

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.13 - phase: 0
(259 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
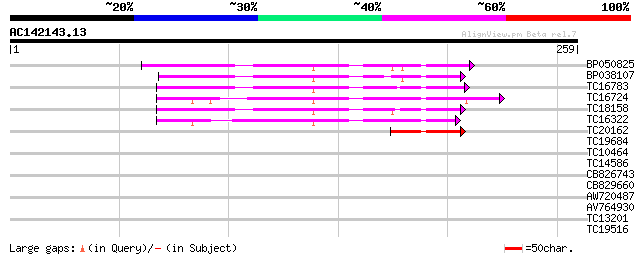
Score E
Sequences producing significant alignments: (bits) Value
BP050825 65 1e-11
BP038107 62 1e-10
TC16783 similar to UP|AAR88435 (AAR88435) NAC domain protein, pa... 59 8e-10
TC16724 similar to GB|BAB20599.1|12060424|AB049070 AtNAC3 {Arabi... 57 2e-09
TC18158 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (34%) 56 5e-09
TC16322 homologue to UP|Q9SQL0 (Q9SQL0) Jasmonic acid 2, partial... 53 5e-08
TC20162 similar to UP|Q8LKN6 (Q8LKN6) Nam-like protein 18, parti... 39 9e-04
TC19684 similar to UP|AAR88435 (AAR88435) NAC domain protein, pa... 37 0.003
TC10464 weakly similar to UP|Q8L8G0 (Q8L8G0) Nam-like protein 1,... 36 0.006
TC14586 similar to UP|Q9FY93 (Q9FY93) NAM-like protein (AT5g1318... 34 0.029
CB826743 33 0.037
CB829660 28 1.6
AW720487 27 3.5
AV764930 26 6.0
TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partia... 26 6.0
TC19516 similar to UP|P93148 (P93148) Cytochrome P450 (Fragment)... 26 7.8
>BP050825
Length = 539
Score = 65.1 bits (157), Expect = 1e-11
Identities = 48/155 (30%), Positives = 68/155 (42%), Gaps = 3/155 (1%)
Frame = +2
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEK 120
G + P LP G +F P D+E++ H K VP +I E + P +
Sbjct: 17 GSQQQPNLPPGFRFHPTDEELVVHYLKKKAASVPLPVAIIAEV--------DLYKFDPWE 172
Query: 121 LPGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII-DGVMK 178
LP G+ +F P + Y G R R + W TG +PV+ +G K
Sbjct: 173 LPAKATFGEQEWYFFSPRDRKYPNGARPNRAATSGY------WKATGTDKPVLTSNGTQK 334
Query: 179 -GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNE 212
G KK LV Y G+ + KTNW+MH+Y L N+
Sbjct: 335 VGVKKALVFYG--GKPPRGIKTNWIMHEYRLADNK 433
>BP038107
Length = 469
Score = 62.0 bits (149), Expect = 1e-10
Identities = 45/143 (31%), Positives = 63/143 (43%), Gaps = 3/143 (2%)
Frame = +3
Query: 69 PAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKDG 128
P G +F P D+E++ H + P P+I E + P LPG+ G
Sbjct: 66 PPGFRFHPTDEELVLHYLCRKCASQPIAVPIIAEI--------DLYKYDPWDLPGLASYG 221
Query: 129 QIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKILV 185
+ +F P + Y G+R R T W TG +P+ G K G KK LV
Sbjct: 222 EKEWYFFSPRDRKYPNGSRPNRAAGTGY------WKATGADKPI---GHPKPVGIKKALV 374
Query: 186 LYTNYGRQKKPEKTNWVMHQYHL 208
Y G+ K +KTNW+MH+Y L
Sbjct: 375 FYA--GKAPKGDKTNWIMHEYRL 437
>TC16783 similar to UP|AAR88435 (AAR88435) NAC domain protein, partial (46%)
Length = 511
Score = 58.9 bits (141), Expect = 8e-10
Identities = 43/144 (29%), Positives = 62/144 (42%), Gaps = 1/144 (0%)
Frame = +1
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D E+++H P P+I E + P +LP +
Sbjct: 97 LPPGFRFHPTDDELVKHYLCAKCASQPINVPIIKEL--------DLYKFDPWQLPDMALY 252
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +F P + Y G+R R T W TG +P+ + G KK LV
Sbjct: 253 GEKEWYFFTPRDRKYPNGSRPNRAAGTGY------WKATGADKPIGKPKTL-GIKKALVF 411
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGS 210
Y G+ K KTNW+MH+Y L +
Sbjct: 412 YA--GKAPKGVKTNWIMHEYRLAN 477
>TC16724 similar to GB|BAB20599.1|12060424|AB049070 AtNAC3 {Arabidopsis
thaliana;}, partial (53%)
Length = 615
Score = 57.4 bits (137), Expect = 2e-09
Identities = 50/165 (30%), Positives = 73/165 (43%), Gaps = 6/165 (3%)
Frame = +2
Query: 68 LPAGVKFDPNDQEIL-QHLEAKVL---FDVPKLHPLIDEFIPTLEGENGICYTHPEKLPG 123
LP G +F P D+E+L Q+L KV F +P + E + P LPG
Sbjct: 134 LPPGFRFFPTDEELLVQYLCRKVAGYHFSLPII------------AEIDLYKFDPWVLPG 277
Query: 124 VKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKK 182
G+ +F P + Y G+R R + W TG + + +G G KK
Sbjct: 278 KAIFGEKEWYFFSPRDRKYPNGSRPNRVAGSGY------WKATGTDKVITTEGRKVGIKK 439
Query: 183 ILVLYTNYGRQKKPEKTNWVMHQYH-LGSNEEEKDGELVVSKKIL 226
LV Y G+ K KTNW+MH+Y L S+ + K G + + +L
Sbjct: 440 ALVFYI--GKAPKGTKTNWIMHEYRLLDSSRKHKLGSSRLDEWVL 568
>TC18158 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (34%)
Length = 476
Score = 56.2 bits (134), Expect = 5e-09
Identities = 44/151 (29%), Positives = 63/151 (41%), Gaps = 10/151 (6%)
Frame = +3
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H K + +P +I E I P +LPG
Sbjct: 78 LPPGFRFHPTDEELILHYLRKKVASIPLPVSIIAEV--------DIYKLDPWELPGKALF 233
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII---------DGVM 177
G+ +F P + Y G R R + W TG + ++ D +
Sbjct: 234 GEKEWYFFSPRDRKYPNGARPNRAAASGY------WKATGTDKTIVASLPGGGRAQDSI- 392
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
G KK LV Y G+ K KTNW+MH+Y L
Sbjct: 393 -GVKKALVFYK--GKPPKGVKTNWIMHEYRL 476
>TC16322 homologue to UP|Q9SQL0 (Q9SQL0) Jasmonic acid 2, partial (40%)
Length = 510
Score = 53.1 bits (126), Expect = 5e-08
Identities = 43/141 (30%), Positives = 60/141 (42%), Gaps = 2/141 (1%)
Frame = +1
Query: 68 LPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKK 126
LP G +F P D+E+L Q+L KV F + E + P LP
Sbjct: 136 LPPGFRFYPTDEELLVQYLCRKVAGH---------NFTLPIIAEIDLYKFDPWVLPSKAI 288
Query: 127 DGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILV 185
G+ +F P + Y G+R R + W TG + + +G G KK LV
Sbjct: 289 FGEKEWYFFSPRDRKYPNGSRPNRVAGSGY------WKATGTDKIITTEGRKVGIKKALV 450
Query: 186 LYTNYGRQKKPEKTNWVMHQY 206
Y G+ K KTNW+MH+Y
Sbjct: 451 FYV--GKAPKGTKTNWIMHEY 507
>TC20162 similar to UP|Q8LKN6 (Q8LKN6) Nam-like protein 18, partial (17%)
Length = 623
Score = 38.9 bits (89), Expect = 9e-04
Identities = 19/34 (55%), Positives = 22/34 (63%)
Frame = +2
Query: 175 GVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
G + G KK LV Y GR K EKTNWVMH++ L
Sbjct: 44 GNLVGMKKTLVFYR--GRAPKGEKTNWVMHEFRL 139
>TC19684 similar to UP|AAR88435 (AAR88435) NAC domain protein, partial (48%)
Length = 535
Score = 37.0 bits (84), Expect = 0.003
Identities = 17/32 (53%), Positives = 21/32 (65%)
Frame = +3
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGS 210
G KK LVLY G+ K KTNW+MH+Y L +
Sbjct: 402 GIKKALVLYV--GKAPKGIKTNWIMHEYRLAN 491
>TC10464 weakly similar to UP|Q8L8G0 (Q8L8G0) Nam-like protein 1, partial
(12%)
Length = 935
Score = 36.2 bits (82), Expect = 0.006
Identities = 16/28 (57%), Positives = 20/28 (71%)
Frame = +1
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQY 206
G KK LV Y +GR K ++TNWVMH+Y
Sbjct: 19 GMKKTLVFY--HGRAPKGKRTNWVMHEY 96
>TC14586 similar to UP|Q9FY93 (Q9FY93) NAM-like protein
(AT5g13180/T19L5_140), partial (18%)
Length = 589
Score = 33.9 bits (76), Expect = 0.029
Identities = 15/33 (45%), Positives = 21/33 (63%)
Frame = +3
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSN 211
G KK LV YT G+ +T+W+MH+Y L +N
Sbjct: 6 GMKKTLVFYT--GKPPHGSRTDWIMHEYRLLTN 98
>CB826743
Length = 513
Score = 33.5 bits (75), Expect = 0.037
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = +2
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQY 206
G KK LV Y GR K ++T+WVMH+Y
Sbjct: 35 GMKKTLVFYA--GRAPKGKRTHWVMHEY 112
>CB829660
Length = 517
Score = 28.1 bits (61), Expect = 1.6
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = +1
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDE 102
LP G +F P D+E++ H + P P+I E
Sbjct: 130 LPPGFRFHPTDEELIVHYLCNQVSSHPSPAPIIPE 234
>AW720487
Length = 575
Score = 26.9 bits (58), Expect = 3.5
Identities = 15/39 (38%), Positives = 19/39 (48%)
Frame = +1
Query: 48 PSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLE 86
P G ++EF G G P V PNDQ +L HL+
Sbjct: 193 PVIGESLEFLSTGW----NGHPGEVHLRPNDQVLLAHLQ 297
>AV764930
Length = 334
Score = 26.2 bits (56), Expect = 6.0
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = +3
Query: 188 TNYGRQKKPEKTNWVMHQYH 207
+N KPE NW+MH +H
Sbjct: 237 SNADSSPKPETLNWLMHLHH 296
>TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partial (22%)
Length = 465
Score = 26.2 bits (56), Expect = 6.0
Identities = 19/61 (31%), Positives = 27/61 (44%)
Frame = +3
Query: 132 HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVLYTNYG 191
+F+ K Y G+ + T+ + W TGK RPV G KK LV + +G
Sbjct: 288 YFYSGLDKKYGKGSSR-----TNRATEKGYWKTTGKDRPVNHANRTVGMKKTLVYH--FG 446
Query: 192 R 192
R
Sbjct: 447 R 449
>TC19516 similar to UP|P93148 (P93148) Cytochrome P450 (Fragment), partial
(73%)
Length = 480
Score = 25.8 bits (55), Expect = 7.8
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = +3
Query: 85 LEAKVLFDVPKLHPLIDEFIPTLEGEN 111
+E K +P+ HPLI +P GEN
Sbjct: 171 MEEKPSITLPRAHPLICVPVPRFSGEN 251
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,536,617
Number of Sequences: 28460
Number of extensions: 59126
Number of successful extensions: 325
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 322
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 323
length of query: 259
length of database: 4,897,600
effective HSP length: 88
effective length of query: 171
effective length of database: 2,393,120
effective search space: 409223520
effective search space used: 409223520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC142143.13


