNFGTDLDRATNELVIRVQPDEAIY 660
Query: 217 RNLDSNVKSGMIRLERSKFWVSPTTTYFKKW-ELQEFDLETVVMHQIGHLLGLDHSSDKE 275
+++ V +RL+RS + Y K+ + E L + + + D
Sbjct: 661 LKINNKVPGLEMRLDRSNLNLHYAARYAKEIPDAYERLLLDAIEGERRLFIRSDELDAAW 840
Query: 276 SIMYPLIDPLQEKKV 290
S+ PL++ L+EKKV
Sbjct: 841 SLFTPLLNELEEKKV 885
>BP047893
Length = 514
Score = 26.6 bits (57), Expect = 6.4
Identities = 17/39 (43%), Positives = 20/39 (50%), Gaps = 10/39 (25%)
Frame = -3
Query: 259 MHQIGHLL----------GLDHSSDKESIMYPLIDPLQE 287
MH I +LL LDHSS+K+ M L PLQE
Sbjct: 230 MHSIAYLLLPSHWFRLSSQLDHSSEKKKEM*QLPSPLQE 114
>TC14173 homologue to UP|RL10_SOLME (P93847) 60S ribosomal protein L10
(EQM), partial (90%)
Length = 611
Score = 26.2 bits (56), Expect = 8.4
Identities = 11/42 (26%), Positives = 19/42 (45%)
Frame = -3
Query: 84 SFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
SF + +W KG + F+P +L G P + + G+
Sbjct: 192 SFSQLTRWTQKGNSSTPFFFIPTS*IRILGSGTPRQKRDLGY 67
>TC10612 homologue to UP|RL10_VITRI (Q9SPB3) 60S ribosomal protein L10 (QM
protein homolog), partial (68%)
Length = 474
Score = 26.2 bits (56), Expect = 8.4
Identities = 10/42 (23%), Positives = 18/42 (42%)
Frame = -1
Query: 84 SFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
SF +W KG + F+P + G P + ++G+
Sbjct: 198 SFSHETRWTQKGNSSTPFFFIPTS*IRIFGSGTPRQKRDFGY 73
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,679,820
Number of Sequences: 28460
Number of extensions: 71173
Number of successful extensions: 283
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 258
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 278
length of query: 339
length of database: 4,897,600
effective HSP length: 91
effective length of query: 248
effective length of database: 2,307,740
effective search space: 572319520
effective search space used: 572319520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144345.11


