

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.6 + phase: 0
(683 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
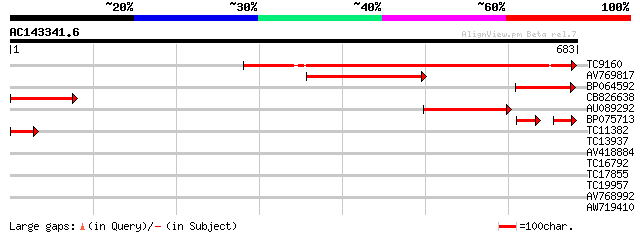
Score E
Sequences producing significant alignments: (bits) Value
TC9160 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partial ... 421 e-118
AV769817 192 1e-49
BP064592 133 8e-32
CB826638 131 3e-31
AU089292 121 4e-28
BP075713 43 4e-09
TC11382 similar to PIR|T01900|T01900 isp4 protein homolog T12H20... 44 7e-05
TC13937 weakly similar to UP|Q9LVE0 (Q9LVE0) Nitrate transporter... 31 0.75
AV418884 30 0.99
TC16792 28 3.7
TC17855 28 4.9
TC19957 28 6.4
AV768992 27 8.3
AW719410 27 8.3
>TC9160 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partial (53%)
Length = 1554
Score = 421 bits (1081), Expect = e-118
Identities = 191/401 (47%), Positives = 271/401 (66%)
Frame = +2
Query: 282 TKTFDIDMESYNNYSKIYLSVTFAFQYGLSFAALTATISHVVLFHGEMIVQMWKKTTTSL 341
T +++++++YN Y K+YLS FA G FA TAT++HV LFHG I++ +
Sbjct: 11 TPDYELNVDAYNQYGKLYLSPLFALSIGSGFARFTATLTHVALFHGSDILRQSRSAMNKA 190
Query: 342 KNQLGDVHTRIMKKNYEQVPEWWFVAILILMVMMALLACEGFGKQLQLPWWGILLSLSIA 401
K DVH R+MK Y+QVPEWWF+ +L + ++L+ + + +QLPWWG+L + ++A
Sbjct: 191 KL---DVHGRLMKA-YKQVPEWWFLILLFGSMALSLVMSFVWKRDVQLPWWGMLFAFALA 358
Query: 402 LVFTLPIGVIEATTSARTGLNVITELVIGFTYPGKPLANVAFKTYGHTSMVQALGFLGDF 461
+ TLPIGVI+ATT+ + G ++I + +IG+ PGKP+AN+ FK YG S V AL FL D
Sbjct: 359 FIVTLPIGVIQATTNQQPGYDIIAQFMIGYVLPGKPIANLLFKIYGRISTVHALSFLSDL 538
Query: 462 KLGHYMKIPPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENICDESLLPKGSPWTCPGDD 521
KLGHYMKIPP+ M+ QLVGT+V+ V+ AWW+L +I++IC + +PWTCP
Sbjct: 539 KLGHYMKIPPRCMYTAQLVGTLVASIVNLAVAWWMLDNIKDICMDDKAHHDNPWTCPKFR 718
Query: 522 AFYNASIIWGVVGPKRMFTNDGVYPGMNWFFLIGLIAPVPVWLLSIKFPNHKWIKLINIP 581
++AS+IWG++GP+R+F G+Y + W FLIG + PVPVW+LS FP KWI LINIP
Sbjct: 719 VTFDASVIWGLIGPRRLFGPGGLYRNLVWLFLIGAVLPVPVWVLSKIFPEKKWIALINIP 898
Query: 582 IIIVGASTIPPRRSVNYITWGIVGMFFNFYVYRKFRVWWARHTYILSAGLDAGVAFMGLV 641
+I G + +PP N +W + GM FNF+V+R + WW ++ Y+LSA LDAG AFMG++
Sbjct: 899 VISYGFAGMPPATPSNIASWLVTGMIFNFFVFRYHQRWWQKYNYVLSAALDAGTAFMGVL 1078
Query: 642 LYFSLQSYGIFGPTWWGLEADHCPLARCPTAPGVYAEGCPV 682
++F+L + G WWG E D CPLA CPTAPG+ +GCPV
Sbjct: 1079IFFALPNAG-HSLKWWGSEMDPCPLASCPTAPGIKVDGCPV 1198
>AV769817
Length = 436
Score = 192 bits (488), Expect = 1e-49
Identities = 82/145 (56%), Positives = 111/145 (76%)
Frame = +2
Query: 358 EQVPEWWFVAILILMVMMALLACEGFGKQLQLPWWGILLSLSIALVFTLPIGVIEATTSA 417
E +P WWF A+L+ + ++L+ C Q+Q+PWWG+L + ++A VFTLPI +I ATT+
Sbjct: 2 EDIPSWWFYALLVATLAISLVLCIFLNDQIQMPWWGLLFAGALAFVFTLPISIITATTNQ 181
Query: 418 RTGLNVITELVIGFTYPGKPLANVAFKTYGHTSMVQALGFLGDFKLGHYMKIPPKSMFIV 477
GLN+ITE V G YPG+P+ANV FKTYG+ SM QA+ FL DFKLGHYMKIPP+SMF+V
Sbjct: 182 TPGLNIITEYVFGLIYPGRPIANVCFKTYGYISMAQAVSFLSDFKLGHYMKIPPRSMFLV 361
Query: 478 QLVGTVVSLSVHFGTAWWLLTSIEN 502
Q +GTV++ +++ G AWWLL SI+N
Sbjct: 362 QFIGTVLAGTINIGVAWWLLNSIDN 436
>BP064592
Length = 531
Score = 133 bits (335), Expect = 8e-32
Identities = 56/72 (77%), Positives = 62/72 (85%)
Frame = -2
Query: 610 FYVYRKFRVWWARHTYILSAGLDAGVAFMGLVLYFSLQSYGIFGPTWWGLEADHCPLARC 669
FYVYR F+ WW TYILSA LDAGVAFMGL+LYF+L S G+FGPTWWGL+ DHCPLARC
Sbjct: 530 FYVYRNFKXWWXSXTYILSAALDAGVAFMGLLLYFALPSNGVFGPTWWGLDNDHCPLARC 351
Query: 670 PTAPGVYAEGCP 681
PT PGVYA+GCP
Sbjct: 350 PTYPGVYAKGCP 315
>CB826638
Length = 575
Score = 131 bits (330), Expect = 3e-31
Identities = 64/81 (79%), Positives = 75/81 (92%)
Frame = +1
Query: 1 MNQFLGYKTNPLKITSVSAQIITLPLGKLMAATLPIKTIQVPFTSLTFSLNPGSFSVKEH 60
+NQFLGY+TNP+ ITSVSAQI+TLPLGKLMAATLP + I+VPFTS +FSLNPG FS+KEH
Sbjct: 331 VNQFLGYRTNPMNITSVSAQIVTLPLGKLMAATLPTRPIRVPFTSWSFSLNPGPFSMKEH 510
Query: 61 VLISIFATSGSSGVYAINIIT 81
VLI+IFA+SGS GVYAI+IIT
Sbjct: 511 VLITIFASSGSGGVYAISIIT 573
>AU089292
Length = 345
Score = 121 bits (303), Expect = 4e-28
Identities = 51/106 (48%), Positives = 76/106 (71%)
Frame = +2
Query: 499 SIENICDESLLPKGSPWTCPGDDAFYNASIIWGVVGPKRMFTNDGVYPGMNWFFLIGLIA 558
+I ++CD S LP PW CP D+ FY+AS+IWG++GP+R+F + GVY +N+FFL G IA
Sbjct: 11 TIPDLCDASKLPTDXPWLCPMDNVFYDASVIWGLLGPRRIFGDQGVYGKVNYFFLGGAIA 190
Query: 559 PVPVWLLSIKFPNHKWIKLINIPIIIVGASTIPPRRSVNYITWGIV 604
P+ VWL FP +W++LI++PI++ S +PP +VNY +W +V
Sbjct: 191 PLLVWLAHKAFPXKEWLRLIHMPIMLGATSMMPPATAVNYTSWLLV 328
>BP075713
Length = 381
Score = 42.7 bits (99), Expect(2) = 4e-09
Identities = 16/27 (59%), Positives = 17/27 (62%)
Frame = -2
Query: 656 WWGLEADHCPLARCPTAPGVYAEGCPV 682
WWG E D CPLA CP PG +GC V
Sbjct: 245 WWGSEMDPCPLASCPLPPGSKVDGCHV 165
Score = 35.4 bits (80), Expect(2) = 4e-09
Identities = 14/29 (48%), Positives = 19/29 (65%)
Frame = -3
Query: 611 YVYRKFRVWWARHTYILSAGLDAGVAFMG 639
+V+R WW ++ Y+LSA LDAG A G
Sbjct: 379 FVFRYPHPWWQKYNYVLSAALDAGPALHG 293
>TC11382 similar to PIR|T01900|T01900 isp4 protein homolog T12H20.7 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(10%)
Length = 606
Score = 44.3 bits (103), Expect = 7e-05
Identities = 19/34 (55%), Positives = 25/34 (72%)
Frame = +3
Query: 1 MNQFLGYKTNPLKITSVSAQIITLPLGKLMAATL 34
+NQF Y+T PL +TS+SAQI +P+G MA TL
Sbjct: 504 VNQFFWYRTEPLTVTSISAQIAVVPIGHFMARTL 605
>TC13937 weakly similar to UP|Q9LVE0 (Q9LVE0) Nitrate transporter
(AT3g21670/MIL23_23), partial (27%)
Length = 558
Score = 30.8 bits (68), Expect = 0.75
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +2
Query: 421 LNVITELVIGFTYPGKPLANVAFKTYGHTSMVQALG-FLGDFKLGHYMKI 469
+N++T LV P A + G ++V LG FL D KLG Y+ I
Sbjct: 194 MNIVTYLVGVLNLPTAYSATIVTNCLGTMNLVSLLGGFLADVKLGRYLTI 343
>AV418884
Length = 418
Score = 30.4 bits (67), Expect = 0.99
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +2
Query: 1 MNQFLGYKTNPLKITSVSAQIITL 24
+NQF Y+T PL IT ++ Q+ TL
Sbjct: 233 LNQFFAYRTEPLIITLITVQVSTL 304
>TC16792
Length = 558
Score = 28.5 bits (62), Expect = 3.7
Identities = 20/76 (26%), Positives = 34/76 (44%)
Frame = +1
Query: 270 TGATYNITRILNTKTFDIDMESYNNYSKIYLSVTFAFQYGLSFAALTATISHVVLFHGEM 329
TG+TY+ ++ T + ++Y Y+ L +A + A TA + VVLF
Sbjct: 22 TGSTYDPDELIETSVEKLGDQTYYKYA---LETPYALTGSHNLAKATAKGNTVVLFVASA 192
Query: 330 IVQMWKKTTTSLKNQL 345
Q W + ++K L
Sbjct: 193 NDQQWPTSQ*TVKAML 240
>TC17855
Length = 543
Score = 28.1 bits (61), Expect = 4.9
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +1
Query: 651 IFGPTWWGLEADHCPLARCPTAPGVYAEGCPV 682
+F ++W L DHC + C G AEG P+
Sbjct: 247 LFSISYWSLFEDHCCIKGCSRFWGCDAEGQPM 342
>TC19957
Length = 642
Score = 27.7 bits (60), Expect = 6.4
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = +1
Query: 134 SLFRAFHEKEKRPKGGTSRLQFFFLVFVASFAYYIVPGYFF 174
SL+ + + K P GG++ FF + S A PG+FF
Sbjct: 100 SLYEQYVGERK*PLGGSNCRLFFLATTLVSGASLAFPGFFF 222
>AV768992
Length = 473
Score = 27.3 bits (59), Expect = 8.3
Identities = 11/18 (61%), Positives = 13/18 (72%), Gaps = 1/18 (5%)
Frame = -1
Query: 616 FRVWWARHTYI-LSAGLD 632
F WWARH +I +S GLD
Sbjct: 395 FSAWWARHPWISISVGLD 342
>AW719410
Length = 538
Score = 27.3 bits (59), Expect = 8.3
Identities = 12/26 (46%), Positives = 19/26 (72%)
Frame = -3
Query: 455 LGFLGDFKLGHYMKIPPKSMFIVQLV 480
+GFLG F++ H ++PPK FI+ L+
Sbjct: 260 VGFLGVFRVRHDARLPPK--FIMSLI 189
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.142 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,458,462
Number of Sequences: 28460
Number of extensions: 210183
Number of successful extensions: 1411
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 1400
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1409
length of query: 683
length of database: 4,897,600
effective HSP length: 96
effective length of query: 587
effective length of database: 2,165,440
effective search space: 1271113280
effective search space used: 1271113280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC143341.6


