

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
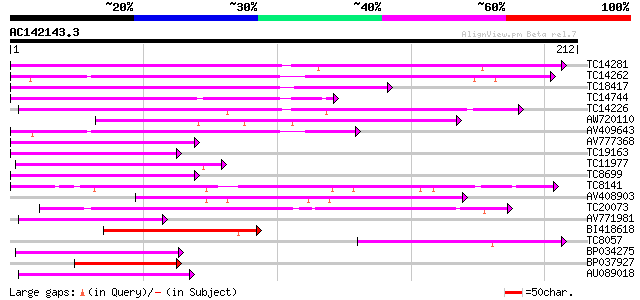
Score E
Sequences producing significant alignments: (bits) Value
TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygen... 128 8e-31
TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete 91 1e-19
TC18417 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, pa... 74 1e-14
TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment)... 64 1e-11
TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete 60 4e-10
AW720110 59 5e-10
AV409643 56 4e-09
AV777368 54 3e-08
TC19163 similar to UP|Q84J65 (Q84J65) Gray pubescence flavonoid ... 53 3e-08
TC11977 weakly similar to UP|C822_SOYBN (O81972) Cytochrome P450... 53 5e-08
TC8699 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, par... 49 8e-07
TC8141 weakly similar to UP|C79B_ARATH (O81346) Cytochrome P450 ... 48 1e-06
AV408903 48 1e-06
TC20073 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 mon... 44 2e-05
AV771981 44 3e-05
BI418618 42 1e-04
TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, par... 40 2e-04
BP034275 40 4e-04
BP037927 40 4e-04
AU089018 39 5e-04
>TC14281 similar to UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1,
partial (97%)
Length = 1768
Score = 128 bits (321), Expect = 8e-31
Identities = 74/214 (34%), Positives = 120/214 (55%), Gaps = 6/214 (2%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT + +F RP +S + Y S + PYG YWRF+KK+C+T+LLS L F+ +R
Sbjct: 324 LKTSEESFCNRPIMIASENLTYGASDYFFIPYGTYWRFLKKLCMTELLSGKTLEHFVSVR 503
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV--IQCL 118
E E++ LK++L S + + + TNN++ M+MG K E+ ++ +
Sbjct: 504 ENEIQAFLKTILEISKTGEAVVMRQELIRHTNNVISMMTMGK---KSEGSKDEIGELRKV 674
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTE--- 175
+RE + ++G+++G + DL GYGKK + + + D ++E LKEHEE TE
Sbjct: 675 IREIGELLGSFNLGDIIGFMRPLDLQGYGKKNKDMHRKLDGMMEKVLKEHEEARATEGAD 854
Query: 176 -DCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D + D+ DILL + + A+ +LTR+ KAF L
Sbjct: 855 SDRKKDLFDILLNLIEADGADNKLTRDSAKAFAL 956
>TC14262 UP|Q9MBE5 (Q9MBE5) Cytochrome P450, complete
Length = 1960
Score = 90.9 bits (224), Expect = 1e-19
Identities = 56/215 (26%), Positives = 115/215 (53%), Gaps = 11/215 (5%)
Frame = +1
Query: 1 MKTHDL-NFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ +FS R Q + Y S + P+ PYW+F++K+ + LL+++ + + +
Sbjct: 355 LQTHEATSFSTRFQTSAIRRLTYDNSVAMV-PFAPYWKFIRKVIMNDLLNATTVNKLRPL 531
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R QE+ K+LK++ + + ++ + TN+ + RM +G + E ++ +V
Sbjct: 532 RSQEIRKVLKAMAQSAESQKPLNVTEELLKWTNSTISRMMLG---------EAEHVKDIV 684
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI------N 173
RE + + + S+ + + PL K + Y K++ +I +FD ++E +K+ +E I N
Sbjct: 685 REVLKIFGEYSLTDFIWPLKKLRVGQYEKRIDEIFNKFDPVIEKVIKKRQEIIKRRKERN 864
Query: 174 TEDCQGD----MMDILLQVYRNENAEVRLTRNDIK 204
E +G+ +D LL+ +EN E+++T+ IK
Sbjct: 865 GELEEGEQSVVFLDTLLEYAADENMEIKITKEQIK 969
>TC18417 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (38%)
Length = 747
Score = 74.3 bits (181), Expect = 1e-14
Identities = 44/143 (30%), Positives = 72/143 (49%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP +S Y + YGPYWR +KK+C T+LLS S++ +R
Sbjct: 297 LKTHDTIFASRPIIQASEYLSYGSKGLAFSEYGPYWRNVKKVCTTQLLSGSKVESSAPLR 476
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
++E+ L+K L+ D+ L NI+ RM +G + N D ++ +V
Sbjct: 477 KEEMGLLVKLLVESGESGEVVDVSEKVAKLIENIVFRMILG-----RSNDDRFDLKGIVH 641
Query: 121 EFMHVGAKLSMGEVLGPLGKFDL 143
E +H+ ++G+ + G DL
Sbjct: 642 EAVHLVGNFNLGDYVHLAGALDL 710
>TC14744 UP|C7DB_LOTJA (O22307) Cytochrome P450 71D11 (Fragment) , complete
Length = 1753
Score = 64.3 bits (155), Expect = 1e-11
Identities = 43/123 (34%), Positives = 63/123 (50%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHD+ F+ RP+ + Y + +PYG YWR ++KIC +LLS ++ IR
Sbjct: 300 MKTHDVTFASRPRSLFTDIVFYGSTDIGFSPYGDYWRQVRKICNVELLSMKRVQSLWPIR 479
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E++ L++ I S E +L +L I R + G K+Y E I C VR
Sbjct: 480 EEEVKNLIQR--IASEEGSVVNLSQAIDSLIFTITSRSAFG----KRYMEQEEFISC-VR 638
Query: 121 EFM 123
E M
Sbjct: 639 EVM 647
>TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete
Length = 1878
Score = 59.7 bits (143), Expect = 4e-10
Identities = 48/198 (24%), Positives = 92/198 (46%), Gaps = 9/198 (4%)
Frame = +3
Query: 4 HDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQE 63
+D+ + RP+F S Y + + YG +WR +++I +LS+ ++ F IR E
Sbjct: 315 NDVVLANRPRFLSGKYIFYNYTTLGSTSYGEHWRNLRRITSLDVLSNHRINSFSPIRRDE 494
Query: 64 LEKLLKSLLICSNENRS-TDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQC----- 117
+L++ L S +N S +L F +T N + RM G + Y D ++ +
Sbjct: 495 TTRLIRKLAEDSAKNFSEVELTSRFFDMTFNNIMRMISGK---RYYGEDCDMTELQEASE 665
Query: 118 ---LVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINT 174
+V E + + + + + L D G K+++ I + D+ L G L+EH ++
Sbjct: 666 FRDMVTELLQLSGANNKADFMPILRLVDFEGLEKRVKGISSKTDRFLRGLLQEHRDK--K 839
Query: 175 EDCQGDMMDILLQVYRNE 192
+ M+D LL + ++
Sbjct: 840 QRTANTMIDHLLTLQESQ 893
>AW720110
Length = 460
Score = 59.3 bits (142), Expect = 5e-10
Identities = 37/148 (25%), Positives = 74/148 (50%), Gaps = 11/148 (7%)
Frame = +2
Query: 33 GPYWRFMKKICVTKLLSSSQLGRFMHIREQELEKLLK---SLLICSNENRSTDLGLD--- 86
GPYWR+M+KI ++LS+ ++ + E+ +K + + E+ +T + +
Sbjct: 2 GPYWRYMRKIATLEVLSTKRIEMLKEVMHSEVMAAVKKSYNYWVMMRESGATKVVTEMEK 181
Query: 87 -FTTLTNNILCRMSMGTTC----FKKYNIDPEVIQCLVREFMHVGAKLSMGEVLGPLGKF 141
F +T N++ R +G + + E I+ VR+F H+ + ++ + + L
Sbjct: 182 WFADITLNVMFRTVLGKRLTGMGCEDDEEENEKIRKAVRDFFHLFGQFTVSDAVPFLRWL 361
Query: 142 DLFGYGKKLRKIVGEFDKILEGFLKEHE 169
DL GY KK++K E D + +L++H+
Sbjct: 362 DLDGYEKKMKKTAKEMDDFAQAWLEQHK 445
>AV409643
Length = 422
Score = 56.2 bits (134), Expect = 4e-09
Identities = 32/132 (24%), Positives = 69/132 (52%), Gaps = 1/132 (0%)
Frame = +1
Query: 1 MKTHDLN-FSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ + F+ R Q + Y S + P+ PYW+F++KI + LL+++ + + +
Sbjct: 55 LQTHEASSFNTRFQTSAIRRLTYDNSVAMV-PFAPYWKFIRKIIMNDLLNATTVNKLRPL 231
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R QE+ K+LK++ + + ++ + TNN + RM +G + E ++ +
Sbjct: 232 RSQEIRKVLKAMAHSAESQQPLNVTEELLKWTNNTISRMMLG---------EAEEVRDIA 384
Query: 120 REFMHVGAKLSM 131
RE + + + S+
Sbjct: 385 REVLKIFGEYSL 420
>AV777368
Length = 604
Score = 53.5 bits (127), Expect = 3e-08
Identities = 28/71 (39%), Positives = 41/71 (57%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
M+THD FS RP S+ LY + A YG WR +K+CV +LLS ++ F IR
Sbjct: 346 MQTHDTVFSSRPHLTSTKALLYGCNDIGFASYGDAWRQKRKLCVLELLSLKRVQSFQFIR 525
Query: 61 EQELEKLLKSL 71
E+E+ L++ +
Sbjct: 526 EEEVAALVEKI 558
>TC19163 similar to UP|Q84J65 (Q84J65) Gray pubescence flavonoid
3'-hydroxylase, partial (36%)
Length = 486
Score = 53.1 bits (126), Expect = 3e-08
Identities = 24/64 (37%), Positives = 34/64 (52%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+K HD NFS RP + Y + APYG WR+++KI L S L F H+R
Sbjct: 291 LKVHDANFSSRPPNAGAKYIAYNYQDLVFAPYGARWRYLRKITNLHLFSGKALDNFKHLR 470
Query: 61 EQEL 64
++E+
Sbjct: 471 QEEV 482
>TC11977 weakly similar to UP|C822_SOYBN (O81972) Cytochrome P450 82A2
(P450 CP4) , partial (25%)
Length = 576
Score = 52.8 bits (125), Expect = 5e-08
Identities = 26/81 (32%), Positives = 47/81 (57%), Gaps = 2/81 (2%)
Frame = +3
Query: 3 THDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQ 62
T+D+ S RP+ ++ Y G+ F A YGPYWR ++KI +L++ ++ + H+R
Sbjct: 297 TNDIVVSTRPKLVATQHMGYNGAMFALALYGPYWRQLRKIVTLGILTNRRVEQLGHVRLS 476
Query: 63 ELEKLLKSL--LICSNENRST 81
E++ +K L + CS +N +
Sbjct: 477 EVQTSIKELYRVWCSQKNEES 539
>TC8699 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (27%)
Length = 561
Score = 48.5 bits (114), Expect = 8e-07
Identities = 24/71 (33%), Positives = 39/71 (54%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD+ F+ RP +S Y I Y YWR M K+C+ +LL+ ++ F +R
Sbjct: 310 LKTHDIVFASRPITQASKYICYDSKGIIFTEYNSYWRNMMKLCMFELLNMPKVQSFAPLR 489
Query: 61 EQELEKLLKSL 71
+E+ ++SL
Sbjct: 490 REEVGLFVESL 522
>TC8141 weakly similar to UP|C79B_ARATH (O81346) Cytochrome P450 79B2 ,
partial (60%)
Length = 1919
Score = 48.1 bits (113), Expect = 1e-06
Identities = 56/230 (24%), Positives = 104/230 (44%), Gaps = 25/230 (10%)
Frame = +2
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITA---PYGPYWRFMKKICVTKLLSSSQLGRFM 57
+K HD +F+ RP+ S+ D G FIT PYG W+ MK++ V LLS + +
Sbjct: 341 LKKHDASFASRPKIMST-DIASDG--FITTVLVPYGEQWKKMKRVLVNNLLSPQKHQWLL 511
Query: 58 HIREQELEKLLKSLL--ICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVI 115
R +E + L+ + C + N D L N + G F+K +
Sbjct: 512 GKRNEEADNLMFYIYNKCCKDVN-------DGPGLVNIRIAAQHYGGNVFRKLIFNSRYF 670
Query: 116 QCLV-------REFMHVGA---------KLSMGEVLGPLGKFDLFGYGKKLRK---IVGE 156
++ E H+ A S+ + + L + DL G+ K+ K I+ +
Sbjct: 671 GKVMEDGGPGFEEVEHINATFTILKYVYAFSISDFVPFLRRLDLDGHRSKIMKAMRIMRK 850
Query: 157 F-DKILEGFLKEHEERINTEDCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
+ D I++ +K+ + + T + D++D+L+++ ++ N + LT ++KA
Sbjct: 851 YHDPIIDDRIKQWNDGLKT--VEEDLLDVLIKL-KDANNKPLLTLKELKA 991
>AV408903
Length = 426
Score = 48.1 bits (113), Expect = 1e-06
Identities = 39/138 (28%), Positives = 65/138 (46%), Gaps = 14/138 (10%)
Frame = +3
Query: 48 LSSSQLGRFMHIREQELEKLLKSLL-ICSNENRS--------TDLGLDFTTLTNNILCRM 98
LSS ++ + IR E+ + L+ + SN + +L F LT N++ R+
Sbjct: 9 LSSRRIEQMSQIRVSEIRTSIHELVDVWSNSKNNEPQSEYVMVELKQWFAQLTLNMILRL 188
Query: 99 SMGTTCFKKYNI---DPEVIQCL--VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKI 153
+G F + + +CL VREFM + ++ + + L D GY K ++K
Sbjct: 189 LVGERYFGATGSAAGEEKAERCLRNVREFMRLMGTFTVADAVPWLRWLDFGGYEKAMKKT 368
Query: 154 VGEFDKILEGFLKEHEER 171
EFDK+L +L EH +R
Sbjct: 369 AEEFDKMLSEWLAEHRQR 422
>TC20073 weakly similar to UP|Q9SML3 (Q9SML3) Cytochrome P450 monooxygenase
(Fragment), partial (10%)
Length = 605
Score = 44.3 bits (103), Expect = 2e-05
Identities = 40/180 (22%), Positives = 76/180 (42%), Gaps = 3/180 (1%)
Frame = +1
Query: 12 PQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQELEKLLKSL 71
P ++H++ + F+ P P W+ ++KIC KL ++ L +R Q+L +L+ +
Sbjct: 4 PDVTTAHNHNHSSLVFL--PVSPLWQDLRKICHHKLFANKTLDASQDLRRQKLLELMSDM 177
Query: 72 LICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVREFMHVGAKLSM 131
S D+G N L + + N+D E + +V + ++
Sbjct: 178 RQSSAGGEVVDIGRAAFKTCINFLSNTFVSQDFVR--NLDDE-YKDIVATLLKATGTPNV 348
Query: 132 GEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED---CQGDMMDILLQV 188
+ + L FD G K + +F IL+ ++E ++ E DM+D LL +
Sbjct: 349 SDFVPALKMFDPQGVRKHTANYLTKFFAILDRLIEE-RSKVRKEKGYVTNNDMLDTLLDI 525
>AV771981
Length = 558
Score = 43.5 bits (101), Expect = 3e-05
Identities = 19/56 (33%), Positives = 33/56 (58%)
Frame = +2
Query: 4 HDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
+D F+ RP+ + Y S F +PYGPYWR+++KI ++LS+ ++ H+
Sbjct: 386 NDKAFASRPKHVAFDVLAYNFSMFGLSPYGPYWRYIRKIASMEVLSNHRIEMLKHV 553
>BI418618
Length = 601
Score = 41.6 bits (96), Expect = 1e-04
Identities = 21/60 (35%), Positives = 38/60 (63%), Gaps = 1/60 (1%)
Frame = +2
Query: 36 WRFMKKICVTKLLSSSQLGRFMHIREQELEKLLKSLLICSNENRSTDLG-LDFTTLTNNI 94
WR +K++C TK+ S+ L +R+++L++LL + SN+ + DLG F+T+ N+I
Sbjct: 371 WRNLKRVCATKVFSTQMLDSTKVLRQEKLKELLDFVKEKSNKGEALDLGEAVFSTVLNSI 550
>TC8057 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (29%)
Length = 1148
Score = 40.4 bits (93), Expect = 2e-04
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 4/82 (4%)
Frame = +2
Query: 131 MGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG----DMMDILL 186
+G+ + +G FDL G +K +K EFD++LE +K+HE D + D D LL
Sbjct: 8 VGDYIPWVGAFDLQGLVRKFKKAHEEFDQMLEQIIKDHEAPSRLSDQKNGQSIDFTDTLL 187
Query: 187 QVYRNENAEVRLTRNDIKAFFL 208
+ + + + ++KA L
Sbjct: 188 SHMKQSKDKHIINKTNLKAILL 253
>BP034275
Length = 524
Score = 39.7 bits (91), Expect = 4e-04
Identities = 20/63 (31%), Positives = 34/63 (53%)
Frame = +3
Query: 3 THDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQ 62
T D F+ RP+ ++ Y AP GP+W+ M++IC+ LL++ +L F R +
Sbjct: 333 TQDDVFASRPRTLAAVHLAYGCGDVALAPLGPHWKRMRRICMEHLLTTKRLESFSRHRHE 512
Query: 63 ELE 65
E +
Sbjct: 513 EAQ 521
>BP037927
Length = 578
Score = 39.7 bits (91), Expect = 4e-04
Identities = 14/40 (35%), Positives = 27/40 (67%)
Frame = +2
Query: 25 SYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQEL 64
S+F ++PYGP WR+M+KI ++LS+ ++ + + E+
Sbjct: 458 SFFGSSPYGPNWRYMRKIATVEVLSTKRIEMLKQVMDSEV 577
>AU089018
Length = 512
Score = 39.3 bits (90), Expect = 5e-04
Identities = 18/66 (27%), Positives = 35/66 (52%)
Frame = +3
Query: 4 HDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQE 63
+D F+ RP+ ++ S F PYG YWR ++KI ++L++ ++ H+ E
Sbjct: 279 NDKAFASRPKTLATEILGLNFSMFAFIPYGSYWRHVRKIATLEVLTTKRIXILKHVMESX 458
Query: 64 LEKLLK 69
++ +K
Sbjct: 459 IKAAMK 476
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,791,235
Number of Sequences: 28460
Number of extensions: 53075
Number of successful extensions: 316
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 309
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 310
length of query: 212
length of database: 4,897,600
effective HSP length: 86
effective length of query: 126
effective length of database: 2,450,040
effective search space: 308705040
effective search space used: 308705040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC142143.3


