

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.11 + phase: 0
(678 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
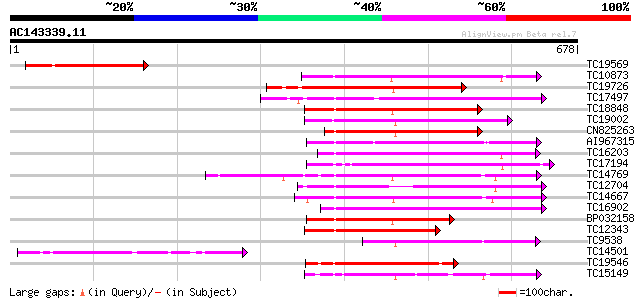
Score E
Sequences producing significant alignments: (bits) Value
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 213 1e-55
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 202 2e-52
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 195 2e-50
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 189 2e-48
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 187 5e-48
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 181 3e-46
CN825263 177 6e-45
AI967315 174 5e-44
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 169 1e-42
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 165 2e-41
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 162 2e-40
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 160 6e-40
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 159 1e-39
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 156 1e-38
BP032158 155 2e-38
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 144 5e-35
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 143 8e-35
TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA),... 142 1e-34
TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine k... 142 2e-34
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 139 1e-33
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 213 bits (541), Expect = 1e-55
Identities = 104/148 (70%), Positives = 119/148 (80%)
Frame = +3
Query: 19 ILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFRV 78
+L+L LL + P T+SL FNITNFDD T A ++Y+GDGKSTNGSIDLNKVSY FRV
Sbjct: 48 LLLLLLLLTTTTTFPTTHSLSFNITNFDDDTTA--IAYEGDGKSTNGSIDLNKVSYFFRV 221
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGG 138
GRA +QPLHL+D T TLT FTTRF+FTI +N T+YGDGF FYLAPLGYQIPPNSAGG
Sbjct: 222 GRAISAQPLHLYDPTTKTLTDFTTRFTFTISAINKTSYGDGFAFYLAPLGYQIPPNSAGG 401
Query: 139 VYGLFNATTNSNFVMNYVVGVEFDTFVG 166
+GLFNATTN+N N+VV VEFDTFVG
Sbjct: 402 TFGLFNATTNNNLPQNHVVAVEFDTFVG 485
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 202 bits (513), Expect = 2e-52
Identities = 120/296 (40%), Positives = 170/296 (56%), Gaps = 10/296 (3%)
Frame = +1
Query: 350 TIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVF 409
T F + EL AT F + ++G GG+G+VYKG L+ G+ VAVK++ D + F
Sbjct: 400 TAAASFGFRELADATRNFKEANLIGEGGFGKVYKGRLT-TGEAVAVKQLSHDGRQGFQEF 576
Query: 410 INEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG---DKKSLAWEVRY 466
+ EV ++S L H NLV+ IG+C + D+ LLV+EYMP GSL+ HLF DK+ L W R
Sbjct: 577 VMEVLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRM 756
Query: 467 KVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RT 525
KVA+G A L YLH A+ V++RD+KSAN+LLD +F+ KL DFG+AKL T T
Sbjct: 757 KVAVGAARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVST 936
Query: 526 DVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDG--DFHVPLMNWVWGL 583
V+GTYGY APEY G+ + +SD+YSFG+V LEL TGRR L++W
Sbjct: 937 RVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPY 1116
Query: 584 YVE----GNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
+ + G+++ D L F + + + C K RP +++ L+
Sbjct: 1117FSDRRRFGHMV---DPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIVVALE 1275
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 195 bits (495), Expect = 2e-50
Identities = 104/246 (42%), Positives = 159/246 (64%), Gaps = 7/246 (2%)
Frame = +3
Query: 308 ILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGF 367
++V +I ++ W+ +K+KR +EG + + K PR F Y EL ATN F
Sbjct: 12 VIVAVIGGLLWWIRLKKKR---EEGSNSQILGTLKSLP----GTPREFNYVELKKATNNF 170
Query: 368 DDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQF 427
D+ LG+GGYG VY+G L VAVK D S F++E+ II+RL H++LV+
Sbjct: 171 DEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELIIINRLRHKHLVRL 350
Query: 428 IGWCHEQDELLLVFEYMPNGSLDTHLFGDK-----KSLAWEVRYKVALGVANALRYLHDD 482
GWCH+ LLLV++YMPNGSLD+H+F ++ L+W +RYK+ GVA+AL YLH++
Sbjct: 351 QGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKIISGVASALNYLHNE 530
Query: 483 AEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLV--DPMLRTQRTDVVGTYGYLAPEYIN 540
+Q V+HRD+K++N++LD++F+ KLGDFG+A+ + + + T+ V GT GY+APE +
Sbjct: 531 YDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTELEGVHGTMGYIAPECFH 710
Query: 541 GGRASK 546
G+A++
Sbjct: 711 TGKATR 728
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 189 bits (479), Expect = 2e-48
Identities = 119/352 (33%), Positives = 189/352 (52%), Gaps = 9/352 (2%)
Frame = +1
Query: 300 VVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKF-----DLDKATIPRR 354
+V +I VI + ++ + I ++K K DEG E GI K D+D ATI
Sbjct: 1471 LVVIIAFVIFITILGLAISTCIQRKKNKRGDEG--EIGIINHWKDKRGDEDIDLATI--- 1635
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F++S + +ATN F LG GG+G VYKG L+ G+ +AVKR+ F NE++
Sbjct: 1636 FDFSTISSATNHFSLSNKLGEGGFGPVYKGLLAN-GQEIAVKRLSNTSGQGMEEFKNEIK 1812
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLA-WEVRYKVALGVA 473
+I+RL HRNLV+ G QDE N + L + L W R ++ G+A
Sbjct: 1813 LIARLQHRNLVKLFGCSVHQDE-----NSHANKKMKILLDSTRSKLVDWNKRLQIIDGIA 1977
Query: 474 NALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKL-VDPMLRTQRTDVVGTYG 532
L YLH D+ ++HRD+K++N+LLD + + K+ DFG+A++ + + + V+GTYG
Sbjct: 1978 RGLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYG 2157
Query: 533 YLAPEYINGGRASKESDMYSFGIVALELATGRRF--FQDGDFHVPLMNWVWGLYVEGNLM 590
Y+ PEY G S +SD++SFG++ LE+ +G++ F D H+ L++ W L++E +
Sbjct: 2158 YMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERPL 2337
Query: 591 CAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPE 642
DE L+ +E+ + V L C + RP ++ +L E LP+
Sbjct: 2338 ELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKELPK 2493
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 187 bits (475), Expect = 5e-48
Identities = 100/216 (46%), Positives = 136/216 (62%), Gaps = 3/216 (1%)
Frame = +2
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R F Y EL AAT GF DD LG GG+G VY G S G +AVK++ A +E F E
Sbjct: 260 RIFTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSD-GLQIAVKKLKAMNSKAEMEFAVE 436
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD---KKSLAWEVRYKVA 469
V ++ R+ H+NL+ G+C D+ L+V++YMPN SL +HL G + L W+ R K+A
Sbjct: 437 VEVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIA 616
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVG 529
+G A + YLH + ++HRDIK++NVLL++DF + DFG AKL+ + T V G
Sbjct: 617 IGSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMTTRVKG 796
Query: 530 TYGYLAPEYINGGRASKESDMYSFGIVALELATGRR 565
T GYLAPEY G+ S+ D+YSFGI+ LEL TGR+
Sbjct: 797 TLGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRK 904
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 181 bits (459), Expect = 3e-46
Identities = 106/254 (41%), Positives = 149/254 (57%), Gaps = 5/254 (1%)
Frame = +1
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R F EL +ATN F+ D LG GG+G VY G L + G +AVKR+ ++ F E
Sbjct: 22 RVFSLKELHSATNNFNYDNKLGEGGFGSVYWGQL-WDGSQIAVKRLKVWSNKADMEFAVE 198
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVA 469
V I++R+ H+NL+ G+C E E L+V++YMPN SL +HL G S L W R +A
Sbjct: 199 VEILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIA 378
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVG 529
+G A + YLH A ++HRDIK++NVLLD+DF ++ DFG AKL+ T V G
Sbjct: 379 IGSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKG 558
Query: 530 TYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHV--PLMNWVWGLYVEG 587
T GYLAPEY G+A++ D++SFGI+ LELA+G++ + V + +W L
Sbjct: 559 TLGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAK 738
Query: 588 NLMCAADEKLNMEF 601
AD +LN E+
Sbjct: 739 KFTEFADPRLNGEY 780
>CN825263
Length = 663
Score = 177 bits (448), Expect = 6e-45
Identities = 93/193 (48%), Positives = 129/193 (66%), Gaps = 4/193 (2%)
Frame = +1
Query: 377 GYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDE 436
G+G VYKG L+ G+ VAVK + D + R F+ EV ++SRL HRNLV+ IG C E+
Sbjct: 1 GFGLVYKGILND-GRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQT 177
Query: 437 LLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIK 493
L++E +PNGS+++HL G K L W R K+ALG A L YLH+D+ CV+HRD K
Sbjct: 178 RCLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFK 357
Query: 494 SANVLLDTDFSTKLGDFGMAK-LVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYS 552
S+N+LL+ DF+ K+ DFG+A+ +D + T V+GT+GYLAPEY G +SD+YS
Sbjct: 358 SSNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYS 537
Query: 553 FGIVALELATGRR 565
+G+V LEL TG +
Sbjct: 538 YGVVLLELLTGTK 576
>AI967315
Length = 1308
Score = 174 bits (440), Expect = 5e-44
Identities = 100/285 (35%), Positives = 158/285 (55%), Gaps = 4/285 (1%)
Frame = +1
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADF--ENSERVFINE 412
F Y EL ATNGF + M+G+GGY +VYKG L G +AVKR+ E E+ F+ E
Sbjct: 112 FSYEELFHATNGFSSENMVGKGGYAEVYKGRLES-GDEIAVKRLTRTCRDERKEKEFLTE 288
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALG 471
+ I + H N++ +G C + L LVFE GS+ + + +K + L W+ RYK+ LG
Sbjct: 289 IGTIGHVCHSNVMPLLGCCIDNG-LYLVFELSTVGSVASLIHDEKMAPLDWKTRYKIVLG 465
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAK-LVDPMLRTQRTDVVGT 530
A L YLH ++ ++HRDIK++N+LL DF ++ DFG+AK L + GT
Sbjct: 466 TARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGT 645
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLM 590
+G+LAPEY G +++D+++FG+ LE+ +GR+ DG H L W + + +
Sbjct: 646 FGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPV-DGS-HQSLHTWAKPILSKWEIE 819
Query: 591 CAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
D +L +DV++ + C ++ RP EV++V++
Sbjct: 820 KLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVME 954
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 169 bits (428), Expect = 1e-42
Identities = 95/276 (34%), Positives = 155/276 (55%), Gaps = 10/276 (3%)
Frame = +2
Query: 369 DDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFEN-SERVFINEVRIISRLIHRNLVQF 427
++ ++G+GG G VY+G++ G VA+KR+ ++ F E+ + ++ HRN+++
Sbjct: 2198 EENIIGKGGAGIVYRGSMPN-GTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRL 2374
Query: 428 IGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANALRYLHDDAEQC 486
+G+ +D LL++EYMPNGSL L G K L WE+RYK+A+ A L Y+H D
Sbjct: 2375 LGYVSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAVEAARGLCYMHHDCSPL 2554
Query: 487 VLHRDIKSANVLLDTDFSTKLGDFGMAK-LVDPMLRTQRTDVVGTYGYLAPEYINGGRAS 545
++HRD+KS N+LLD DF + DFG+AK L DP + + G+YGY+APEY +
Sbjct: 2555 IIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVD 2734
Query: 546 KESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVE-------GNLMCAADEKLN 598
++SD+YSFG+V LEL GR+ + V ++ WV E ++ D +L+
Sbjct: 2735 EKSDVYSFGVVLLELIIGRKPVGEFGDGVDIVGWVNKTMSELSQPSDTALVLAVVDPRLS 2914
Query: 599 MEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
+ ++ + + + + C RP EV+ +L
Sbjct: 2915 -GYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHML 3019
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 165 bits (418), Expect = 2e-41
Identities = 103/306 (33%), Positives = 159/306 (51%), Gaps = 9/306 (2%)
Frame = +1
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F Y EL ATN F D +G+GG+G VY L GK A+K++ D + S F+ E++
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELR--GKKTAIKKM--DVQASTE-FLCELK 1215
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD-KKSLAWEVRYKVALGVA 473
+++ + H NLV+ IG+C E L LV+E++ NG+L +L G K+ L W R ++AL A
Sbjct: 1216 VLTHVHHLNLVRLIGYCVE-GSLFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDAA 1392
Query: 474 NALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGY 533
L Y+H+ +HRD+KSAN+L+D + K+ DFG+ KL++ T +T +VGT+GY
Sbjct: 1393 RGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTFGY 1572
Query: 534 LAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEG------ 587
+ PEY G S + D+Y+FG+V EL + + V + L+ E
Sbjct: 1573 MPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKSDP 1752
Query: 588 --NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPL 645
L D +L + + + + +G CT N RP ++ L M L L
Sbjct: 1753 CDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLVVAL---MTLSSLTE 1923
Query: 646 DMHDRA 651
D D +
Sbjct: 1924 DCDDES 1941
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 162 bits (409), Expect = 2e-40
Identities = 127/419 (30%), Positives = 200/419 (47%), Gaps = 18/419 (4%)
Frame = +3
Query: 235 ISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSL 294
+SY DLM KLPE + + L N S E +N+ S T +
Sbjct: 1755 LSYN-DLMGKLPESI-VKLPHLKSLYFGCNEHMSPEDPANMNSSLINTDYGRCKGKESRF 1928
Query: 295 KTVIVVVAVIVPVILVFLIASIVGWVIVKRK---------RKNCDEGLDEYGIPFPKKFD 345
VIV+ A+ +L+ L ++ ++K +K E + +P F
Sbjct: 1929 GQVIVIGAITCGSLLITLAFGVLFVCRYRQKLIPWEGFAGKKYPMETNIIFSLPSKDDFF 2108
Query: 346 LDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENS 405
+ +I + F + AT + ++G GG+G VY+G L+ G+ VAVK +
Sbjct: 2109 IKSVSI-QAFTLEYIEVATERYKT--LIGEGGFGSVYRGTLND-GQEVAVKVRSSTSTQG 2276
Query: 406 ERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD---KKSLAW 462
R F NE+ ++S + H NLV +G+C+E D+ +LV+ +M NGSL L+G+ +K L W
Sbjct: 2277 TREFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDW 2456
Query: 463 EVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRT 522
R +ALG A L YLH + V+HRDIKS+N+LLD K+ DFG +K +
Sbjct: 2457 PTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDS 2636
Query: 523 Q-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNW-- 579
+V GT GYL PEY + S++SD++SFG+V LE+ +GR + + P W
Sbjct: 2637 YVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGR---EPLNIKRPRTEWSL 2807
Query: 580 -VWGL-YVEGNLMC-AADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
W Y+ G+ + D + + M ++ V L C RP +++ L+
Sbjct: 2808 VEWATPYIRGSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRELE 2984
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 160 bits (405), Expect = 6e-40
Identities = 102/325 (31%), Positives = 165/325 (50%), Gaps = 27/325 (8%)
Frame = +1
Query: 345 DLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFEN 404
D+D ATI F++S + + TN F + LG GG+G VYKG L+ G+ +AVKR+
Sbjct: 1396 DIDLATI---FDFSTISSTTNHFSESNKLGEGGFGPVYKGVLAN-GQEIAVKRLSNTSGQ 1563
Query: 405 SERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEV 464
F NEV++I+RL HRNLV+ +G DE+LL++E+M N SLD +F
Sbjct: 1564 GMEEFKNEVKLIARLQHRNLVKLLGCSIHHDEMLLIYEFMHNRSLDYFIF---------- 1713
Query: 465 RYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKL-VDPMLRTQ 523
D+ ++HRD+K++N+LLD++ + K+ DFG+A++ + +
Sbjct: 1714 -----------------DSRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAK 1842
Query: 524 RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRF--FQDGDFHVPLMNW-- 579
V+GTYGY++PEY G S +SD++SFG++ LE+ +G++ F D H L++
Sbjct: 1843 TKRVMGTYGYMSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSS 2022
Query: 580 ----------------------VWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCT 617
W L++E + DE L+ +E+ + + L C
Sbjct: 2023 NFAVFLIKALRICMFENVKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLCV 2202
Query: 618 HSNDKERPKAYEVIKVLQLEMALPE 642
+ RP V+ +L E LP+
Sbjct: 2203 QQRPEYRPDMLSVVLMLNGEKELPK 2277
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 159 bits (403), Expect = 1e-39
Identities = 106/319 (33%), Positives = 158/319 (49%), Gaps = 18/319 (5%)
Frame = +2
Query: 341 PKKFDLDKATIPRR---FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKR 397
P K + KA P EL T+ F ++G G YG+VY L+ G VAVK+
Sbjct: 272 PVKHETQKAPPPIEVPALSLDELKEKTDNFGSKALIGEGSYGRVYYATLND-GNAVAVKK 448
Query: 398 IFADFE-NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD 456
+ E + F+ +V ++SRL + N V+ G+C E + +L +E+ GSL L G
Sbjct: 449 LDVSSEPETNNEFLTQVSMVSRLKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGR 628
Query: 457 K--------KSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLG 508
K +L W R ++A+ A L YLH+ + ++HRDI+S+NVL+ D+ K+
Sbjct: 629 KGVQGAQPGPTLDWIQRVRIAVDAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIA 808
Query: 509 DFGMAKLVDPML-RTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF 567
DF ++ M R T V+GT+GY APEY G+ +++SD+YSFG+V LEL TGR+
Sbjct: 809 DFNLSNQAPDMAARLHSTRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRK-- 982
Query: 568 QDGDFHVP-----LMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDK 622
D +P L+ W E + D KL E+ + L V C +
Sbjct: 983 -PVDHTMPRGQQSLVTWATPRLSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAE 1159
Query: 623 ERPKAYEVIKVLQLEMALP 641
RP V+K LQ + P
Sbjct: 1160FRPNMSIVVKALQPLLKTP 1216
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 156 bits (394), Expect = 1e-38
Identities = 94/275 (34%), Positives = 144/275 (52%), Gaps = 4/275 (1%)
Frame = +3
Query: 372 MLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWC 431
+LG GG+G+VY G + G VA+KR E F E+ ++S+L HR+LV IG+C
Sbjct: 12 LLGVGGFGKVYYGEVDG-GTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYC 188
Query: 432 HEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANALRYLHDDAEQCVLHR 490
E E++LV+++M G+L HL+ +K L W+ R ++ +G A L YLH A+ ++HR
Sbjct: 189 EENTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHR 368
Query: 491 DIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV-GTYGYLAPEYINGGRASKESD 549
D+K+ N+LLD + K+ DFG++K + T + VV G++GYL PEY + + +SD
Sbjct: 369 DVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSD 548
Query: 550 MYSFGIVALELATGRRFFQD--GDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMK 607
+YSFG+V E+ R V L W Y +G L D L + K
Sbjct: 549 VYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFK 728
Query: 608 SLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPE 642
+ C ERP +V+ L+ + L E
Sbjct: 729 KFAETAMKCVSDQGIERPSMGDVLWNLEFALQLQE 833
>BP032158
Length = 555
Score = 155 bits (393), Expect = 2e-38
Identities = 82/181 (45%), Positives = 113/181 (62%), Gaps = 3/181 (1%)
Frame = +2
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F ++ AATN FD +G GG+G VYKG LS G ++AVK++ + + R FINE+
Sbjct: 14 FSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSE-GDVIAVKQLSSKSKQGNREFINEIG 190
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD---KKSLAWEVRYKVALG 471
+IS L H NLV+ G C E ++LLLV+EYM N SL LFG+ K +L W R K+ +G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTY 531
+A L YLH+++ ++HRDIK+ NVLLD D K+ DFG+AKL + T + GT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAGTI 550
Query: 532 G 532
G
Sbjct: 551 G 553
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 144 bits (363), Expect = 5e-35
Identities = 72/165 (43%), Positives = 104/165 (62%), Gaps = 2/165 (1%)
Frame = +3
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
+ F Y LVAAT F LG GG+G V+KG L+ G+ +AVK++ FINE
Sbjct: 201 KTFSYETLVAATKNFHAVNKLGEGGFGPVFKGKLND-GREIAVKKLSRRSNQGRTQFINE 377
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS--LAWEVRYKVAL 470
++++R+ HRN+V G+C E LLV+EY+P SLD LF +K L W+ R+ +
Sbjct: 378 AKLLTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKEQLDWKRRFDIIS 557
Query: 471 GVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKL 515
GVA L YLH+D+ C++HRDIK+AN+LLD + K+ DFG+A++
Sbjct: 558 GVARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARI 692
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 143 bits (361), Expect = 8e-35
Identities = 84/226 (37%), Positives = 125/226 (55%), Gaps = 14/226 (6%)
Frame = +2
Query: 423 NLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS------------LAWEVRYKVAL 470
N+V+ + + +LLV+EY+ N SLD L KS L W R K+A+
Sbjct: 5 NIVRLLCCISNEASMLLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAI 184
Query: 471 GVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAK-LVDPMLRTQRTDVVG 529
G A L Y+H D ++HRD+K++N+LLD F+ K+ DFG+A+ L+ P + V+G
Sbjct: 185 GAAQGLSYMHHDCSPPIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMSTVIG 364
Query: 530 TYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNL 589
T+GY+APEY+ R S++ D+YSFG+V LEL TG+ GD H L W W + G+
Sbjct: 365 TFGYIAPEYVQTTRISEKVDVYSFGVVLLELTTGKE-ANYGDQHSSLAEWAWRHILIGSN 541
Query: 590 MCAADEKLNMEFD-VSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
+ K ME + EM S+ +G+ CT + RP EV+ +L
Sbjct: 542 VXDLLXKDVMEASYIDEMCSVFKLGVMCTATLPATRPSMKEVLPIL 679
>TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA), partial
(86%)
Length = 1066
Score = 142 bits (359), Expect = 1e-34
Identities = 103/278 (37%), Positives = 142/278 (51%), Gaps = 3/278 (1%)
Frame = +1
Query: 10 SNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKS-TNGSID 68
SN Q+ + LF LL + L S FN T F D ++ QGD K +GS+
Sbjct: 58 SNASTQNFLLACLFFLLAINVNLAAAIS--FNFTEFTDD---GSLIIQGDAKIWADGSLA 222
Query: 69 L-NKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPL 127
L S F RA Y+ P+ +WD T + SF T FSF I D DG VF+LAP
Sbjct: 223 LPTDPSVGFTTSRALYATPVPIWDSTTGNVASFVTSFSFIIKDFEDYNPADGLVFFLAPF 402
Query: 128 GYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDNALTSVAF 187
G +IP S GG +G+ N N +V VEFDTF+ P D +H+GID N+L S+
Sbjct: 403 GTEIPKESTGGRFGIINGKD----AFNQIVAVEFDTFINPWDSSPRHIGIDVNSLISLKT 570
Query: 188 GKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPE 247
+ +K G + V I Y+S K L V + +G S+IS +IDL LPE
Sbjct: 571 VPW--NKVAGSLEKVTIIYDSQTKTLSVL------VIHENGQI--STISQEIDLKVVLPE 720
Query: 248 FVNIGFSA-STGLSTESNVIHSWEFSSNLEDSNSTTSL 284
V++GFSA +T E + I+SW F+S L + +T ++
Sbjct: 721 EVSVGFSATTTSGGRERHDIYSWSFTSTLNTNGATENI 834
>TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine kinase ,
partial (46%)
Length = 731
Score = 142 bits (358), Expect = 2e-34
Identities = 68/183 (37%), Positives = 117/183 (63%)
Frame = +1
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
++ Y ++ AT F +LG+G +G VYK + G++VAVK + + + E F EV
Sbjct: 196 KYSYKDIQKATQNFTT--ILGQGSFGTVYKATMP-TGEVVAVKVLAPNSKQGEHEFQTEV 366
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVA 473
++ RL HRNLV +G+C ++ + +LV+++M NGSL L+G++K L+W+ R ++A+ ++
Sbjct: 367 HLLGRLHHRNLVNLVGFCVDKGQRILVYQFMSNGSLANLLYGEEKELSWDERLQIAMDIS 546
Query: 474 NALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGY 533
+ + YLH+ A V+HRD+KSAN+LLD + DFG++K + + + + + GTYGY
Sbjct: 547 HGIEYLHEGAVPPVIHRDLKSANILLDDSMRAMVADFGLSK--EEIFDGRNSGLKGTYGY 720
Query: 534 LAP 536
+ P
Sbjct: 721 MDP 729
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 139 bits (350), Expect = 1e-33
Identities = 100/293 (34%), Positives = 150/293 (51%), Gaps = 10/293 (3%)
Frame = +3
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R F+ +L+ A+ +LG+G +G YK L G +VAVKR+ D SE+ F ++
Sbjct: 249 RGFDLEDLLRASA-----EVLGKGTFGTAYKAVLE-TGLVVAVKRL-KDVTISEKEFKDK 407
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS----LAWEVRYKV 468
+ + + H++LV + +DE LLV++YMP GSL L G+K + L WE+R +
Sbjct: 408 IETVGAMDHQSLVPLRAYYFSRDEKLLVYDYMPMGSLSALLHGNKGAGRTPLNWEIRSGI 587
Query: 469 ALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV 528
ALG A + YLH V H +IK++N+LL + K+ DFG+A LV P R
Sbjct: 588 ALGAARGIEYLHSQGPN-VSHGNIKASNILLTKSYEAKVSDFGLAHLVGPSSTPNR---- 752
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRR-----FFQDGDFHVPLMNWVWGL 583
GY APE + + S+++D+YSFG++ LEL TG+ ++G V L WV L
Sbjct: 753 -VAGYRAPEVTDPRKVSQKADVYSFGVLLLELLTGKAPTHALLNEEG---VDLPRWVQSL 920
Query: 584 YVEGNLMCAAD-EKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
E D E L + EM LL + + C RP EV + ++
Sbjct: 921 VREEWTSEVFDLELLRYQNVEEEMVQLLQLAVDCAAPYPDRRPSIAEVTRSIE 1079
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,728,512
Number of Sequences: 28460
Number of extensions: 141019
Number of successful extensions: 1254
Number of sequences better than 10.0: 300
Number of HSP's better than 10.0 without gapping: 1067
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1074
length of query: 678
length of database: 4,897,600
effective HSP length: 96
effective length of query: 582
effective length of database: 2,165,440
effective search space: 1260286080
effective search space used: 1260286080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC143339.11


