

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141863.5 - phase: 0
(139 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
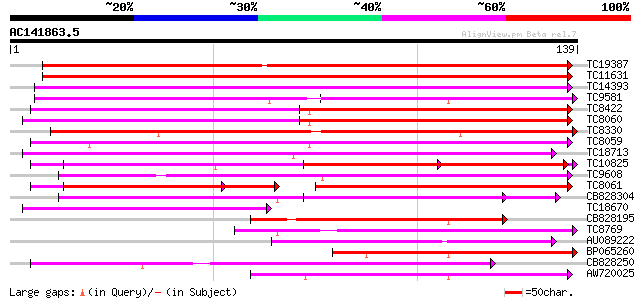
Score E
Sequences producing significant alignments: (bits) Value
TC19387 weakly similar to UP|Q9FYK2 (Q9FYK2) F21J9.28, partial (... 106 2e-24
TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-rela... 102 3e-23
TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 95 5e-21
TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete 93 2e-20
TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete 88 6e-19
TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regu... 87 8e-19
TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodu... 82 3e-17
TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus ann... 79 3e-16
TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13... 77 8e-16
TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Ar... 70 1e-13
TC9608 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial ... 59 4e-10
TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete 48 1e-09
CB828304 49 4e-07
TC18670 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, p... 48 7e-07
CB828195 47 9e-07
TC8769 similar to UP|Q42438 (Q42438) Calcium-dependent protein k... 46 2e-06
AU089222 44 1e-05
BP065260 40 1e-04
CB828250 40 1e-04
AW720025 39 3e-04
>TC19387 weakly similar to UP|Q9FYK2 (Q9FYK2) F21J9.28, partial (45%)
Length = 631
Score = 106 bits (264), Expect = 2e-24
Identities = 51/130 (39%), Positives = 82/130 (62%)
Frame = +1
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
+ + FD +GDGK+S +EL+ + +G + +E + +D +GDG++ L+E
Sbjct: 58 IFKKFDRNGDGKISCSELKDLMAALGSKTTAEEVRRMMAELDRNGDGHIDLKEF-GEFHR 234
Query: 69 GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGL 128
GG K++REAFEMYD +K G I+ K L ++KK+GE S+ +C+ MI + D+DGDG
Sbjct: 235 GGGGGNSKEIREAFEMYDLDKNGLISAKELHSVMKKLGEKCSLGDCRKMIGNVDVDGDGN 414
Query: 129 LSFDEFITMM 138
++F+EF MM
Sbjct: 415 VNFEEFKKMM 444
>TC11631 weakly similar to UP|TCH2_ARATH (P25070) Calmodulin-related protein
2, touch-induced, partial (68%)
Length = 530
Score = 102 bits (253), Expect = 3e-23
Identities = 49/130 (37%), Positives = 79/130 (60%)
Frame = +3
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
+ FD++GDGK+S EL++ L +G + +E + + +D +GDGY+ L+E
Sbjct: 111 IFNKFDKNGDGKISCTELKEMLCALGTKTSSEEVKRMMAEIDQNGDGYIDLKEFAEFHLH 290
Query: 69 GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGL 128
E K+LR+A ++YD +K G I+ L +LKK+GE S+ +C+ MI + D DGDG
Sbjct: 291 DAGEDDSKELRDACDLYDLDKNGLISANELHSVLKKLGEKCSLGDCRKMISNVDADGDGN 470
Query: 129 LSFDEFITMM 138
++F+EF MM
Sbjct: 471 VNFEEFKKMM 500
>TC14393 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(85%)
Length = 786
Score = 94.7 bits (234), Expect = 5e-21
Identities = 49/132 (37%), Positives = 76/132 (57%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ V FD +GDGK+S EL LR +G + +E + + +D D DG+++L E A
Sbjct: 156 KRVFTRFDTNGDGKISVNELDNVLRSLGSGVPPEELKRVMVDLDGDHDGFINLSEFAAFC 335
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+ +L +AFE+YD +K G I+ L ++L ++G + S +EC MIK D DGD
Sbjct: 336 RSDTADGGASELHDAFELYDQDKNGLISAAELCKVLNRLGMNCSEEECHKMIKSVDSDGD 515
Query: 127 GLLSFDEFITMM 138
G ++F+EF MM
Sbjct: 516 GNVNFEEFKKMM 551
>TC9581 UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, complete
Length = 1049
Score = 92.8 bits (229), Expect = 2e-20
Identities = 50/139 (35%), Positives = 84/139 (59%), Gaps = 6/139 (4%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEEL---- 62
+ V + FD +GDG+++ EL L +G I KE IE +D +GDG + ++E
Sbjct: 354 KRVFQMFDRNGDGRITKKELNDSLENLGIFIPDKELTQMIERIDVNGDGCVDIDEFGELY 533
Query: 63 IALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAMIKH 120
++M+E EE+ D+REAF ++D GFIT + L+ +L +G + +++++CK MI
Sbjct: 534 QSIMDERDEEE---DMREAFNVFDQNGDGFITVEELRTVLASLGIKQGRTVEDCKKMIMK 704
Query: 121 FDLDGDGLLSFDEFITMMQ 139
D+DGDG++ + EF MM+
Sbjct: 705 VDVDGDGMVDYKEFKQMMK 761
Score = 42.0 bits (97), Expect = 4e-05
Identities = 22/63 (34%), Positives = 33/63 (51%)
Frame = +3
Query: 77 DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFIT 136
+L+ F+M+D G IT K L L+ +G E MI+ D++GDG + DEF
Sbjct: 348 ELKRVFQMFDRNGDGRITKKELNDSLENLGIFIPDKELTQMIERIDVNGDGCVDIDEFGE 527
Query: 137 MMQ 139
+ Q
Sbjct: 528 LYQ 536
>TC8422 UP|AAQ63461 (AAQ63461) Calmodulin 4 (Fragment), complete
Length = 832
Score = 87.8 bits (216), Expect = 6e-19
Identities = 42/134 (31%), Positives = 79/134 (58%), Gaps = 1/134 (0%)
Frame = +1
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E + L
Sbjct: 172 FKEAFSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLNL 351
Query: 66 MEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
M ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D+D
Sbjct: 352 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 531
Query: 125 GDGLLSFDEFITMM 138
GDG ++++EF+ +M
Sbjct: 532 GDGQINYEEFVKVM 573
Score = 51.6 bits (122), Expect = 5e-08
Identities = 21/67 (31%), Positives = 41/67 (60%)
Frame = +1
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ ++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 154 DDQISEFKEAFSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 333
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 334 PEFLNLM 354
>TC8060 UP|O49184 (O49184) Calmodulin (ESTS AU081349) (Auxin-regulated
calmodulin) (Calmodulin TACAM1-1) (Calmodulin TACAM1-2)
(Calmodulin TACAM1-3) (Calmodulin TACAM3-1) (Calmodulin
TACAM3-2) (Calmodulin TACAM3-3) (Calmodulin TACAM4-1)
(CaM protein), complete
Length = 615
Score = 87.4 bits (215), Expect = 8e-19
Identities = 43/136 (31%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Frame = +3
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 99 AEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 278
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
LM ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 279 NLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 458
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+ +M
Sbjct: 459 VDGDGQINYEEFVKVM 506
Score = 51.2 bits (121), Expect = 6e-08
Identities = 21/67 (31%), Positives = 41/67 (60%)
Frame = +3
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ ++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 87 DDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 266
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 267 PEFLNLM 287
>TC8330 weakly similar to GB|AAM61257.1|21536925|AY084696 calmodulin-like
protein {Arabidopsis thaliana;} , partial (72%)
Length = 1174
Score = 82.4 bits (202), Expect = 3e-17
Identities = 46/131 (35%), Positives = 81/131 (61%), Gaps = 2/131 (1%)
Frame = +3
Query: 11 RYFDEDGDGKVSPAELRQRLRIMGE-EILLKEAEMAIEAMDSDGDGYLSLEELIALMEEG 69
R D D DG VS A+L L +G ++ +E E+ + +D DG G +S+E L+ + G
Sbjct: 309 RLIDRDNDGVVSRADLVAVLTRLGALQLSDEEVEVMLREVDGDGRGSISVEALMERVGPG 488
Query: 70 GEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESK-SIDECKAMIKHFDLDGDGL 128
+ + +LREAFE++D+++ G I+ + L R+ +G+ + ++DEC+ MI+ D +GDG
Sbjct: 489 SDPET--ELREAFEVFDTDRDGRISAEELLRVFTAIGDERCTLDECRRMIEGVDRNGDGF 662
Query: 129 LSFDEFITMMQ 139
+ F++F MM+
Sbjct: 663 VCFEDFSRMME 695
>TC8059 GB|AAB68399.1|1773321|HAU79736 calmodulin {Helianthus annuus;} ,
partial (83%)
Length = 836
Score = 79.0 bits (193), Expect = 3e-16
Identities = 42/135 (31%), Positives = 77/135 (56%), Gaps = 2/135 (1%)
Frame = +2
Query: 6 FEHVLRYFDEDGD-GKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
F H L DE D G ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 122 FAHTLTQADEAADPGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLN 301
Query: 65 LMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
LM ++ ++L+EAF ++D ++ GFI+ L+ ++ +GE + +E MI+ D+
Sbjct: 302 LMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADV 481
Query: 124 DGDGLLSFDEFITMM 138
DGDG ++++EF+ +M
Sbjct: 482 DGDGQINYEEFVKVM 526
>TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13E7_5
{Arabidopsis thaliana;}, partial (88%)
Length = 868
Score = 77.4 bits (189), Expect = 8e-16
Identities = 42/135 (31%), Positives = 69/135 (51%), Gaps = 4/135 (2%)
Frame = +1
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A + R FD + DG ++ EL LR +G + ++ E I+ D++ +G + E +
Sbjct: 211 AELREIFRSFDRNNDGSLTQLELSSLLRSLGLKPSAEQVEGFIQKADTNSNGLIEFSEFV 390
Query: 64 ALMEE----GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIK 119
A++ + LR+ F ++D + GFIT L + K+G + S +E MIK
Sbjct: 391 AVVAPELLPARSPYTEEQLRQLFRVFDRDGNGFITAAELAHSMAKLGHALSAEELTGMIK 570
Query: 120 HFDLDGDGLLSFDEF 134
D DGDG++SF EF
Sbjct: 571 EADADGDGMISFQEF 615
>TC10825 similar to GB|AAP68339.1|31711966|BT008900 At1g74740 {Arabidopsis
thaliana;}, partial (32%)
Length = 1091
Score = 70.5 bits (171), Expect = 1e-13
Identities = 38/126 (30%), Positives = 66/126 (52%)
Frame = +1
Query: 14 DEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEEQ 73
D D DG+VS EL+ LR +G ++ +E +M +E D DG+G L E +A+ + +
Sbjct: 43 DTDKDGRVSYEELKAGLRKVGSQLADQEIKMLMEVADVDGNGVLDYGEFVAVTIHLQKME 222
Query: 74 KLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDE 133
+ +AF+ +D + G+I L+ L D +++ D D DG +S++E
Sbjct: 223 NDEHFHKAFKFFDKDGSGYIESGELQEALADESGVTDADVLNDIMREVDTDKDGRISYEE 402
Query: 134 FITMMQ 139
F+ MM+
Sbjct: 403 FVAMMK 420
Score = 45.1 bits (105), Expect = 5e-06
Identities = 33/104 (31%), Positives = 52/104 (49%), Gaps = 3/104 (2%)
Frame = +1
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAM---DSDGDGYLSLEEL 62
F ++FD+DG G + EL++ L +E + +A++ + M D+D DG +S EE
Sbjct: 235 FHKAFKFFDKDGSGYIESGELQEAL---ADESGVTDADVLNDIMREVDTDKDGRISYEEF 405
Query: 63 IALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG 106
+A+M+ G D R+A Y E+ KSL L K G
Sbjct: 406 VAMMKAG------TDWRKASRQYSRERF-----KSLSLNLMKDG 504
Score = 44.3 bits (103), Expect = 8e-06
Identities = 18/65 (27%), Positives = 43/65 (65%)
Frame = +1
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
++++ +++ F + D++K G ++ + LK L+K+G + E K +++ D+DG+G+L +
Sbjct: 4 EEVEIIKDMFTLMDTDKDGRVSYEELKAGLRKVGSQLADQEIKMLMEVADVDGNGVLDYG 183
Query: 133 EFITM 137
EF+ +
Sbjct: 184 EFVAV 198
>TC9608 similar to UP|Q94AZ4 (Q94AZ4) At1g12310/F5O11_2, partial (97%)
Length = 817
Score = 58.5 bits (140), Expect = 4e-10
Identities = 34/127 (26%), Positives = 64/127 (49%), Gaps = 1/127 (0%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD DGDGK++P+EL +R +G +A++ + + + LM + +
Sbjct: 162 FDTDGDGKIAPSELGILMRSLGGN--PTQAQLKSIVAEENLTSPFDFPRFLDLMAKHMKP 335
Query: 73 QKL-KDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ + LR+AF++ D + GF++ L+ +L +GE E I+ D+ DG + +
Sbjct: 336 EPFDRQLRDAFKVIDKDSTGFVSVTELRHILTSIGEKLEPAEFDEWIREVDVGSDGKIRY 515
Query: 132 DEFITMM 138
++FI M
Sbjct: 516 EDFIARM 536
Score = 35.4 bits (80), Expect = 0.004
Identities = 25/70 (35%), Positives = 34/70 (47%), Gaps = 4/70 (5%)
Frame = +3
Query: 1 MKNAGFEHVLR----YFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGY 56
MK F+ LR D+D G VS ELR L +GE++ E + I +D DG
Sbjct: 327 MKPEPFDRQLRDAFKVIDKDSTGFVSVTELRHILTSIGEKLEPAEFDEWIREVDVGSDGK 506
Query: 57 LSLEELIALM 66
+ E+ IA M
Sbjct: 507 IRYEDFIARM 536
>TC8061 UP|AAA34013 (AAA34013) Calmodulin, complete
Length = 568
Score = 48.1 bits (113), Expect(2) = 1e-09
Identities = 20/63 (31%), Positives = 39/63 (61%)
Frame = +2
Query: 76 KDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFI 135
K R+ + D ++ GFI+ L+ ++ +GE + +E MI+ D+DGDG ++++EF+
Sbjct: 296 KSSRKLSALLDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEFV 475
Query: 136 TMM 138
+M
Sbjct: 476 KVM 484
Score = 42.7 bits (99), Expect = 2e-05
Identities = 20/53 (37%), Positives = 33/53 (61%)
Frame = +2
Query: 14 DEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
D+D +G +S AELR + +GE++ +E + I D DGDG ++ EE + +M
Sbjct: 326 DKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVDGDGQINYEEFVKVM 484
Score = 32.0 bits (71), Expect = 0.040
Identities = 14/46 (30%), Positives = 25/46 (53%)
Frame = +1
Query: 93 ITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFITMM 138
+ P++L +G ++ C+ MI D DG+G + F EF+ +M
Sbjct: 127 LPPRNLGL*CGHLGRTQLRLSCRHMINEVDADGNGTIDFPEFLNLM 264
Score = 28.5 bits (62), Expect(2) = 1e-09
Identities = 15/48 (31%), Positives = 24/48 (49%)
Frame = +3
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDG 53
F+ FD+DGDG ++ EL +R +G+ E ++A D G
Sbjct: 81 FKEAFSLFDKDGDGCITTKELGTVMRSLGQ----NPTEAELQAHDQRG 212
Score = 24.3 bits (51), Expect = 8.4
Identities = 14/53 (26%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = +1
Query: 33 MGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM-EEGGEEQKLKDLREAFEM 84
+G L I +D+DG+G + E + LM + + ++L+EAF +
Sbjct: 163 LGRTQLRLSCRHMINEVDADGNGTIDFPEFLNLMARKMKDTDSEEELKEAFRV 321
>CB828304
Length = 489
Score = 48.5 bits (114), Expect = 4e-07
Identities = 32/115 (27%), Positives = 53/115 (45%), Gaps = 5/115 (4%)
Frame = +1
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA-----LME 67
FD D DG ++ EL LR +G + + + MDS+G+G + +EL+ L
Sbjct: 142 FDMDSDGSLTMLELAALLRSLGLKPSGDQLHDLLSNMDSNGNGSVEFDELVRTILPDLKN 321
Query: 68 EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+ L + F+ +D + GFI+ L + KMG+ + E MI+ D
Sbjct: 322 NAEVLLNQEQLLDVFKCFDRDSNGFISAAELAGAMAKMGQPLTYKELTEMIREAD 486
Score = 43.5 bits (101), Expect = 1e-05
Identities = 20/63 (31%), Positives = 35/63 (54%)
Frame = +1
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
Q+L LRE F +D + G +T L +L+ +G S D+ ++ + D +G+G + FD
Sbjct: 106 QQLNQLREIFARFDMDSDGSLTMLELAALLRSLGLKPSGDQLHDLLSNMDSNGNGSVEFD 285
Query: 133 EFI 135
E +
Sbjct: 286 ELV 294
Score = 29.6 bits (65), Expect = 0.20
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +1
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDS 51
V + FD D +G +S AEL + MG+ + KE I D+
Sbjct: 361 VFKCFDRDSNGFISAAELAGAMAKMGQPLTYKELTEMIREADT 489
>TC18670 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(41%)
Length = 329
Score = 47.8 bits (112), Expect = 7e-07
Identities = 24/61 (39%), Positives = 35/61 (57%)
Frame = +3
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A E V FD + DGK+S EL LR +G + +E + +E +D+D DG++SL E
Sbjct: 111 AELERVFNRFDANSDGKISVDELDSVLRTLGSGVPPEELQQVMEDLDTDRDGFISLPEFA 290
Query: 64 A 64
A
Sbjct: 291 A 293
>CB828195
Length = 551
Score = 47.4 bits (111), Expect = 9e-07
Identities = 24/65 (36%), Positives = 43/65 (65%), Gaps = 2/65 (3%)
Frame = +3
Query: 60 EELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAM 117
+EL L E+ +E L+++++AF+++D + GFI + L R+L +G E I++CK M
Sbjct: 360 KELSELFED--QEPSLEEVKQAFDVFDENRDGFIDARELHRVLCVLGLKEEAGIEKCKIM 533
Query: 118 IKHFD 122
I++FD
Sbjct: 534 IRNFD 548
>TC8769 similar to UP|Q42438 (Q42438) Calcium-dependent protein kinase ,
partial (21%)
Length = 1144
Score = 46.2 bits (108), Expect = 2e-06
Identities = 28/88 (31%), Positives = 48/88 (53%), Gaps = 4/88 (4%)
Frame = +2
Query: 56 YLSLEELIA----LMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSI 111
YL+ E +A L + G +E +K AF+ +D+ + G+I + L+ L E+ S
Sbjct: 2 YLNYGEFVAISVHLRKMGNDEHLIK----AFKFFDANQSGYIEIEELRDALSDEVETNSE 169
Query: 112 DECKAMIKHFDLDGDGLLSFDEFITMMQ 139
+ A++ D D DG +S++EF TMM+
Sbjct: 170 EVISAIMHDVDTDKDGKISYEEFATMMK 253
>AU089222
Length = 345
Score = 43.5 bits (101), Expect = 1e-05
Identities = 22/70 (31%), Positives = 39/70 (55%)
Frame = +3
Query: 65 LMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
L++ + Q + D F+ +D+ G I+ L LK +G S + +E + M+ D+D
Sbjct: 39 LIKMADDPQDVADRERIFKRFDANGDGQISSSELGEALKTLG-SVTAEEVQRMMDEIDID 215
Query: 125 GDGLLSFDEF 134
GDG +S++EF
Sbjct: 216 GDGFISYEEF 245
>BP065260
Length = 446
Score = 40.4 bits (93), Expect = 1e-04
Identities = 20/63 (31%), Positives = 40/63 (62%), Gaps = 3/63 (4%)
Frame = -1
Query: 80 EAFEMYD-SEKCGFITPKSLKRMLKKMG--ESKSIDECKAMIKHFDLDGDGLLSFDEFIT 136
+AF+++ ++ + P+ L R+L +G E I++C+ MI++FD + DG + F EF+
Sbjct: 257 QAFDIFG*EQRWVLLMPRELHRVLCVLGLKEEPGIEKCQIMIRNFDKNPDGRIDFLEFVK 78
Query: 137 MMQ 139
+M+
Sbjct: 77 IME 69
>CB828250
Length = 485
Score = 40.0 bits (92), Expect = 1e-04
Identities = 30/120 (25%), Positives = 53/120 (44%), Gaps = 6/120 (5%)
Frame = +1
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLR------IMGEEILLKEAEMAIEAMDSDGDGYLSL 59
F D D DGK+S +LR G++++ +A D+D DG++
Sbjct: 118 FRPAFDVLDADCDGKISRDDLRAFYAGIYGGVASGDDVIGAMMTVA----DTDKDGFVEY 285
Query: 60 EELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIK 119
EE ++ E + + E F++ D + G ++ LK + G S + ++ AMIK
Sbjct: 286 EEFERVVTENNKPLGCGAMEEVFKVMDKDGDGKLSHHDLKSYMASAGFSATDEDINAMIK 465
Score = 35.8 bits (81), Expect = 0.003
Identities = 24/96 (25%), Positives = 45/96 (46%), Gaps = 2/96 (2%)
Frame = +1
Query: 45 AIEAMDSDGDGYLSLEELIALMEE--GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRML 102
A + +D+D DG +S ++L A GG + + D++K GF+ + +R++
Sbjct: 127 AFDVLDADCDGKISRDDLRAFYAGIYGGVASGDDVIGAMMTVADTDKDGFVEYEEFERVV 306
Query: 103 KKMGESKSIDECKAMIKHFDLDGDGLLSFDEFITMM 138
+ + + + K D DGDG LS + + M
Sbjct: 307 TENNKPLGCGAMEEVFKVMDKDGDGKLSHHDLKSYM 414
>AW720025
Length = 563
Score = 39.3 bits (90), Expect = 3e-04
Identities = 25/81 (30%), Positives = 45/81 (54%), Gaps = 2/81 (2%)
Frame = +3
Query: 60 EELIALMEEGGE-EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG-ESKSIDECKAM 117
E+L+ +M E + E + +L F++ + G I+ +SL R +G + + ++ + M
Sbjct: 141 EDLLPVMAEKLDVETFVSELCGGFKLLSDPETGLISSESLMRNSALLGMDGMTKEDAEEM 320
Query: 118 IKHFDLDGDGLLSFDEFITMM 138
+K DLDGDG L+ EF +M
Sbjct: 321 VKQGDLDGDGSLNETEFCILM 383
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.138 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,590,942
Number of Sequences: 28460
Number of extensions: 14303
Number of successful extensions: 162
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 119
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 139
length of database: 4,897,600
effective HSP length: 81
effective length of query: 58
effective length of database: 2,592,340
effective search space: 150355720
effective search space used: 150355720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 50 (23.9 bits)
Medicago: description of AC141863.5


