

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
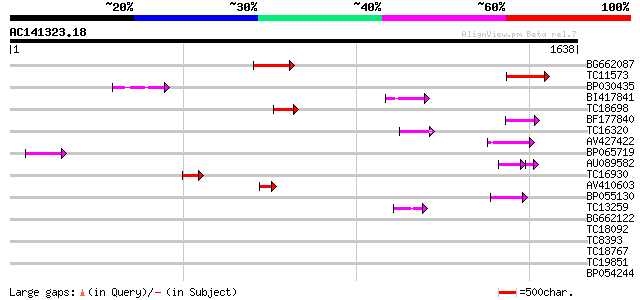
Query= AC141323.18 - phase: 0
(1638 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662087 164 1e-40
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 120 2e-27
BP030435 113 3e-25
BI417841 72 9e-13
TC18698 68 1e-11
BF177840 60 2e-09
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 58 1e-08
AV427422 54 2e-07
BP065719 52 1e-06
AU089582 48 1e-06
TC16930 49 8e-06
AV410603 47 2e-05
BP055130 47 3e-05
TC13259 42 6e-04
BG662122 42 0.001
TC18092 33 0.36
TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenyla... 32 1.1
TC18767 32 1.1
TC19851 similar to UP|Q9SKK6 (Q9SKK6) At2g25430 protein (At2g254... 31 1.4
BP054244 31 1.8
>BG662087
Length = 373
Score = 164 bits (414), Expect = 1e-40
Identities = 75/120 (62%), Positives = 95/120 (78%)
Frame = +1
Query: 703 SGKWRMCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAP 762
SGKWRM VDYTDLN ACPKD YPLPSID L+D AS + LS MDAYSGY+QIKM P D
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 763 KTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSH 822
KTAFMT + NY Y+ + FGL+NAGAT+Q MD +F + +GRN+EVY+D+++VK++ + +H
Sbjct: 193 KTAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 120 bits (301), Expect = 2e-27
Identities = 57/124 (45%), Positives = 81/124 (64%)
Frame = +2
Query: 1435 DNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDAKGLWAE 1494
+NG QF S + +FC +GI+ +F SV HPQ NGQ E+ANK+I+ IK++L +A+G W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1495 QLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEIDTPTWRREHFSEESNEVGIRCTMD 1554
+L VLWSY+T P S+ ETPF + YGAD MLPVEI +W + EE N + + +
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEISNFSW*TKARHEEENRMNMAVGWE 367
Query: 1555 MIDE 1558
+ E
Sbjct: 368 KLPE 379
>BP030435
Length = 533
Score = 113 bits (282), Expect = 3e-25
Identities = 67/171 (39%), Positives = 98/171 (57%), Gaps = 5/171 (2%)
Frame = +1
Query: 296 SKTMGQDKNAWCKYHLIEGHNTDDCVHLKREIEKLLLNGKLRGYAKEKHHSERREDKPNP 355
++ G+ C YH GH T++C L+RE+ KL+ GK R + P
Sbjct: 70 TRISGEPGGE*CVYHQAMGHITEECRTLQREVGKLIATGK----------PTRIQGGNAP 219
Query: 356 EPKHTLHTISGGFAGGGESSNLRKKYVRQV-----MLLGDSPSSPKKYPHVTFSSEDFGR 410
PK +HTI+GGF GGG +S RK+Y R V +L G S +P +TFSS+DF R
Sbjct: 220 VPKG-VHTIAGGFGGGGITSAARKRYARAVNTVTEVLFGFS------HPVITFSSDDFIR 378
Query: 411 VLPHDDDPLVISVQLFNWEIKRVLIDTGSSADVLYFDAFSKMGLSEEQLQP 461
+ PH D+P+VI +++ ++RVL+D GSSAD++Y F ++GL+E L P
Sbjct: 379 IKPHLDNPIVILLRVNQLNVQRVLLDQGSSADIIYGGVFDRLGLNEADLTP 531
>BI417841
Length = 617
Score = 71.6 bits (174), Expect = 9e-13
Identities = 43/133 (32%), Positives = 70/133 (52%), Gaps = 4/133 (3%)
Frame = +1
Query: 1085 LSVDGSS--NIKGSGAGIIL--EGPGDLLIEQSLKFDFKASNNQAEYEALIAGMLLAQEM 1140
L DGSS N +GAG +L E + + + + +NNQAEY LI G+ A E
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVG---NQTNNQAEYRGLILGLKHAHEQ 312
Query: 1141 GAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFFEAIYVPRESNSRAD 1200
G +++ + DSQL+ Q+ G ++ ++ ++ ++L FQ F+ +VPR+ NS AD
Sbjct: 313 GYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEAD 492
Query: 1201 LLAKLASTKKPGN 1213
+ A L G+
Sbjct: 493 VQANLGVNLPAGH 531
>TC18698
Length = 808
Score = 67.8 bits (164), Expect = 1e-11
Identities = 32/71 (45%), Positives = 46/71 (64%)
Frame = -2
Query: 763 KTAFMTNQKNYHYRVMSFGLRNAGATFQRSMDTIFSNQIGRNLEVYIDDLVVKTSEKQSH 822
KT N+ NY+Y+VM GL+N T+QR MD IF QI +N+EVY++D++VK+S++ H
Sbjct: 801 KTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
Query: 823 SVDLKEIFQQI 833
DL I
Sbjct: 621 RGDLSRDLSDI 589
>BF177840
Length = 410
Score = 60.5 bits (145), Expect = 2e-09
Identities = 30/99 (30%), Positives = 53/99 (53%)
Frame = +2
Query: 1432 IVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDAKGL 1491
IV+D T+F S ++G + + + HPQ +GQ E NK + ++ LE +
Sbjct: 17 IVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLKM 196
Query: 1492 WAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPVEI 1530
W L + ++Y+ HSTT +PF +VYG + + P+++
Sbjct: 197 WETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 57.8 bits (138), Expect = 1e-08
Identities = 30/99 (30%), Positives = 54/99 (54%)
Frame = +2
Query: 1127 YEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQQLSKYLARVRKLAGDFQFF 1186
Y LI G+ A + G K+++ + DS L+ NQ+ G ++ K+Q ++ + ++L F F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1187 EAIYVPRESNSRADLLAKLASTKKPGNNRTVIQEVISAP 1225
+ ++PRE NS AD+ A + + G ++EV P
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQ----VEEVREVP 286
>AV427422
Length = 417
Score = 54.3 bits (129), Expect = 2e-07
Identities = 35/136 (25%), Positives = 67/136 (48%), Gaps = 1/136 (0%)
Frame = +1
Query: 1381 PQAPGQLKFLIVDVDYFTKWVEAEAVSK-ITAERVVKFYWKKIICHFGLPKYIVTDNGTQ 1439
P++ G L+V VD +K+ + TA+ + + ++++ G+P IV+D
Sbjct: 13 PKSKGYEAVLVV-VDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPL 189
Query: 1440 FASSKVVNFCKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDAKGLWAEQLHEV 1499
F S+ K G + K + HP+++GQ E N+ + ++ + D WA +
Sbjct: 190 FMSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWA 369
Query: 1500 LWSYHTTPHSTTGETP 1515
+ Y+T+ H +TG+TP
Sbjct: 370 EYWYNTSYHVSTGQTP 417
>BP065719
Length = 567
Score = 51.6 bits (122), Expect = 1e-06
Identities = 34/120 (28%), Positives = 56/120 (46%), Gaps = 2/120 (1%)
Frame = +3
Query: 46 FSTEIWNAPVPDNFKPPQLPTFDGRSDPS--EHVTVFNTRMSVYGVADSLKCKLLAGTLA 103
F E+ + +P +K P+ F G S S EH+ + + ++LK K +L
Sbjct: 135 FIEEVLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLT 314
Query: 104 DAALRWYMSLPRFSIVGYQDLTKKFTQQFSGTRHRKVLSTSLFNVRQGPNESLREYLARF 163
A W+ +L S+ + L + F +QF KV L +V++ P ES+ +YL RF
Sbjct: 315 KNAFTWFTTLAPRSVHTWAQLERIFHEQFF-RGECKVSXKDLASVKRKPAESIDDYLNRF 491
>AU089582
Length = 383
Score = 47.8 bits (112), Expect(2) = 1e-06
Identities = 24/80 (30%), Positives = 42/80 (52%)
Frame = +2
Query: 1411 AERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNFCKQLGIETKFVSVIHPQANGQA 1470
A + K Y +I+ G+P I++D G QF S +F LG K + HPQ +GQ+
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 1471 ESANKMIVNDIKKKLEDAKG 1490
E +++ + ++ + D +G
Sbjct: 191 ERTIQILEDMLRACVXDLRG 250
Score = 23.1 bits (48), Expect(2) = 1e-06
Identities = 10/40 (25%), Positives = 20/40 (50%)
Frame = +3
Query: 1489 KGLWAEQLHEVLWSYHTTPHSTTGETPFTMVYGADAMLPV 1528
+G W + L + ++Y+ + S+ PF +YG P+
Sbjct: 246 EGSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPI 365
>TC16930
Length = 520
Score = 48.5 bits (114), Expect = 8e-06
Identities = 20/60 (33%), Positives = 37/60 (61%)
Frame = +1
Query: 499 YLVINSPSSYNVIIGRPSINLLDAFVSTKHLLMKYPLDSGRVGVVHGDQKIARECYHASL 558
+++ N+ +SYN I+ PS+N+ D + T+ L K+ D+ + + GDQ++A CY S+
Sbjct: 10 FMLANNLTSYNAILC*PSLNVFDMCILTRRLASKFSTDNTEIVTIQGDQEVAPNCYIMSM 189
>AV410603
Length = 162
Score = 47.4 bits (111), Expect = 2e-05
Identities = 21/49 (42%), Positives = 31/49 (62%)
Frame = +1
Query: 721 KDPYPLPSIDHLIDNASGYKTLSFMDAYSGYNQIKMDPLDAPKTAFMTN 769
KD +P+P++D L+D G + S +D SGY+QI + P D KT F T+
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTH 156
>BP055130
Length = 567
Score = 46.6 bits (109), Expect = 3e-05
Identities = 31/108 (28%), Positives = 52/108 (47%), Gaps = 1/108 (0%)
Frame = +2
Query: 1390 LIVDVDYFTKWVE-AEAVSKITAERVVKFYWKKIICHFGLPKYIVTDNGTQFASSKVVNF 1448
+IV VD +K+ A + T+ +V + + I+ G+P+ IV+D F S+ +F
Sbjct: 191 IIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHF 370
Query: 1449 CKQLGIETKFVSVIHPQANGQAESANKMIVNDIKKKLEDAKGLWAEQL 1496
K G S HPQ +GQ E+ NK + ++ + + LW L
Sbjct: 371 FKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYL 514
>TC13259
Length = 506
Score = 42.4 bits (98), Expect = 6e-04
Identities = 31/99 (31%), Positives = 50/99 (50%)
Frame = +1
Query: 1108 LLIEQSLKFDFKASNNQAEYEALIAGMLLAQEMGAKNLRARSDSQLMTNQISGEYQTKDQ 1167
+L S K+ S +E AL G+ L+ E+G + D+Q++ N G +
Sbjct: 58 VLATASSKWPRTVSVATSEAMALRWGLQLSLELGFFEVEVEIDNQVVVNSWKGT-KKHVS 234
Query: 1168 QLSKYLARVRKLAGDFQFFEAIYVPRESNSRADLLAKLA 1206
L+ +A V +++ F+FF YVPR S+ AD LA+ A
Sbjct: 235 YLANVIADVVRISQCFRFFSLAYVPRLSHLAADYLAQFA 351
>BG662122
Length = 386
Score = 41.6 bits (96), Expect = 0.001
Identities = 28/86 (32%), Positives = 40/86 (45%), Gaps = 6/86 (6%)
Frame = +2
Query: 271 LNTRPEKILKEVFESKIIP------PPPFNRSKTMGQDKNAWCKYHLIEGHNTDDCVHLK 324
LN IL++V + ++ PPP G D C+Y H+ DDC LK
Sbjct: 101 LNAHLTDILQDVKAAHMVGKSGQS*PPP-----RRGIDTTK*CEYRRSVVHDIDDCFTLK 265
Query: 325 REIEKLLLNGKLRGYAKEKHHSERRE 350
REIEKL+ G+L+ Y + R+
Sbjct: 266 REIEKLIKMGRLKQYDRGSRQQGDRQ 343
>TC18092
Length = 519
Score = 33.1 bits (74), Expect = 0.36
Identities = 17/35 (48%), Positives = 23/35 (65%)
Frame = -2
Query: 708 MCVDYTDLNMACPKDPYPLPSIDHLIDNASGYKTL 742
MCV TD+N CP+ PL SI+ L+D +S Y+ L
Sbjct: 101 MCVC*TDINKDCPR--XPLTSINALVDYSSSYEYL 3
>TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenylated Rab
receptor 2 {Arabidopsis thaliana;}, partial (78%)
Length = 1012
Score = 31.6 bits (70), Expect = 1.1
Identities = 22/92 (23%), Positives = 42/92 (44%), Gaps = 9/92 (9%)
Frame = -3
Query: 158 EYLARFNDSTIKVSNPNQEVFVGAFQNGLRAGQFNESLAQKPADS---------MEEIIA 208
++ +R+ +ST+ + +P+ R G A++ +S EEI+
Sbjct: 782 DFTSRYQNSTLPLPDPSDPQSNATGDGRNRGGGDGSRSAEEGEESGGLRGLLLIKEEILR 603
Query: 209 RAECYVKGEESNAEKRARDVKEKGSSGGERRN 240
AEC ++ + +AE +RD E G+ GE +
Sbjct: 602 DAECAMEANKGDAEHESRD--EDGTDTGEEND 513
>TC18767
Length = 1004
Score = 31.6 bits (70), Expect = 1.1
Identities = 27/97 (27%), Positives = 42/97 (42%), Gaps = 5/97 (5%)
Frame = +2
Query: 337 RGYAKEKHHSERREDKPNPEPKH-----TLHTISGGFAGGGESSNLRKKYVRQVMLLGDS 391
RG+ +E+ + R+D P +P H + H GG SS+ + R L D+
Sbjct: 713 RGHYREQRLGDLRDDGPPGDPGHSSLQYSFHPRFGGHESIPRSSSAGRSQDRNRSPLHDA 892
Query: 392 PSSPKKYPHVTFSSEDFGRVLPHDDDPLVISVQLFNW 428
SP+ + + GR+ P D D S +L NW
Sbjct: 893 -ESPRPFSFHSLHYTSSGRLSPMDRD----SGRLENW 988
>TC19851 similar to UP|Q9SKK6 (Q9SKK6) At2g25430 protein
(At2g25430/F13B15.9), partial (30%)
Length = 642
Score = 31.2 bits (69), Expect = 1.4
Identities = 21/73 (28%), Positives = 27/73 (36%), Gaps = 3/73 (4%)
Frame = +1
Query: 40 EPEPQPFSTEIWNAPVPDNFKPPQLPTFDGRSDPSEHVTVFNTRMSVYGVADS---LKCK 96
E EP P EI P P+N+ PP P + + P + N R D
Sbjct: 13 EEEPVPDMNEIKALPPPENYTPPPQPEPEPKPQPQVQEDLVNLREDAVTADDQGNRFAXA 192
Query: 97 LLAGTLADAALRW 109
L AG A+ W
Sbjct: 193 LFAGAPANGNGSW 231
>BP054244
Length = 465
Score = 30.8 bits (68), Expect = 1.8
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Frame = -1
Query: 148 VRQGPNESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLRAG-QFNESLAQKPADSMEE 205
++QG ESLR++L +F V++ + +V + +R G F S A P+ M+E
Sbjct: 453 LKQGKKESLRDFLTKFYKEATLVTSLDLKVRLHLLCEAIRMGTSFYTSSAXTPSQDMDE 277
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,762,223
Number of Sequences: 28460
Number of extensions: 366664
Number of successful extensions: 1824
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1822
length of query: 1638
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1535
effective length of database: 1,966,220
effective search space: 3018147700
effective search space used: 3018147700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC141323.18


