

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
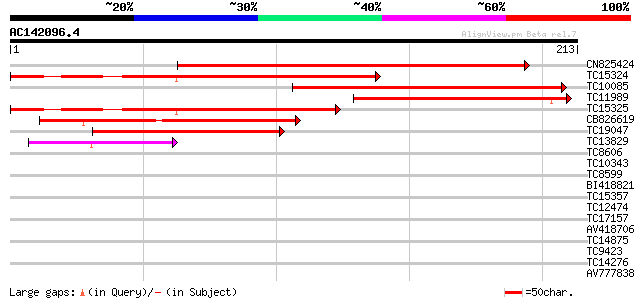
Query= AC142096.4 - phase: 0 /pseudo
(213 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CN825424 212 4e-56
TC15324 similar to UP|O82268 (O82268) At2g31160 protein (At2g311... 186 3e-48
TC10085 similar to UP|AAR92340 (AAR92340) At5g28490, partial (46%) 162 3e-41
TC11989 homologue to UP|Q9LW68 (Q9LW68) Gb|AAC63835.1, partial (... 160 2e-40
TC15325 similar to UP|O82268 (O82268) At2g31160 protein (At2g311... 151 9e-38
CB826619 116 3e-27
TC19047 similar to UP|Q9FGH2 (Q9FGH2) Gb|AAC63835.1, partial (22%) 74 2e-14
TC13829 similar to UP|P93321 (P93321) Cdc2MsD protein, partial (... 39 5e-04
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 38 0.001
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 38 0.001
TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%) 37 0.002
BI418821 37 0.003
TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial... 37 0.003
TC12474 weakly similar to UP|O82576 (O82576) Blue copper protein... 36 0.004
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 36 0.004
AV418706 36 0.004
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 36 0.006
TC9423 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protein... 35 0.007
TC14276 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (... 35 0.007
AV777838 35 0.007
>CN825424
Length = 639
Score = 212 bits (539), Expect = 4e-56
Identities = 96/132 (72%), Positives = 113/132 (84%)
Frame = +2
Query: 64 SRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGH 123
SRYE+QK+RDWNTFGQYLRN RPP++LS+C+ HVLEFLRYLDQFGKTKVH+ C F+G
Sbjct: 8 SRYESQKKRDWNTFGQYLRNQRPPVALSQCNSNHVLEFLRYLDQFGKTKVHSQGCLFFGQ 187
Query: 124 PNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKA 183
PP C CPL+QAWGSLDALIGRLRAA+EENGG PE NPF + ++R++L EVRD QAKA
Sbjct: 188 TEPPGPCTCPLKQAWGSLDALIGRLRAAYEENGGLPETNPFASGSIRVHLHEVRDFQAKA 367
Query: 184 RGISYEKKKRKR 195
RGI Y+KKK+KR
Sbjct: 368 RGIPYKKKKKKR 403
>TC15324 similar to UP|O82268 (O82268) At2g31160 protein
(At2g31160/T16B12.3), partial (39%)
Length = 627
Score = 186 bits (471), Expect = 3e-48
Identities = 91/140 (65%), Positives = 108/140 (77%), Gaps = 1/140 (0%)
Frame = +2
Query: 1 MNSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
M+S+QEF + N+ + ++TN +S T+T S +S S A++SP ++ ST
Sbjct: 245 MDSIQEFIGTCNS------DSTCSLTNNTSMTTLT-------SLASGSGASSSPSASNST 385
Query: 61 T-TPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICP 119
T SRYENQKRRDWNTFGQYL+NHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVH+ +CP
Sbjct: 386 AATSSRYENQKRRDWNTFGQYLKNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHSAMCP 565
Query: 120 FYGHPNPPASCPCPLRQAWG 139
FYGHPNPPA CPCPLRQAWG
Sbjct: 566 FYGHPNPPAPCPCPLRQAWG 625
>TC10085 similar to UP|AAR92340 (AAR92340) At5g28490, partial (46%)
Length = 737
Score = 162 bits (411), Expect = 3e-41
Identities = 74/103 (71%), Positives = 86/103 (82%)
Frame = +3
Query: 107 QFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGA 166
QFGKTKVH CPF+G PNPPA CPCPLRQAWGSLDALIGRLRAA+EENGG+PE NPFGA
Sbjct: 3 QFGKTKVHNHPCPFFGLPNPPAPCPCPLRQAWGSLDALIGRLRAAYEENGGRPETNPFGA 182
Query: 167 RAVRLYLREVRDSQAKARGISYEKKKRKRPPQPPPPPPSNNAT 209
RAVR+YLR+VRD Q KARG+SYEKK+++ P+ P + +T
Sbjct: 183 RAVRIYLRDVRDFQTKARGVSYEKKRKRPKPKMITTPAATAST 311
>TC11989 homologue to UP|Q9LW68 (Q9LW68) Gb|AAC63835.1, partial (38%)
Length = 564
Score = 160 bits (404), Expect = 2e-40
Identities = 78/86 (90%), Positives = 78/86 (90%), Gaps = 4/86 (4%)
Frame = +2
Query: 130 CPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYE 189
CPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYE
Sbjct: 5 CPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYE 184
Query: 190 KKKRKRPPQPPPP----PPSNNATIT 211
KKKRKRPP PPPP PPSN A T
Sbjct: 185 KKKRKRPPHPPPPPAALPPSNGARAT 262
>TC15325 similar to UP|O82268 (O82268) At2g31160 protein
(At2g31160/T16B12.3), partial (32%)
Length = 645
Score = 151 bits (381), Expect = 9e-38
Identities = 77/125 (61%), Positives = 94/125 (74%), Gaps = 1/125 (0%)
Frame = +2
Query: 1 MNSLQEFESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
M+S+QEF + N+ + ++TN +S T+T S +S S A++SP ++ ST
Sbjct: 308 MDSIQEFIGTCNS------DSTCSLTNNTSMTTLT-------SLASGSGASSSPSASNST 448
Query: 61 T-TPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICP 119
T SRYENQKRRDWNTFGQYL+NHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVH+ +CP
Sbjct: 449 AATSSRYENQKRRDWNTFGQYLKNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHSAMCP 628
Query: 120 FYGHP 124
FYGHP
Sbjct: 629 FYGHP 643
>CB826619
Length = 535
Score = 116 bits (291), Expect = 3e-27
Identities = 60/103 (58%), Positives = 75/103 (72%), Gaps = 5/103 (4%)
Frame = +2
Query: 12 NNKDMMNTNPMINIT-----NPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRY 66
NNK ++ + I IT ++S +T P+ S +++++ S+ T + P T+ TTPSRY
Sbjct: 233 NNKPIVQIHRSIFITMDLVSESTTSTPITPPTMSISASATGSSGTVTTP--TAATTPSRY 406
Query: 67 ENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFG 109
ENQKRRDWNTF QYLRNHRPPLSLS CSGAHVLEFL YLDQFG
Sbjct: 407 ENQKRRDWNTFCQYLRNHRPPLSLSLCSGAHVLEFLHYLDQFG 535
>TC19047 similar to UP|Q9FGH2 (Q9FGH2) Gb|AAC63835.1, partial (22%)
Length = 473
Score = 73.9 bits (180), Expect = 2e-14
Identities = 34/72 (47%), Positives = 49/72 (67%)
Frame = +2
Query: 32 MTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLS 91
M T +S+ +A+ + + S + SRYE+QKRRDWNTF QYLRNH+PPL+L+
Sbjct: 257 MEATSGGASSEAAAPAVSHQAEQSSPAAAAPLSRYESQKRRDWNTFLQYLRNHKPPLTLA 436
Query: 92 RCSGAHVLEFLR 103
CSG +V+EF++
Sbjct: 437 WCSGTNVIEFMK 472
>TC13829 similar to UP|P93321 (P93321) Cdc2MsD protein, partial (43%)
Length = 515
Score = 39.3 bits (90), Expect = 5e-04
Identities = 22/58 (37%), Positives = 33/58 (55%), Gaps = 2/58 (3%)
Frame = +2
Query: 8 ESSTNNKDMMNTNPMINITNPS--SSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP 63
+SS + K + ++ PS S + P+ ST+SASS S+ +T PP TT+TT P
Sbjct: 164 KSSLSRKPVSRWTKKASLRPPSAKSPSSRCSPNPSTSSASSKSSTSTKPPKTTTTTPP 337
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 38.1 bits (87), Expect = 0.001
Identities = 18/55 (32%), Positives = 40/55 (72%)
Frame = +1
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTP 63
S++++ +++ + + + PSSS + SSS++S+SSSS+++ S PS++S+++P
Sbjct: 271 SASSSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSP 435
Score = 32.0 bits (71), Expect = 0.083
Identities = 16/43 (37%), Positives = 31/43 (71%)
Frame = +1
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYEN 68
++ SSS + ++ SSST S+SSSS ++S S++S+++ S + +
Sbjct: 280 SSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSS 408
Score = 31.2 bits (69), Expect = 0.14
Identities = 14/36 (38%), Positives = 29/36 (79%)
Frame = +1
Query: 29 SSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+SS + + SSS +S+S+ S++++SPPS++S+++ S
Sbjct: 274 ASSSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSS 381
Score = 28.9 bits (63), Expect = 0.70
Identities = 15/46 (32%), Positives = 27/46 (58%)
Frame = +1
Query: 31 SMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNT 76
S + + SSS++S SSSST ++S S S+++ S + W++
Sbjct: 271 SASSSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSS 408
Score = 28.5 bits (62), Expect = 0.92
Identities = 15/39 (38%), Positives = 28/39 (71%)
Frame = +1
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
++ S S + T SSS++ SSSS++++S S++S ++PS
Sbjct: 298 SSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPS 414
Score = 27.3 bits (59), Expect = 2.0
Identities = 19/56 (33%), Positives = 34/56 (59%), Gaps = 1/56 (1%)
Frame = +1
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASS-SSTATTSPPSTTSTTTP 63
SS+++ +T + + PSSS + + SSS++S SS SS++++SP S+ P
Sbjct: 295 SSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSGPAPAP 462
Score = 25.8 bits (55), Expect = 5.9
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = +1
Query: 28 PSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTT 61
PSS + PS + S SSSS+ ++S PS+ S +
Sbjct: 526 PSSDDKSSSPSPNGPSPSSSSSPSSSLPSSPSAS 627
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 38.1 bits (87), Expect = 0.001
Identities = 29/75 (38%), Positives = 36/75 (47%), Gaps = 15/75 (20%)
Frame = -1
Query: 12 NNKDMMNTNPMINITNPSSSMTMTIPSSST--------TSASSSSTATTSP-------PS 56
NN + + + T SSS T SSST TS SSSS++ TSP P
Sbjct: 249 NNNSAPSLHSSSSATTASSSATPASSSSSTATSCLKRGTSTSSSSSSITSPSSSITPSPK 70
Query: 57 TTSTTTPSRYENQKR 71
+T+TTT SR N R
Sbjct: 69 STTTTTTSRRRNHLR 25
Score = 28.9 bits (63), Expect = 0.70
Identities = 15/45 (33%), Positives = 27/45 (59%)
Frame = -1
Query: 16 MMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTST 60
+ N N ++++ ++S SSS T+ASSS+T +S ST ++
Sbjct: 285 LSNENTLMSVRLNNNSAPSLHSSSSATTASSSATPASSSSSTATS 151
Score = 28.9 bits (63), Expect = 0.70
Identities = 17/57 (29%), Positives = 33/57 (57%), Gaps = 1/57 (1%)
Frame = -1
Query: 9 SSTNNKDMMNTNPMIN-ITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
S T N+ + N + ++N ++ M++ + ++S S SSS+ATT+ S T ++ S
Sbjct: 336 SFTRNQILSNKSIFFTTLSNENTLMSVRLNNNSAPSLHSSSSATTASSSATPASSSS 166
>TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%)
Length = 632
Score = 37.4 bits (85), Expect = 0.002
Identities = 23/76 (30%), Positives = 37/76 (48%), Gaps = 4/76 (5%)
Frame = +3
Query: 12 NNKDMMNTNPMINITNPS--SSMTMTIPSSSTTSAS--SSSTATTSPPSTTSTTTPSRYE 67
++ ++ N P +NI+ PS S + I S TS S SSS++++S T Y
Sbjct: 216 SSSNLSNRKPSLNISIPSINSGLKSNISQSKETSFSHQSSSSSSSSDHQTQEEDNNKHYR 395
Query: 68 NQKRRDWNTFGQYLRN 83
+RR W F +R+
Sbjct: 396 GVRRRPWGKFAAEIRD 443
>BI418821
Length = 614
Score = 37.0 bits (84), Expect = 0.003
Identities = 25/68 (36%), Positives = 35/68 (50%), Gaps = 5/68 (7%)
Frame = +3
Query: 2 NSLQEFESSTNNKDMMNTNPMI-----NITNPSSSMTMTIPSSSTTSASSSSTATTSPPS 56
+S F ST ++ +++PM + TNP S T T PSS S+SS S PP+
Sbjct: 78 DSAASFSGSTMSRVSASSSPMTAEKISSSTNPPSDPTATEPSSKAISSSSPS-----PPA 242
Query: 57 TTSTTTPS 64
TT+ PS
Sbjct: 243 TTTRPKPS 266
Score = 26.2 bits (56), Expect = 4.5
Identities = 8/10 (80%), Positives = 9/10 (90%)
Frame = -3
Query: 195 RPPQPPPPPP 204
+PP PPPPPP
Sbjct: 393 QPPPPPPPPP 364
Score = 25.8 bits (55), Expect = 5.9
Identities = 8/9 (88%), Positives = 8/9 (88%)
Frame = -3
Query: 196 PPQPPPPPP 204
PP PPPPPP
Sbjct: 387 PPPPPPPPP 361
>TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial (69%)
Length = 1578
Score = 36.6 bits (83), Expect = 0.003
Identities = 24/62 (38%), Positives = 33/62 (52%)
Frame = +1
Query: 28 PSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPP 87
P SS+ T PS+ + S+SS S T SP + TSTTT S +K W++ R R
Sbjct: 568 PRSSLRTTNPSTPSPSSSSKSPRTPSP-TPTSTTTLSTSP*RKPCTWSSHSMTQRRKRSE 744
Query: 88 LS 89
+S
Sbjct: 745 IS 750
>TC12474 weakly similar to UP|O82576 (O82576) Blue copper protein
(Fragment), partial (44%)
Length = 818
Score = 36.2 bits (82), Expect = 0.004
Identities = 25/51 (49%), Positives = 28/51 (54%), Gaps = 10/51 (19%)
Frame = +1
Query: 23 INITNPSS-----SMTMTIPSSSTTSASSSSTATTSP-----PSTTSTTTP 63
IN+ + SS S T T PSSSTT SST T SP PS +ST TP
Sbjct: 445 INVASGSSAATPPSSTATPPSSSTTPPPPSSTTTPSPSSPTSPSPSSTVTP 597
Score = 33.9 bits (76), Expect = 0.022
Identities = 20/42 (47%), Positives = 25/42 (58%)
Frame = +1
Query: 23 INITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
++I S S T PSS+ T SSS T+PP +STTTPS
Sbjct: 439 LSINVASGSSAATPPSSTATPPSSS----TTPPPPSSTTTPS 552
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 36.2 bits (82), Expect = 0.004
Identities = 17/39 (43%), Positives = 32/39 (81%)
Frame = -3
Query: 27 NPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSR 65
+PSSS + + SSS++S+SSSS++++S PS++S++ S+
Sbjct: 876 SPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSISSK 760
Score = 31.2 bits (69), Expect = 0.14
Identities = 15/42 (35%), Positives = 31/42 (73%)
Frame = -3
Query: 21 PMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTT 62
P++ + PS S + + SSS++S+SSSS++++S S +S+++
Sbjct: 900 PLLLLLFPSPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSS 775
Score = 29.6 bits (65), Expect = 0.41
Identities = 15/48 (31%), Positives = 29/48 (60%)
Frame = -3
Query: 7 FESSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSP 54
F S +++ +++ + ++ SSS + + PSSS++S SS S + SP
Sbjct: 882 FPSPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSISSKSLCSASP 739
Score = 27.3 bits (59), Expect = 2.0
Identities = 14/36 (38%), Positives = 27/36 (74%)
Frame = -3
Query: 29 SSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
SSS + + SSS++S+SSSS++++ S++S ++ S
Sbjct: 864 SSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSISSKS 757
Score = 27.3 bits (59), Expect = 2.0
Identities = 15/39 (38%), Positives = 27/39 (68%)
Frame = -3
Query: 21 PMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTS 59
P + ++ SSS + + SSS++S+SSSS+ ++S S +S
Sbjct: 879 PSPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSISS 763
Score = 26.9 bits (58), Expect = 2.7
Identities = 15/45 (33%), Positives = 30/45 (66%)
Frame = -3
Query: 20 NPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+P + ++ SSS + + SSS++S+SSS ++++S S+ S + S
Sbjct: 876 SPSSSSSSSSSSSSSSSSSSSSSSSSSSPSSSSSSISSKSLCSAS 742
>AV418706
Length = 374
Score = 36.2 bits (82), Expect = 0.004
Identities = 22/60 (36%), Positives = 36/60 (59%), Gaps = 12/60 (20%)
Frame = +1
Query: 21 PMINITNPSSSMTMTIP-------SSSTTSASSSSTATTSPPSTTSTT-----TPSRYEN 68
P + + +P ++ T+ SSS++S+SSSS+A +P ST+STT +PSR+ N
Sbjct: 109 PTVPVLHPLAAAAATVAAAHSSAGSSSSSSSSSSSSAPPAPSSTSSTTLNALPSPSRHSN 288
Score = 27.3 bits (59), Expect = 2.0
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +1
Query: 196 PPQPPPPPPSNNA 208
PP PPP PPS+N+
Sbjct: 307 PPPPPPSPPSSNS 345
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 35.8 bits (81), Expect = 0.006
Identities = 21/42 (50%), Positives = 31/42 (73%), Gaps = 2/42 (4%)
Frame = +1
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTA--TTSPPSTTSTTTPSR 65
T+P + T T PSSST++A+ S++A TT+P + TSTT+P R
Sbjct: 196 TSPLTPPT-TSPSSSTSTAAPSASAPLTTTPTTATSTTSPRR 318
Score = 35.4 bits (80), Expect = 0.007
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = +1
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+NP +S + P+S S S + TTSP S+TST PS
Sbjct: 142 SNPKTSTSTPTPASPPASTSPLTPPTTSPSSSTSTAAPS 258
Score = 33.5 bits (75), Expect = 0.028
Identities = 17/41 (41%), Positives = 28/41 (67%), Gaps = 2/41 (4%)
Frame = +1
Query: 26 TNPSSSMTMTIPSSST--TSASSSSTATTSPPSTTSTTTPS 64
T+PSSS + PS+S T+ +++T+TTSP T +++PS
Sbjct: 220 TSPSSSTSTAAPSASAPLTTTPTTATSTTSPRRQTPSSSPS 342
>TC9423 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protein B13-1-9,
partial (53%)
Length = 969
Score = 35.4 bits (80), Expect = 0.007
Identities = 17/56 (30%), Positives = 34/56 (60%)
Frame = +2
Query: 9 SSTNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPS 64
+S+ +++ + ++ S+S++ PSSS + +S ++TT PP+TT+ TT S
Sbjct: 173 ASSGRLSSQSSSSSPSYSSSSTSLSNHAPSSSMSPTPNSHSSTTPPPTTTTATTTS 340
Score = 32.3 bits (72), Expect = 0.063
Identities = 22/58 (37%), Positives = 39/58 (66%), Gaps = 2/58 (3%)
Frame = +2
Query: 8 ESSTNNKDMMNTNPMINITNPSSSMTMTIPS-SSTTSASSSSTATTSPPSTT-STTTP 63
+SS+++ +++ ++ PSSSM+ T S SSTT +++TATT+ +TT S+T+P
Sbjct: 197 QSSSSSPSYSSSSTSLSNHAPSSSMSPTPNSHSSTTPPPTTTTATTTSFATTLSSTSP 370
Score = 32.0 bits (71), Expect = 0.083
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 12/74 (16%)
Frame = +2
Query: 10 STNNKDMMNTNPMINITNPSSSMTMTIPSSSTTSASSSSTATT----SPP--------ST 57
S+++ + N P +++ +S + T P +TT+A+++S ATT SPP ST
Sbjct: 224 SSSSTSLSNHAPSSSMSPTPNSHSSTTPPPTTTTATTTSFATTLSSTSPPATRTKSSTST 403
Query: 58 TSTTTPSRYENQKR 71
T+ PSR R
Sbjct: 404 TTRWRPSRSTTASR 445
>TC14276 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (16%)
Length = 624
Score = 35.4 bits (80), Expect = 0.007
Identities = 23/46 (50%), Positives = 29/46 (63%), Gaps = 5/46 (10%)
Frame = +3
Query: 23 INITNPSSSMTMTIPSSSTTSASSSSTATTSP-----PSTTSTTTP 63
IN+T SS T T PSS++ S S SS+ T SP PS +S+TTP
Sbjct: 153 INVTGGSS--TATPPSSASPSPSPSSSTTPSPSSSTTPSPSSSTTP 284
>AV777838
Length = 565
Score = 35.4 bits (80), Expect = 0.007
Identities = 20/47 (42%), Positives = 27/47 (56%)
Frame = +1
Query: 25 ITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKR 71
IT P SS + + SSS++S+SSSS T+ TT T PS + R
Sbjct: 202 ITGPPSSSSSSSSSSSSSSSSSSSMRATAAGHTTITAPPSSSSSSMR 342
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,965,795
Number of Sequences: 28460
Number of extensions: 109842
Number of successful extensions: 5999
Number of sequences better than 10.0: 759
Number of HSP's better than 10.0 without gapping: 2898
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4859
length of query: 213
length of database: 4,897,600
effective HSP length: 86
effective length of query: 127
effective length of database: 2,450,040
effective search space: 311155080
effective search space used: 311155080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC142096.4


