

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.5 + phase: 0
(485 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
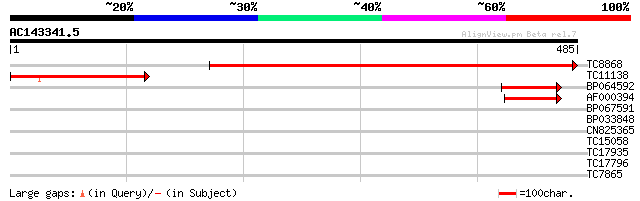
Score E
Sequences producing significant alignments: (bits) Value
TC8868 similar to UP|AAR06294 (AAR06294) Adenylosuccinate syntha... 573 e-164
TC11138 homologue to UP|PURA_ARATH (Q96529) Adenylosuccinate syn... 198 1e-51
BP064592 71 3e-13
AF000394 70 8e-13
BP067591 29 1.5
BP033848 29 1.5
CN825365 28 3.3
TC15058 27 5.7
TC17935 27 5.7
TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 27 5.7
TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinas... 27 5.7
>TC8868 similar to UP|AAR06294 (AAR06294) Adenylosuccinate synthase ,
partial (62%)
Length = 1273
Score = 573 bits (1478), Expect = e-164
Identities = 281/314 (89%), Positives = 297/314 (94%)
Frame = +1
Query: 172 QTVDGLREAELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAAL 231
Q VDGLR AEL+KSFIGTTKRGIGPCY+SK NRNG+RV DLRYM+T PQKL++LLSDAA
Sbjct: 1 QEVDGLR*AELAKSFIGTTKRGIGPCYASKVNRNGLRVSDLRYMDTFPQKLEVLLSDAAE 180
Query: 232 RFKDFKYGPDVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDID 291
RFK F YGP+VL+EEVEKYKRYAERLEP+IADTVHVMNEAITQKKK+LVEGGQATMLDID
Sbjct: 181 RFKKFNYGPNVLKEEVEKYKRYAERLEPFIADTVHVMNEAITQKKKILVEGGQATMLDID 360
Query: 292 FGTYPFVTSSSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTEILGPGGDLLR 351
FGTYPFVTSSSPSAGGIC GLGIAPRV+GDL+GVVKAYTTRVGSGPFPTE LG GGDLLR
Sbjct: 361 FGTYPFVTSSSPSAGGICIGLGIAPRVVGDLIGVVKAYTTRVGSGPFPTENLGSGGDLLR 540
Query: 352 FAGQEFGTTTGRPRRCGWLDIVALSYSCQINGFSSLNLTKLDVLSDLDEIQLGVSYKHAD 411
F GQEFGTTTGRPRRCGWLDIVAL YSCQINGFSSLNLTKLDVLS+LDEIQLGVSYKH D
Sbjct: 541 FTGQEFGTTTGRPRRCGWLDIVALRYSCQINGFSSLNLTKLDVLSELDEIQLGVSYKHVD 720
Query: 412 GTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIH 471
GTP+QSFPSDL LLEQLKVEYE LPGWK+DISSIRNYSDLP+AAQ YVERIEELVGVPIH
Sbjct: 721 GTPIQSFPSDLSLLEQLKVEYEVLPGWKSDISSIRNYSDLPKAAQQYVERIEELVGVPIH 900
Query: 472 YIGVGPGRDALIFK 485
YIGVGPGRDALIFK
Sbjct: 901 YIGVGPGRDALIFK 942
>TC11138 homologue to UP|PURA_ARATH (Q96529) Adenylosuccinate synthetase,
chloroplast precursor (IMP--aspartate ligase) (AdSS)
(AMPSase) , partial (17%)
Length = 429
Score = 198 bits (504), Expect = 1e-51
Identities = 100/121 (82%), Positives = 105/121 (86%), Gaps = 2/121 (1%)
Frame = +1
Query: 1 MNISSLRLDSHAICTHQPQRPFAF--HNRFPRNVVVCAAKPVAPPPTKLAAADSSLRRIE 58
M+ISSL LDSHAIC +PQRPF+ H R RN VVC+AKPVAPP +KL AADSS RI
Sbjct: 28 MSISSLALDSHAICNPKPQRPFSTFNHYRTTRNFVVCSAKPVAPPASKLTAADSSASRIG 207
Query: 59 SLSQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPS 118
SLSQVSGVLGCQWGDEGKGKLVDILAQHF IVARCQGGANAGHTIYNAEGKKFALHLVPS
Sbjct: 208 SLSQVSGVLGCQWGDEGKGKLVDILAQHFDIVARCQGGANAGHTIYNAEGKKFALHLVPS 387
Query: 119 G 119
G
Sbjct: 388 G 390
Score = 33.1 bits (74), Expect = 0.10
Identities = 28/95 (29%), Positives = 38/95 (39%), Gaps = 2/95 (2%)
Frame = +2
Query: 39 PVAPPPTKLAAADSSLRRIESLSQVSGVLGCQWGDEGK--GKLVDILAQHFQIVARCQGG 96
P +PPPT A R + SG + + L+ +LA +++
Sbjct: 167 PSSPPPTPPPLASGRSARSPASWVASGAMRAKGSSLTS*LSTLISLLAVRAELML----- 331
Query: 97 ANAGHTIYNAEGKKFALHLVPSGILNEDTLCVIGN 131
G +GK L ILNEDTLCVIGN
Sbjct: 332 ---GIPFIMRKGKSSPFILFHPVILNEDTLCVIGN 427
>BP064592
Length = 531
Score = 71.2 bits (173), Expect = 3e-13
Identities = 34/52 (65%), Positives = 42/52 (80%)
Frame = -2
Query: 421 DLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIHY 472
++ LL + VEYE L GWK DISS RNYS+LP+AA+ YVERI+ELVGVPI +
Sbjct: 257 NIHLLFPISVEYEVLLGWKFDISSFRNYSELPKAAKQYVERIKELVGVPIRH 102
>AF000394
Length = 515
Score = 70.1 bits (170), Expect = 8e-13
Identities = 34/49 (69%), Positives = 40/49 (81%)
Frame = -3
Query: 424 LLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELVGVPIHY 472
LL + VEYE L GWK DISS RNYS+LP+AA+ YVERI+ELVGVPI +
Sbjct: 249 LLFPISVEYEVLLGWKFDISSFRNYSELPKAAKQYVERIKELVGVPIRH 103
>BP067591
Length = 358
Score = 29.3 bits (64), Expect = 1.5
Identities = 16/64 (25%), Positives = 32/64 (50%), Gaps = 3/64 (4%)
Frame = -2
Query: 3 ISSLRLDSHAICTHQPQRPFAF--HNRFPRNVVVCAAKPVAPPP-TKLAAADSSLRRIES 59
+S+L +H+ C H+ Q+PF+ F RN + P P K++ D++ ++
Sbjct: 216 VSALGYHTHSTCLHRYQQPFSVCSGKNFSRNGNIVVTPPAIPTSYPKISPPDAATMQVIK 37
Query: 60 LSQV 63
++ V
Sbjct: 36 IASV 25
>BP033848
Length = 525
Score = 29.3 bits (64), Expect = 1.5
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = +2
Query: 392 LDVLSDLDEIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPGW 438
LD++SDL I K + P+ RLLE +V++E LP W
Sbjct: 185 LDIVSDLSRILQTFDLK------LDLNPASRRLLEAEEVDHEGLPSW 307
>CN825365
Length = 511
Score = 28.1 bits (61), Expect = 3.3
Identities = 16/62 (25%), Positives = 29/62 (45%)
Frame = +1
Query: 71 WGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGILNEDTLCVIG 130
WG +G V++ A HF ++ + A H + G++ L ++P IL + L +
Sbjct: 148 WGSKGLAITVELQAHHFGVMGHLRSQYYATHVDLDG-GQREHLQIIPLYILGQTMLIMTI 324
Query: 131 NG 132
G
Sbjct: 325 KG 330
>TC15058
Length = 716
Score = 27.3 bits (59), Expect = 5.7
Identities = 14/54 (25%), Positives = 24/54 (43%)
Frame = +3
Query: 288 LDIDFGTYPFVTSSSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTE 341
+D+D T+ ++ P + P ++ DLV ++ V SG F TE
Sbjct: 144 VDVDIATHLTISDDGPDRSLLYVETADRPGLLLDLVQIITDINIAVESGEFDTE 305
>TC17935
Length = 543
Score = 27.3 bits (59), Expect = 5.7
Identities = 14/54 (25%), Positives = 24/54 (43%)
Frame = +2
Query: 288 LDIDFGTYPFVTSSSPSAGGICTGLGIAPRVIGDLVGVVKAYTTRVGSGPFPTE 341
+D+D T+ ++ P + P ++ DLV ++ V SG F TE
Sbjct: 32 VDVDIATHISISDDGPDRSLLYVEAADRPGLLVDLVKIITDIDLAVESGEFDTE 193
>TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 594
Score = 27.3 bits (59), Expect = 5.7
Identities = 15/41 (36%), Positives = 19/41 (45%)
Frame = -2
Query: 29 PRNVVVCAAKPVAPPPTKLAAADSSLRRIESLSQVSGVLGC 69
PRN+ + A PV P T L + + SLS V G C
Sbjct: 509 PRNIAITVALPVFVPFTTLELIGCNNSLMNSLSSVVGATCC 387
>TC7865 homologue to UP|SAHH_ARATH (O23255) Adenosylhomocysteinase
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase) ,
complete
Length = 1852
Score = 27.3 bits (59), Expect = 5.7
Identities = 20/67 (29%), Positives = 32/67 (46%), Gaps = 4/67 (5%)
Frame = -2
Query: 128 VIGNGVVVHLPGLFQEIDNLESNG----VSCKGRILISDRAHLLFDFHQTVDGLREAELS 183
V NG+V+HLP + D L + G +S + IL + L FH+++ G LS
Sbjct: 1104 VADNGIVLHLPHVVNHDDVLVTGGGDKDISLRDNIL---QGQNLKTFHESLKGTDGINLS 934
Query: 184 KSFIGTT 190
+ T+
Sbjct: 933 HNDTSTS 913
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,176,320
Number of Sequences: 28460
Number of extensions: 114807
Number of successful extensions: 556
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 552
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 555
length of query: 485
length of database: 4,897,600
effective HSP length: 94
effective length of query: 391
effective length of database: 2,222,360
effective search space: 868942760
effective search space used: 868942760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC143341.5


