

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.7 - phase: 0
(358 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
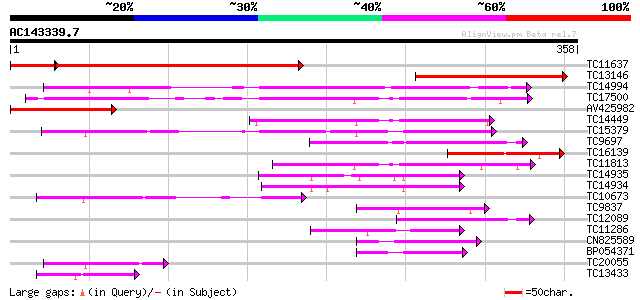
Sequences producing significant alignments: (bits) Value
TC11637 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-lik... 249 3e-76
TC13146 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-lik... 159 9e-40
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 127 2e-30
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 112 7e-26
AV425982 108 2e-24
TC14449 similar to UP|O24078 (O24078) Protein phosphatase 2C, pa... 100 4e-22
TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete 91 4e-19
TC9697 similar to GB|AAM63159.1|21554078|AY085949 protein phosph... 89 1e-18
TC16139 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-lik... 86 1e-17
TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-lik... 77 3e-15
TC14935 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-lik... 69 9e-13
TC14934 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-lik... 66 8e-12
TC10673 homologue to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, ... 56 8e-09
TC9837 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2... 52 2e-07
TC12089 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partia... 52 2e-07
TC11286 homologue to GB|AAO63448.1|28951049|BT005384 At4g03415 {... 51 3e-07
CN825589 49 1e-06
BP054371 47 6e-06
TC20055 45 2e-05
TC13433 similar to UP|Q8VZD9 (Q8VZD9) AT5g53140/MFH8_8, partial ... 44 3e-05
>TC11637 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-like protein
(AT4G31860/F11C18_60), partial (52%)
Length = 677
Score = 249 bits (637), Expect(2) = 3e-76
Identities = 118/156 (75%), Positives = 136/156 (86%)
Frame = +1
Query: 30 QGWRATMEDAHAAHLDVDSSTSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGT 89
+GWRA+MEDAHAA +D STS+F VYDGHGGKAV+KFCAK+LHQQ+LK E YIAGD+GT
Sbjct: 208 RGWRASMEDAHAALPCLDESTSYFAVYDGHGGKAVSKFCAKYLHQQMLKHEAYIAGDIGT 387
Query: 90 SLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKG 149
SL K FLRMDEMMRGQRGWRELAVLGDK++ +G++EGLI SPRS++ D+ DDWA E+G
Sbjct: 388 SLQKTFLRMDEMMRGQRGWRELAVLGDKMEKLSGMLEGLIWSPRSSEANDRVDDWALEEG 567
Query: 150 PHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVI 185
PHSDF GPNSGSTACVA+IR N L VANAGDSRCVI
Sbjct: 568 PHSDFTGPNSGSTACVAVIRGNKLVVANAGDSRCVI 675
Score = 52.4 bits (124), Expect(2) = 3e-76
Identities = 25/31 (80%), Positives = 26/31 (83%)
Frame = +3
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQG 31
MG LS PKTEK S+DGEND LRYGLSSMQG
Sbjct: 120 MGIYLSSPKTEKASDDGENDRLRYGLSSMQG 212
>TC13146 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-like protein
(AT4G31860/F11C18_60), partial (20%)
Length = 462
Score = 159 bits (401), Expect = 9e-40
Identities = 78/96 (81%), Positives = 87/96 (90%)
Frame = +1
Query: 257 IVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSLAVG 316
IVD HDEDEF+V+ACDGIWDCLSSQ LVDFVRQ+L+LETKLS +CERVLDRCLAPS+AVG
Sbjct: 1 IVDRHDEDEFLVLACDGIWDCLSSQPLVDFVRQQLVLETKLSVICERVLDRCLAPSIAVG 180
Query: 317 DGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASE 352
G DNMTMILVQFK+P Q++A AQEQSSSNEQ A E
Sbjct: 181 VGFDNMTMILVQFKRPSQSTALAQEQSSSNEQYACE 288
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 127 bits (320), Expect = 2e-30
Identities = 99/327 (30%), Positives = 149/327 (45%), Gaps = 19/327 (5%)
Frame = +3
Query: 22 LRYGLSSMQGWRATMEDAHAAHLDVDSS----------TSFFGVYDGHGGKAVAKFCAKH 71
+R G + G R MED H D+ S ++F+GV+DGHGG A + K+
Sbjct: 492 IRSGSFADIGPRRYMEDEHVRIDDLSSHLGSLYNFPNPSAFYGVFDGHGGPEAAAYIRKN 671
Query: 72 LHQ---------QVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFN 122
+ + Q + ++ +V SL KAFL D +
Sbjct: 672 VIKFFFEDVSFPQTSEVDKVFLQEVENSLRKAFLLADSAL-------------------- 791
Query: 123 GIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSR 182
+DD + +SG+TA A I LL VANAGD R
Sbjct: 792 ------------------ADDCSVNT---------SSGTTALTAFIFGRLLMVANAGDCR 890
Query: 183 CVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFL 242
V+SR G+A ++S DH+P E+ R+ GG+I G +NG L++ RA+GD D K
Sbjct: 891 AVLSRKGEAIDMSEDHRPIYPSERRRVEDLGGYIEDGYLNGVLSVTRALGDWDMK---LP 1061
Query: 243 SAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCE 302
+ A P+ + L ++DEF++I CDGIWD +SSQ V VR+ L + + E
Sbjct: 1062KGAPSPLIAEPEFRQMVLTEDDEFLIIGCDGIWDVMSSQHAVSLVRKGL----RRHDDPE 1229
Query: 303 RVLDRCLAPSLAVGDGCDNMTMILVQF 329
R + +L + + DN+T+I+V F
Sbjct: 1230RCARDLVMEALRL-NTFDNLTVIIVCF 1307
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 112 bits (281), Expect = 7e-26
Identities = 92/329 (27%), Positives = 151/329 (44%), Gaps = 9/329 (2%)
Frame = +3
Query: 11 EKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHGGKAVAKFCAK 70
E+ +DG L YG S+ G R MEDA + + +F V+DGHGG VA+ C +
Sbjct: 366 ERKRDDGV---LPYGSVSVVGSRKEMEDAVSVETGCVTKCDYFAVFDGHGGAQVAEACRE 536
Query: 71 HLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIR 130
L++ V + E V + W E ++EG R
Sbjct: 537 RLYRLVAEEVERCGNGVE----------------EVDWEE-------------VMEGCFR 629
Query: 131 SPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQ 190
N + + + + A GSTA VA++ + +AN GD R V+ R G+
Sbjct: 630 ----NMDGEVAGNAALR----------TVGSTAVVAVVAAAEVVIANCGDCRAVLGRGGE 767
Query: 191 AYNLSRDHKPELVIEKERIYKAGGFI---HAGRINGSLNLARAIGDVDFKNNRFLSAEKQ 247
A +LS DHKP+ E RI +AGG + + R+ G L +R+IGD ++L +
Sbjct: 768 AVDLSSDHKPDRPDELMRIEEAGGKVINWNGQRVLGVLATSRSIGD------QYL---RP 920
Query: 248 VVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDR 307
V + P++ + +DEF+++A DG+WD +SS+ VR+ L ++ +C R
Sbjct: 921 YVISKPEVTVTKRSSKDEFLILASDGLWDVISSEMACQVVRK--CLNGQIRRICNENQSR 1094
Query: 308 C-----LAPSLAVGDGC-DNMTMILVQFK 330
L +A+ G DN ++I+++ +
Sbjct: 1095ASEAATLLAEIALAKGSRDNTSVIVIELR 1181
>AV425982
Length = 291
Score = 108 bits (269), Expect = 2e-24
Identities = 52/67 (77%), Positives = 57/67 (84%)
Frame = +3
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKTEK S+DGEND LRYGLSSMQGWRA+MEDAHAA +D STS+F VYDGHG
Sbjct: 90 MGIYLSSPKTEKASDDGENDRLRYGLSSMQGWRASMEDAHAALPCLDESTSYFAVYDGHG 269
Query: 61 GKAVAKF 67
GKAV+KF
Sbjct: 270 GKAVSKF 290
>TC14449 similar to UP|O24078 (O24078) Protein phosphatase 2C, partial (63%)
Length = 1000
Score = 100 bits (249), Expect = 4e-22
Identities = 63/163 (38%), Positives = 93/163 (56%), Gaps = 8/163 (4%)
Frame = +3
Query: 152 SDF--DGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPELVIEKERI 209
SDF + + GS A+IRN L V+NAGD R VISR G A L+ DH+P E+ERI
Sbjct: 195 SDFLKEDLHGGSCCVTALIRNGNLVVSNAGDCRAVISRGGVAEALTTDHRPSREDERERI 374
Query: 210 YKAGGFIH----AGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDE 265
GG++ RI GSL ++R+IGD R L KQ VTA P+ ++ + E +
Sbjct: 375 ETLGGYVDLCRGVWRIQGSLAVSRSIGD------RHL---KQWVTAEPETKVIKIEPEHD 527
Query: 266 FIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV--CERVLD 306
+++A DG+WD +S+Q+ VD R + K + C++++D
Sbjct: 528 LLILASDGLWDKVSNQEAVDLARPLCVGNNKPQSLLACKKLVD 656
Score = 35.4 bits (80), Expect = 0.015
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +2
Query: 47 DSSTSFFGVYDGHGGKAVAKFCAKHLHQ 74
++ +FFGV+DGHGG A+F A +L +
Sbjct: 35 ENKLAFFGVFDGHGGAKAAEFAANNLEE 118
>TC15379 UP|Q9ZPL8 (Q9ZPL8) Protein phosphatase type 2C, complete
Length = 1338
Score = 90.5 bits (223), Expect = 4e-19
Identities = 78/296 (26%), Positives = 124/296 (41%), Gaps = 9/296 (3%)
Frame = +1
Query: 21 NLRYGLSSMQGWRA-TMEDAHAAHLDV--DSSTSFFGVYDGHGGKAVAKFCAKHLHQQVL 77
N+ +G ++G MED A ++ F ++DGH G V + +L +L
Sbjct: 316 NITHGYHLVKGRSDHAMEDYVVAQFKTVDNNELGLFAIFDGHSGHNVPDYLQSNLFDNIL 495
Query: 78 KSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRSNDN 137
K ++ V ++ KA++ D + + G
Sbjct: 496 KEPDFWTKPV-EAVKKAYVDTDSTILEKSG------------------------------ 582
Query: 138 KDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRN-NLLFVANAGDSRCVISRNGQAYNLSR 196
E G GSTA AI+ N L VAN GDSR V+ +NG+A LS
Sbjct: 583 ---------ELG--------RGGSTAVTAILINCQKLVVANLGDSRAVLCKNGEAIPLSV 711
Query: 197 DHKPELVIEKERIYKAGGFIH-----AGRINGSLNLARAIGDVDFKNNRFLSAEKQVVTA 251
DH+P E E I GGF+ R++G L ++RA GD K + +++
Sbjct: 712 DHEP--ATESEDIRNRGGFVSNFPGDVPRVDGQLAVSRAFGDKSLKKH---------LSS 858
Query: 252 NPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERVLDR 307
P + + + D+ EFI++A DG+W +S+Q+ VD +R + + E L R
Sbjct: 859 EPHVTVELIDDDAEFIILASDGLWKVMSNQEAVDAIRNVKDARSAAKNLTEEALKR 1026
>TC9697 similar to GB|AAM63159.1|21554078|AY085949 protein phosphatase-2C
{Arabidopsis thaliana;}, partial (33%)
Length = 833
Score = 89.0 bits (219), Expect = 1e-18
Identities = 49/139 (35%), Positives = 82/139 (58%), Gaps = 1/139 (0%)
Frame = +2
Query: 190 QAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQVV 249
+A ++S DH+P + E+ R+ + GGFI G +NG L++ RA+GD DFK + A ++
Sbjct: 2 KAVDMSHDHRPSYLPERRRVEELGGFIDDGYLNGYLSVTRALGDWDFKLP--IGAASPLI 175
Query: 250 TANPDINIVDLHDEDEFIVIACDGIWDCLSSQ-QLVDFVRQELLLETKLSEVCERVLDRC 308
A PD+ ++ L ++DEF++I CDGIWD +SSQ V VR+ L + ++
Sbjct: 176 -AEPDVQLITLTEDDEFLIIGCDGIWDVMSSQVSPVSLVRRGLRRHDDPQQCARELVKEA 352
Query: 309 LAPSLAVGDGCDNMTMILV 327
L + + DN+T+I++
Sbjct: 353 LRLNTS-----DNLTVIVI 394
>TC16139 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-like protein
(AT4G31860/F11C18_60), partial (15%)
Length = 802
Score = 85.5 bits (210), Expect = 1e-17
Identities = 45/76 (59%), Positives = 56/76 (73%), Gaps = 2/76 (2%)
Frame = +1
Query: 277 CLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFKKPL--Q 334
C+SSQQLVDF++ +L E KLS VCE+V DRCLAP+ A G+GCDNMTMIL+QFK
Sbjct: 1 CMSSQQLVDFIQGQLKTENKLSVVCEKVFDRCLAPT-AGGEGCDNMTMILIQFKSSWNPD 177
Query: 335 TSAPAQEQSSSNEQDA 350
TSA Q +SS+ +A
Sbjct: 178 TSAANQTESSAQPSEA 225
>TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-like protein,
partial (36%)
Length = 917
Score = 77.4 bits (189), Expect = 3e-15
Identities = 57/181 (31%), Positives = 92/181 (50%), Gaps = 15/181 (8%)
Frame = +1
Query: 167 IIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFI---HAGRING 223
+I ++ + V+N GDSR V+ R + LS DHKP E RI AGG + + R+ G
Sbjct: 16 LISSSHIIVSNCGDSRAVLCRGKEPLALSVDHKPNREDEYARIEAAGGKVIQWNGHRVFG 195
Query: 224 SLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQL 283
L ++R+IGD R+L K + P++ + EDE +++A DG+WD +++++
Sbjct: 196 VLAMSRSIGD------RYL---KPWIIPEPEVRFLPRAKEDECLILASDGLWDVMTNEEA 348
Query: 284 VDFVRQELLLETK------LSEVCERVLDRCLAPSLAVGDGC------DNMTMILVQFKK 331
D R+ +LL K SE E V A + + + DN+T+I+V K
Sbjct: 349 CDLARKRILLWYKKNGFELSSERGEGVDPAAQAAAEFLSNRALPKGSKDNITVIVVDLKP 528
Query: 332 P 332
P
Sbjct: 529 P 531
>TC14935 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-like protein,
partial (56%)
Length = 1089
Score = 69.3 bits (168), Expect = 9e-13
Identities = 51/160 (31%), Positives = 79/160 (48%), Gaps = 30/160 (18%)
Frame = +1
Query: 158 NSGSTACVAIIRNNLLFVANAGDSRCVISRNG------QAYNLSRDHKPELVIEKERIYK 211
N+GS V +I LFVANAGDSR V+ + A LS +H L +E + +
Sbjct: 88 NAGSCCLVGVIFQQTLFVANAGDSRVVLGKKVGNTDGVAAIQLSTEHNANLEAIREELRE 267
Query: 212 AGGFIHAG------------RINGSLNLARAIGDVDFKNNRF----LSAEKQ-------- 247
+H ++ G + ++R+IGDV K+ RF L+A+ +
Sbjct: 268 ----LHPNDPQIVVLKYGVWKVKGIIQVSRSIGDVYMKDARFNREPLAAKFRLPEPMNMP 435
Query: 248 VVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFV 287
++TANP I L D F++ A DG+W+ LS+++ VD V
Sbjct: 436 IMTANPTILSHSLQPNDLFLIFASDGLWEHLSNEKAVDIV 555
>TC14934 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-like protein,
partial (57%)
Length = 932
Score = 66.2 bits (160), Expect = 8e-12
Identities = 46/154 (29%), Positives = 70/154 (44%), Gaps = 26/154 (16%)
Frame = +1
Query: 160 GSTACVAIIRNNLLFVANAGDSRCVISRNG------QAYNLSRDHK-------------- 199
G+ V +I LFVAN GDSRCV+ + A LS +H
Sbjct: 10 GTCCLVGVIFQQTLFVANLGDSRCVLGKKVGNTGGVAAIQLSTEHNANLEDIRQELKELH 189
Query: 200 ---PELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSAEKQ---VVTANP 253
P++V+ K +++ G I R G + A + + N +F E +++ANP
Sbjct: 190 PDDPQIVVLKHGVWRVKGIIQVSRSIGDSYMKHAQFNREPINPKFRLPEPMNMPIMSANP 369
Query: 254 DINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFV 287
I L D F++ A DG+W+ LS++Q VD V
Sbjct: 370 TILSHPLQPSDSFLIFASDGLWEHLSNEQAVDIV 471
>TC10673 homologue to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, partial
(48%)
Length = 669
Score = 56.2 bits (134), Expect = 8e-09
Identities = 44/173 (25%), Positives = 73/173 (41%), Gaps = 3/173 (1%)
Frame = +3
Query: 18 ENDNLRYGLSSMQGWRATMEDAHAAHLD--VDSSTSFFGVYDGHGGKAVAKFCAKHLHQQ 75
+N YG +S G R++MED + +D FGV+DGHGG A++ ++L
Sbjct: 294 QNGKFSYGYASSPGKRSSMEDFYETRIDGVEGEVVGLFGVFDGHGGARAAEYVKQNLFSN 473
Query: 76 VLKSEEYIAGDVGTSLTKAFLRMD-EMMRGQRGWRELAVLGDKVKGFNGIIEGLIRSPRS 134
++ ++I+ D +++ A+ D E ++ + +
Sbjct: 474 LISHPKFIS-DTKSAIADAYNHTDSEFLKSE----------------------------N 566
Query: 135 NDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISR 187
N N+D +GSTA AI+ + L VAN GDS+ VI R
Sbjct: 567 NQNRD-------------------AGSTASTAILVGDRLLVANVGDSKAVICR 668
>TC9837 similar to UP|Q9M3V0 (Q9M3V0) Protein phosphatase 2C (PP2C) ,
partial (43%)
Length = 727
Score = 52.0 bits (123), Expect = 2e-07
Identities = 29/103 (28%), Positives = 52/103 (50%), Gaps = 19/103 (18%)
Frame = +3
Query: 220 RINGSLNLARAIGDVDFKNNRFLSA------------EKQVVTANPDINIVDLHDEDEFI 267
R+ G + ++R+IGDV K F ++ ++++ P I + L D+FI
Sbjct: 15 RVKGIIQISRSIGDVYLKKAEFNREPLYAKFRLREPFKRPILSSEPSITVHQLQPHDQFI 194
Query: 268 VIACDGIWDCLSSQQLVDFVRQ-------ELLLETKLSEVCER 303
+ A DG+W+ LS+Q+ VD V+ L++T L E ++
Sbjct: 195 IFASDGLWEHLSNQEAVDIVQNSPRSGSARRLVKTALQEAAKK 323
>TC12089 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partial (25%)
Length = 631
Score = 51.6 bits (122), Expect = 2e-07
Identities = 28/87 (32%), Positives = 44/87 (50%)
Frame = +3
Query: 245 EKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERV 304
+K V A PDI +DL D + FI++ CDG+W VDFV++ L ++ V R+
Sbjct: 21 QKVGVIATPDIQSLDLTDREHFIILGCDGLWGVFGPSDAVDFVQKLLNEGLPVTTVSRRL 200
Query: 305 LDRCLAPSLAVGDGCDNMTMILVQFKK 331
+ + DN T I++ FK+
Sbjct: 201 VREAIRERQCQ----DNCTAIVIVFKR 269
>TC11286 homologue to GB|AAO63448.1|28951049|BT005384 At4g03415 {Arabidopsis
thaliana;}, partial (46%)
Length = 996
Score = 51.2 bits (121), Expect = 3e-07
Identities = 37/110 (33%), Positives = 53/110 (47%), Gaps = 13/110 (11%)
Frame = +3
Query: 191 AYNLSRDHKPELVIEKERIYKAGGFIHAGRINGS-------------LNLARAIGDVDFK 237
A L+ D KP+L E ERI + G + A + L +ARA GD K
Sbjct: 21 AVQLTVDLKPDLPREAERIKRCKGRVFALQDEPEVPRVWLPFDDAPGLAMARAFGDFCLK 200
Query: 238 NNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFV 287
+S P+ + L D D+FIVIA DG+WD LS++++V+ V
Sbjct: 201 EYGVISI--------PEFSHRQLTDRDQFIVIASDGVWDVLSNEEVVEIV 326
>CN825589
Length = 544
Score = 49.3 bits (116), Expect = 1e-06
Identities = 27/79 (34%), Positives = 43/79 (54%)
Frame = +3
Query: 220 RINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLS 279
R+ G L ++RAIGD + K V + P++ + D DEDE +++A DG+WD +S
Sbjct: 42 RVLGVLAMSRAIGD---------NYLKPYVISEPEVTVTDRTDEDECLILASDGLWDVVS 194
Query: 280 SQQLVDFVRQELLLETKLS 298
+ VR L + +LS
Sbjct: 195 NDTACGVVRMCLKAQKQLS 251
>BP054371
Length = 567
Score = 46.6 bits (109), Expect = 6e-06
Identities = 20/70 (28%), Positives = 42/70 (59%)
Frame = -2
Query: 220 RINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLS 279
R+NG L ++RA GD + K++ + ++PDI ++ + EF+++A DG+W ++
Sbjct: 563 RVNGQLAVSRAFGDQNLKSH---------LRSDPDIRHDEIDSDTEFLILASDGLWKVMA 411
Query: 280 SQQLVDFVRQ 289
+ + VD ++
Sbjct: 410 NPEAVDIAKK 381
>TC20055
Length = 556
Score = 45.1 bits (105), Expect = 2e-05
Identities = 29/82 (35%), Positives = 40/82 (48%), Gaps = 3/82 (3%)
Frame = +2
Query: 22 LRYGLSSMQGW-RATMEDAHAAHLDV--DSSTSFFGVYDGHGGKAVAKFCAKHLHQQVLK 78
++YG S ++G MED H A F +YDGH G V + KHL +LK
Sbjct: 245 VKYGFSLVKGKANHPMEDFHVAKFVEFKGQELGLFAIYDGHLGDTVPAYLQKHLFPNILK 424
Query: 79 SEEYIAGDVGTSLTKAFLRMDE 100
E++ D S+ KA+L DE
Sbjct: 425 EEDF-WNDPFMSICKAYLSTDE 487
>TC13433 similar to UP|Q8VZD9 (Q8VZD9) AT5g53140/MFH8_8, partial (20%)
Length = 557
Score = 44.3 bits (103), Expect = 3e-05
Identities = 23/68 (33%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Frame = +3
Query: 18 ENDNLRYGLSSMQGWRATMEDAH---AAHLDVDSSTSFFGVYDGHGGKAVAKFCAKHLHQ 74
E+ L G SS +G R TMED + +++D S FG++DGHGG A + HL
Sbjct: 336 EDKRLSCGYSSFRGKRVTMEDFYDIKTSNID-GHSVCLFGIFDGHGGSRAAXYLKXHLFD 512
Query: 75 QVLKSEEY 82
++ + +
Sbjct: 513 NLMNIQSF 536
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,295,806
Number of Sequences: 28460
Number of extensions: 64350
Number of successful extensions: 356
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 337
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 345
length of query: 358
length of database: 4,897,600
effective HSP length: 91
effective length of query: 267
effective length of database: 2,307,740
effective search space: 616166580
effective search space used: 616166580
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC143339.7


