

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
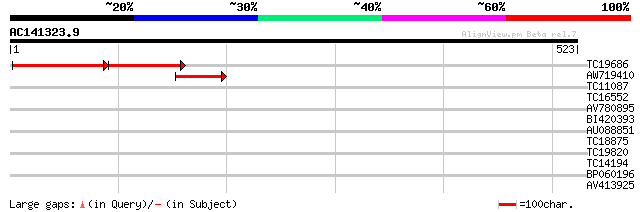
Query= AC141323.9 - phase: 0
(523 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19686 weakly similar to UP|Q9FIN8 (Q9FIN8) Genomic DNA, chromo... 150 3e-67
AW719410 47 8e-06
TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protei... 37 0.008
TC16552 similar to UP|Q94IE2 (Q94IE2) APG1, partial (31%) 31 0.43
AV780895 31 0.57
BI420393 28 2.8
AU088851 28 3.7
TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, pa... 28 4.8
TC19820 similar to GB|AAO63279.1|28950711|BT005215 At3g56740 {Ar... 27 8.2
TC14194 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific... 27 8.2
BP060196 27 8.2
AV413925 27 8.2
>TC19686 weakly similar to UP|Q9FIN8 (Q9FIN8) Genomic DNA, chromosome 5,
TAC clone:K16H17 (At5g24340), partial (20%)
Length = 487
Score = 150 bits (379), Expect(2) = 3e-67
Identities = 73/90 (81%), Positives = 81/90 (89%), Gaps = 1/90 (1%)
Frame = +3
Query: 3 PPNQKPLEIHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQ 62
P NQKP E+H VT+TDSPEFT L+R+LTQTS++GLDAEWKPVRTHQ SFP VSLLQIACQ
Sbjct: 3 PHNQKPFEVHLVTSTDSPEFTLLSRTLTQTSIIGLDAEWKPVRTHQTSFPAVSLLQIACQ 182
Query: 63 L-GDDEVVFLLDLISLPLSSIWEPLREMLV 91
L G D VVFLLDL+SLPLSS+WEPLREMLV
Sbjct: 183LRGGDSVVFLLDLLSLPLSSLWEPLREMLV 272
Score = 122 bits (306), Expect(2) = 3e-67
Identities = 60/71 (84%), Positives = 61/71 (85%)
Frame = +1
Query: 92 SADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKNGRIASKQN 151
S DILKLGFRFKQDLVYLSSTFC GC PG DKVEPYLDITSVYN LQ KK GRIASKQ
Sbjct: 274 SPDILKLGFRFKQDLVYLSSTFCAHGCEPGIDKVEPYLDITSVYNQLQPKKQGRIASKQT 453
Query: 152 KSLSTICGELL 162
KS STICGE+L
Sbjct: 454 KSSSTICGEVL 486
>AW719410
Length = 538
Score = 47.0 bits (110), Expect = 8e-06
Identities = 22/47 (46%), Positives = 33/47 (69%)
Frame = +3
Query: 154 LSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFK 200
L+ + ++LG L+K + S+W QRPLTE Q+ YAA+DA L+ IF+
Sbjct: 39 LAGLTEKILGARLNKTRRNSNWEQRPLTENQLEYAALDAVVLIHIFR 179
>TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protein
(OJ1118_G09.13 protein), partial (6%)
Length = 689
Score = 37.0 bits (84), Expect = 0.008
Identities = 48/192 (25%), Positives = 79/192 (41%), Gaps = 11/192 (5%)
Frame = +3
Query: 12 HFVTTTDSPEFTHLTRSLTQTS---------LVGLDAEWKPVRTHQNSFPTVSLLQIACQ 62
H + T + + +H+ L++T+ +VGLD EW+P + P V+LLQ+
Sbjct: 81 HTIQTLFTSDPSHVDSWLSETTRHRNNKLPFIVGLDIEWRPNTQAFKNNP-VALLQLCV- 254
Query: 63 LGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGF 122
D + +I P SI + L L + +G + D+ L N
Sbjct: 255 ---DHRCLVFQIIHAP--SIPDSLSSFLSNPQHTFVGVGIQGDVDKLLKDRSFTVANA-- 413
Query: 123 DKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSK--ELQCSDWSQRPL 180
V+ VY + K G L + +LG+ + K ++ S W R L
Sbjct: 414 --VDLRTLAAEVYGDPEMMKAG---------LKALTQRVLGMNVEKPKKISTSKWDDRYL 560
Query: 181 TEEQMTYAAMDA 192
+ EQ+ YAA+DA
Sbjct: 561 SVEQVQYAAIDA 596
>TC16552 similar to UP|Q94IE2 (Q94IE2) APG1, partial (31%)
Length = 433
Score = 31.2 bits (69), Expect = 0.43
Identities = 19/66 (28%), Positives = 28/66 (41%), Gaps = 15/66 (22%)
Frame = +1
Query: 461 IEAAKGFQKIPNCLFNKNLEFWQCMDCHQLYWEVLFTCS---------------VCCELL 505
+ + K + + N LF+ FW+ H LYW LFTC V CE
Sbjct: 172 VRSKKMLKSL*NLLFSSCAFFWESWLLHGLYW-FLFTCGSKIRLFRNLSQSKMFVVCETF 348
Query: 506 FTIALS 511
+T++ S
Sbjct: 349 YTVSCS 366
>AV780895
Length = 445
Score = 30.8 bits (68), Expect = 0.57
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = -1
Query: 475 FNKNLEFWQCMDCHQLYWEVLFTCS 499
FN FW CM+ QL+W+++ C+
Sbjct: 145 FNSVGLFWSCMNIFQLFWKIVVLCT 71
>BI420393
Length = 515
Score = 28.5 bits (62), Expect = 2.8
Identities = 18/52 (34%), Positives = 23/52 (43%), Gaps = 7/52 (13%)
Frame = -3
Query: 447 NGRFIQKPLSTEEAIEAAK---GFQKIPNCLFNKNLEFWQ----CMDCHQLY 491
NGR +Q PL T EAA+ I N +FW+ C CH L+
Sbjct: 414 NGRIMQHPLPTPPKGEAARNSYSTNSIKPVSILHNSKFWKPFIICFSCHLLF 259
>AU088851
Length = 663
Score = 28.1 bits (61), Expect = 3.7
Identities = 16/48 (33%), Positives = 25/48 (51%)
Frame = +3
Query: 52 PTVSLLQIACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLG 99
P L A ++ D+ +FL SLP ++W ++E+L S D K G
Sbjct: 387 PPSPRLSFASRMWADKTLFL----SLPSYNLWLLIKELLASLDFXKQG 518
>TC18875 similar to UP|CTNS_ARATH (P57758) Cystinosin homolog, partial (58%)
Length = 567
Score = 27.7 bits (60), Expect = 4.8
Identities = 17/52 (32%), Positives = 23/52 (43%), Gaps = 1/52 (1%)
Frame = +3
Query: 463 AAKGFQKIPNCLFNKNLEFWQCMDCHQLYWEVLFTCSVCCEL-LFTIALSFI 513
+A G IP+C NL W+ YW L + CC L L+ SF+
Sbjct: 354 SADGDYIIPDC----NL*TWKSESVQNFYWNCLCCLADCCNLFLYCFV*SFL 497
>TC19820 similar to GB|AAO63279.1|28950711|BT005215 At3g56740 {Arabidopsis
thaliana;}, partial (23%)
Length = 571
Score = 26.9 bits (58), Expect = 8.2
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 1/25 (4%)
Frame = -3
Query: 443 CTK-CNGRFIQKPLSTEEAIEAAKG 466
CTK C F+ KP+ T AIE++ G
Sbjct: 215 CTKACLAEFLSKPIETRVAIESSDG 141
>TC14194 weakly similar to UP|Q8GT67 (Q8GT67) Xyloglucan-specific fungal
endoglucanase inhibitor protein precursor, partial (11%)
Length = 892
Score = 26.9 bits (58), Expect = 8.2
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +2
Query: 475 FNKNLEFWQCMDCHQLYWEVLFTCSVCCELLFTIALSFILNKC 517
F K+ W+ D H +WE FTC C + + L IL++C
Sbjct: 632 FFKHQRHWE--DLH--WWETKFTCQCCQNWICSDIL*RILHQC 748
>BP060196
Length = 317
Score = 26.9 bits (58), Expect = 8.2
Identities = 14/33 (42%), Positives = 21/33 (63%)
Frame = +2
Query: 75 ISLPLSSIWEPLREMLVSADILKLGFRFKQDLV 107
I + +SS WEPL+ VS+D L L F +D++
Sbjct: 53 IYIKVSSTWEPLKVSRVSSDSLSL---FWEDII 142
>AV413925
Length = 402
Score = 26.9 bits (58), Expect = 8.2
Identities = 14/42 (33%), Positives = 18/42 (42%)
Frame = +2
Query: 463 AAKGFQKIPNCLFNKNLEFWQCMDCHQLYWEVLFTCSVCCEL 504
+A G IP+C NL W+ YW L + CC L
Sbjct: 260 SADGDYIIPDC----NL*TWKSESVQNFYWNCLCCLADCCNL 373
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,745,929
Number of Sequences: 28460
Number of extensions: 144661
Number of successful extensions: 768
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 765
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 766
length of query: 523
length of database: 4,897,600
effective HSP length: 94
effective length of query: 429
effective length of database: 2,222,360
effective search space: 953392440
effective search space used: 953392440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC141323.9


