

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142096.6 - phase: 0 /pseudo
(930 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
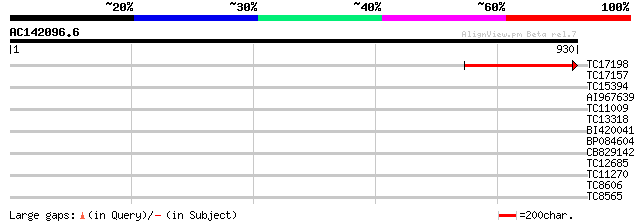
TC17198 weakly similar to UP|Q9LQB3 (Q9LQB3) F4N2.4, partial (20%) 217 6e-57
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 39 0.003
TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%) 35 0.042
AI967639 33 0.21
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 33 0.21
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 32 0.35
BI420041 31 0.79
BP084604 30 1.3
CB829142 30 2.3
TC12685 weakly similar to UP|Q7QA92 (Q7QA92) AgCP13961 (Fragment... 29 3.9
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 28 5.1
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 28 5.1
TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, pa... 28 6.7
>TC17198 weakly similar to UP|Q9LQB3 (Q9LQB3) F4N2.4, partial (20%)
Length = 917
Score = 217 bits (553), Expect = 6e-57
Identities = 125/184 (67%), Positives = 142/184 (76%)
Frame = +2
Query: 747 YGQFQGKCISYRCRNPTRIYQCL**LEFVSRNIFTNFEITT*DSRTEKYAKCITG*N*RC 806
Y F+G C+++ CRN TRIYQ L L+ +S N+ TNFEI T DSRTEKY KCI G*N*RC
Sbjct: 125 YCNFKG*CVTFSCRNSTRIYQHLRRLKLIS*NVLTNFEIVTRDSRTEKYTKCIAG*N*RC 304
Query: 807 G*AD*VESG*TSYFEATTSNAETKASSNQITES*V*GKLR*G*RL*SRPCAS*KKKIEER 866
G*A *VES *TS EATT NA+T+AS+NQI ES V*GKL G RL* R K+KI ER
Sbjct: 305 G*AH*VESR*TSLLEATTPNAQTEASTNQIIESQV*GKLCEGKRL*PRS*TGSKEKIGER 484
Query: 867 GKA*S*RGCS*NT*RQLLLA*CEGKRKVFNGESKS*EVWKD*SFPSRARTCFQIWTTRKR 926
+A*S RGCS* T*RQL LA* EG+R+VF G+ KS EVWK *SFPSRARTCF +WTTRKR
Sbjct: 485 FEA*SQRGCS*IT*RQLFLA*GEGEREVFKGKRKSREVWKS*SFPSRARTCF*VWTTRKR 664
Query: 927 EEEK 930
+EEK
Sbjct: 665 KEEK 676
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 39.3 bits (90), Expect = 0.003
Identities = 30/127 (23%), Positives = 55/127 (42%), Gaps = 7/127 (5%)
Frame = +2
Query: 81 KRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSK---- 136
K K + K + + + F ++ E+D+ +DD L++ E ++ S+ + K
Sbjct: 542 KNTKPITKTEKCESFFNFFNPPQVPEDDDDIDDDAVEELQNLMEHDYDIGSTIRDKIIPH 721
Query: 137 ---YHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG* 193
+ E + +FE I+ D EDE +DDDE +E + + D+ K +
Sbjct: 722 AVSWFTGEAEQSDFEDIEEDDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSG 901
Query: 194 KSKRKGK 200
R GK
Sbjct: 902 SKARPGK 922
>TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%)
Length = 1093
Score = 35.4 bits (80), Expect = 0.042
Identities = 40/162 (24%), Positives = 71/162 (43%)
Frame = +2
Query: 8 SIANGTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGD 67
S + + S + SKK+K P+ KA ++K +++ + G+K K
Sbjct: 278 SASASLSKSKQPASKKRKTENEKPDTKG-KASSKKQTDKSSKALVKDQGKSKSGKKAKAV 454
Query: 68 TKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQL 127
R + ++ V+ +LKE + +T + I R +G + FG ++ R+
Sbjct: 455 PSREAM-HAVVVD-----ILKEVDFNTATLSDILRPLGTH------FGVDLMH----RKA 586
Query: 128 NVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDE 169
VK + +++DE E DG G D +D G+DDDE
Sbjct: 587 EVKDIITDVINNMSDEEDEGEEADGDGDADKDDN--GDDDDE 706
>AI967639
Length = 335
Score = 33.1 bits (74), Expect = 0.21
Identities = 28/91 (30%), Positives = 49/91 (53%)
Frame = +1
Query: 122 QRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDE 181
+R+++ ++ SS + EED++E +G + + EDE +DDD+ D+ E SD
Sbjct: 61 KRKQRRDISSSSEG----DEEDEEEDQGDEY----NLEDEEEDDDDDDDDDLSISEDSDS 216
Query: 182 RSYCKKQVLQG*KSKRKGKR*RPAGRIR*GL 212
K + L G +++R+ KR R I+ GL
Sbjct: 217 DKPRKVKQLPG-RTRRETKR-RSVDEIQSGL 303
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 33.1 bits (74), Expect = 0.21
Identities = 18/41 (43%), Positives = 24/41 (57%)
Frame = +2
Query: 141 EEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDE 181
E+DDD+ E DG DD +D G DDD+ +E +EG E
Sbjct: 29 EDDDDDGEDDDG-DEDDDDDAPGGGDDDDDEEDEEEEGGVE 148
Score = 32.7 bits (73), Expect = 0.27
Identities = 15/43 (34%), Positives = 26/43 (59%)
Frame = +2
Query: 135 SKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQE 177
S+ +S ++DD+ +G D G +D +D+ G DD+ DE +E
Sbjct: 5 SRTRMSGDEDDDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEEE 133
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 32.3 bits (72), Expect = 0.35
Identities = 18/53 (33%), Positives = 31/53 (57%)
Frame = +3
Query: 129 VKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDE 181
V++ + + EE+DDE E D DD +D+ E++D+ D+ +EGS+E
Sbjct: 237 VQTDNEEEEDDDEEEDDEDEDDDD---DDDDDDDGEEEEDDDDDDEEEEGSEE 386
>BI420041
Length = 452
Score = 31.2 bits (69), Expect = 0.79
Identities = 14/46 (30%), Positives = 25/46 (53%)
Frame = +2
Query: 139 LSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSY 184
+ EE+DDEF+ + L +D + + +DD++ +E E D Y
Sbjct: 245 MDEEEDDEFDYEEPLNDEDEDGDEYYDDDEQEEEEFEDEDEDSVPY 382
>BP084604
Length = 463
Score = 30.4 bits (67), Expect = 1.3
Identities = 25/94 (26%), Positives = 43/94 (45%), Gaps = 9/94 (9%)
Frame = -2
Query: 8 SIANGTNTSNKSTSKKKK-------NNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVM 60
S+ANG + K++S KKK ++ + P + A K K S+ F+ +
Sbjct: 462 SVANGEISKIKASSTKKKDKTPAKPSSSLSPGSEASKLNYIKQKLQGLTSMLEASDFKSL 283
Query: 61 GQKRKGDTKRMGLARSLA--VEKRKKTLLKEYEQ 92
K K +++ GL + ++ VE L +EY Q
Sbjct: 282 DTKAKLESEMKGLLQDVSKMVESSSS*LTQEYWQ 181
>CB829142
Length = 534
Score = 29.6 bits (65), Expect = 2.3
Identities = 14/24 (58%), Positives = 16/24 (66%)
Frame = -3
Query: 421 CRFRR*R*YQCWFRRRR*GRRKSC 444
CR RR R ++C RRRR G RK C
Sbjct: 76 CRSRRGRRFRCRRRRRRRGLRKGC 5
>TC12685 weakly similar to UP|Q7QA92 (Q7QA92) AgCP13961 (Fragment), partial
(9%)
Length = 547
Score = 28.9 bits (63), Expect = 3.9
Identities = 21/58 (36%), Positives = 31/58 (53%)
Frame = +2
Query: 135 SKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG 192
++Y + EEDD+EFE D DD+ D+ +D E N E D+ KK+ L+G
Sbjct: 155 TRYKVDEEDDEEFE--DAKEFDDYNDD---DDAPENGYAGNFE-EDDGFARKKRKLRG 310
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 28.5 bits (62), Expect = 5.1
Identities = 23/109 (21%), Positives = 50/109 (45%), Gaps = 4/109 (3%)
Frame = +3
Query: 68 TKRMGLARSLAVEKRKKTLLKE--YEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRER 125
T + L + + + R + +++ ++ K E +++ + E +E +D + E+
Sbjct: 222 TTSITLMKRMRMRMRMRMMMRGGGRKKKLKLVEKLNQGVEEEEEDEED--------EEEQ 377
Query: 126 QLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDF--EDEMLGEDDDETDE 172
+ + H EED++E G + ++ EDE ED+DE +E
Sbjct: 378 TPHPTRGRSRVEHAQEEDEEEENGEEEEEEEEVVVEDEQGNEDEDEDEE 524
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 28.5 bits (62), Expect = 5.1
Identities = 18/58 (31%), Positives = 33/58 (56%)
Frame = -2
Query: 141 EEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG*KSKRK 198
++DDDE + + G +D E+E + E+D+E +E ++ DE + LQ K ++K
Sbjct: 392 DDDDDEDDDEEDDGGEDEEEEGVEEEDNEDEE---EDEEDE------EALQPPKKRKK 246
>TC8565 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial (97%)
Length = 1913
Score = 28.1 bits (61), Expect = 6.7
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +1
Query: 532 NHTCCRKSKESTSILWSIIAVLCCFGK*ET 561
N +CCRKS+ ++ +W VL CF +* +
Sbjct: 1423 NLSCCRKSRSNSRCIW----VLVCFSE*RS 1500
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.377 0.171 0.654
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,841,696
Number of Sequences: 28460
Number of extensions: 179151
Number of successful extensions: 3683
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1518
Number of HSP's successfully gapped in prelim test: 130
Number of HSP's that attempted gapping in prelim test: 2006
Number of HSP's gapped (non-prelim): 1798
length of query: 930
length of database: 4,897,600
effective HSP length: 99
effective length of query: 831
effective length of database: 2,080,060
effective search space: 1728529860
effective search space used: 1728529860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC142096.6


