

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
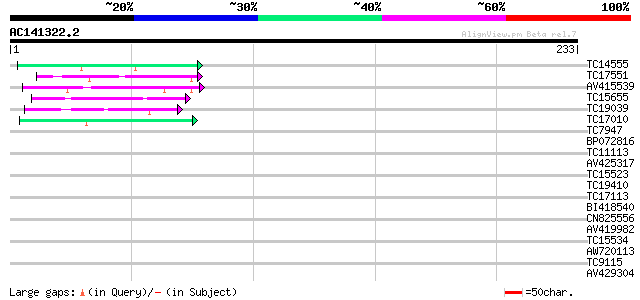
Query= AC141322.2 - phase: 0
(233 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 41 2e-04
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 40 3e-04
AV415539 40 5e-04
TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein... 40 5e-04
TC19039 similar to UP|Q8L7W5 (Q8L7W5) AT4g12110/F16J13_180, part... 39 8e-04
TC17010 similar to UP|BAC84225 (BAC84225) RRM-containing RNA-bin... 39 8e-04
TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, parti... 39 0.001
BP072816 38 0.002
TC11113 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich... 37 0.003
AV425317 37 0.003
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 37 0.003
TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial... 37 0.004
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 37 0.004
BI418540 36 0.005
CN825556 36 0.005
AV419982 36 0.007
TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28... 36 0.007
AW720113 36 0.007
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 35 0.011
AV429304 33 0.055
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 41.2 bits (95), Expect = 2e-04
Identities = 27/81 (33%), Positives = 32/81 (39%), Gaps = 5/81 (6%)
Frame = +2
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNI---PPLQQQPLPLTLNIPQPSHPPP--PPPQDVVD 58
S P QS P P TP + P + PP+Q P P + P P+ PPP PPP
Sbjct: 284 SPPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPP 463
Query: 59 NHAPVRSRRRGPPRNGPIPTP 79
A P P P P
Sbjct: 464 PAATPPPALTPVPATSPAPAP 526
Score = 37.7 bits (86), Expect = 0.002
Identities = 28/78 (35%), Positives = 34/78 (42%), Gaps = 4/78 (5%)
Frame = +2
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPP----PPPQDVVDN 59
S PT S P T Q A + PP Q P P+ + P S PPP PPP V +
Sbjct: 194 SNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPP---VSS 364
Query: 60 HAPVRSRRRGPPRNGPIP 77
PV+S P + P P
Sbjct: 365 PPPVQSTPPPAPASTPPP 418
Score = 35.0 bits (79), Expect = 0.011
Identities = 24/74 (32%), Positives = 29/74 (38%), Gaps = 3/74 (4%)
Frame = +2
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPP---PPPQDVVDNHAPV 63
PT + P P A + PP+Q P P + P S PPP PPP AP
Sbjct: 224 PTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSPPPVQSTPPPAPA 403
Query: 64 RSRRRGPPRNGPIP 77
+ PP P P
Sbjct: 404 ST----PPPASPPP 433
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 40.4 bits (93), Expect = 3e-04
Identities = 29/71 (40%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Frame = +1
Query: 12 PLPMLTPFQRMFAHPNIPPL-QQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGP 70
P P L P F P +PPL PLP L P P PPPPPP + +R R P
Sbjct: 169 PRPTLFP---PFPLPRLPPLVPPPPLPRLLRPPPP--PPPPPPLRPRPHRHRLRPLLRHP 333
Query: 71 PRN--GPIPTP 79
PR P+P P
Sbjct: 334 PRRQMEPLPLP 366
Score = 28.9 bits (63), Expect = 0.80
Identities = 21/80 (26%), Positives = 26/80 (32%), Gaps = 4/80 (5%)
Frame = +3
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPP----PPPPQDVVDN 59
+ P SLP P + P P + P PS PP PPPP +
Sbjct: 171 ASHPLPSLPSPSSAAVGPTSSTPPTSPTSAAAATSSATPPSPSPPPTSASPPPPSSTPNG 350
Query: 60 HAPVRSRRRGPPRNGPIPTP 79
PP + P TP
Sbjct: 351 T---------PPSSSPATTP 383
Score = 22.7 bits (47), Expect(2) = 4.4
Identities = 12/38 (31%), Positives = 16/38 (41%), Gaps = 7/38 (18%)
Frame = +3
Query: 48 PPPPPPQDVVDNHAP-------VRSRRRGPPRNGPIPT 78
PPPPPP N+ P S P + P+P+
Sbjct: 78 PPPPPPSPNHRNNLPPPFPPSLTTSFSTSSPASHPLPS 191
Score = 21.9 bits (45), Expect(2) = 4.4
Identities = 7/10 (70%), Positives = 8/10 (80%)
Frame = +3
Query: 44 QPSHPPPPPP 53
Q + PPPPPP
Sbjct: 63 QTTPPPPPPP 92
>AV415539
Length = 419
Score = 39.7 bits (91), Expect = 5e-04
Identities = 30/90 (33%), Positives = 37/90 (40%), Gaps = 15/90 (16%)
Frame = +3
Query: 6 QPTQSLPLPMLTPFQRM--FAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAP- 62
+P + L ++ P R HP PP PLPL P P HP PPP + H P
Sbjct: 3 EPQPLMGLHVVLPLHRHSHLPHPLPPP---PPLPLAPPPPPPPHPTRPPPGNPQPLHVPH 173
Query: 63 --------VRSRRRGPPRN----GPIPTPF 80
R RRR PR+ P+ PF
Sbjct: 174LRRYFHRNTRLRRRRDPRHALVLAPLQNPF 263
>TC15655 similar to UP|Q8LE99 (Q8LE99) Transketolase-like protein, partial
(17%)
Length = 575
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/65 (43%), Positives = 31/65 (47%)
Frame = +1
Query: 10 SLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRG 69
+LPLP+ P P PPL PLP L P P H P P P NH+P SRRR
Sbjct: 46 NLPLPLSLPSPPHA--PLFPPLLHAPLPPRLLRPPPIHHPLPLPLP-HSNHSPPPSRRRQ 216
Query: 70 PPRNG 74
R G
Sbjct: 217 SLRCG 231
Score = 26.2 bits (56), Expect = 5.2
Identities = 16/50 (32%), Positives = 19/50 (38%)
Frame = +3
Query: 29 PPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPT 78
P L QP P +L+ S P P P P + PP P PT
Sbjct: 30 PWLLPQPPPPSLSPKPSSRAPLPSPPPRASPSPPSPASANPPPSPSPSPT 179
>TC19039 similar to UP|Q8L7W5 (Q8L7W5) AT4g12110/F16J13_180, partial (61%)
Length = 763
Score = 38.9 bits (89), Expect = 8e-04
Identities = 29/68 (42%), Positives = 31/68 (44%), Gaps = 3/68 (4%)
Frame = +3
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDV---VDNHAPV 63
P SLPLP L P P P Q PLPL + P+ P PP V D HA V
Sbjct: 267 PQHSLPLPHLLPRPS----PLRLPRDQTPLPLR-RLQDPTQGSPLPP*HVRLLQDRHAHV 431
Query: 64 RSRRRGPP 71
RRR PP
Sbjct: 432 LPRRRSPP 455
>TC17010 similar to UP|BAC84225 (BAC84225) RRM-containing RNA-binding
protein-like, partial (4%)
Length = 569
Score = 38.9 bits (89), Expect = 8e-04
Identities = 24/77 (31%), Positives = 30/77 (38%), Gaps = 4/77 (5%)
Frame = +1
Query: 5 EQPTQSLPLPMLTPFQRMFAHPNIPP----LQQQPLPLTLNIPQPSHPPPPPPQDVVDNH 60
+QP Q P PM+ P + H P QQP+P P P H P Q NH
Sbjct: 58 KQPMQHYPRPMMPPPPGQYHHHQYYPPYGGYMQQPVPPYQQYPPPYHAVVAPSQPPAANH 237
Query: 61 APVRSRRRGPPRNGPIP 77
S + G + G P
Sbjct: 238 PYQHSMQPGSSQTGSAP 288
>TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, partial (81%)
Length = 1294
Score = 38.5 bits (88), Expect = 0.001
Identities = 27/92 (29%), Positives = 37/92 (39%), Gaps = 18/92 (19%)
Frame = +2
Query: 2 NDSEQPTQSLPLPMLTPFQRMFAH----------------PNIPPLQQQP--LPLTLNIP 43
+D QPT S + +L F + H P+ P QP P +L+ P
Sbjct: 38 SDPTQPTSSFRISLLPSFLQWLLHLL*YPPLLSPLSPSAPPHPSPSSPQPPNSPSSLSSP 217
Query: 44 QPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGP 75
PS PPPPPP + + P PP + P
Sbjct: 218 DPSSPPPPPPPSPPNANPPTTLT---PPSSTP 304
>BP072816
Length = 492
Score = 37.7 bits (86), Expect = 0.002
Identities = 20/43 (46%), Positives = 24/43 (55%), Gaps = 3/43 (6%)
Frame = +2
Query: 25 HPNIPPLQQQPLP-LTLNIPQPSHP--PPPPPQDVVDNHAPVR 64
H P QQ+P P L L +P P HP PPPPP + H P+R
Sbjct: 182 HHRPPRRQQRPHPHLRLQLPLPIHPPSPPPPPSRLPPPHLPLR 310
Score = 26.2 bits (56), Expect = 5.2
Identities = 18/55 (32%), Positives = 21/55 (37%), Gaps = 6/55 (10%)
Frame = +1
Query: 29 PPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAP------VRSRRRGPPRNGPIP 77
PPL QP + S P P PP + HAP S PP + P P
Sbjct: 85 PPLSHQPCHSPSSSSSSSSPLPSPPTTL---HAPSTSPSSAPSATAAPPPSTPTP 240
>TC11113 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (10%)
Length = 309
Score = 37.0 bits (84), Expect = 0.003
Identities = 25/75 (33%), Positives = 31/75 (41%), Gaps = 6/75 (8%)
Frame = +1
Query: 11 LPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGP 70
LP P+ P + P+ P + PLP P PS P PPP +H P S P
Sbjct: 70 LPFPISAPLRSQ--PPSTPAVHHSPLPYLP--PSPSLPNSPPPPPRSGHHHPSPSEPPPP 237
Query: 71 PRNGP------IPTP 79
P+ P PTP
Sbjct: 238 PQTTPPFTNPQTPTP 282
>AV425317
Length = 398
Score = 37.0 bits (84), Expect = 0.003
Identities = 22/79 (27%), Positives = 33/79 (40%)
Frame = +1
Query: 6 QPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRS 65
+P +LPLP+ PF + + P + P+ P+ PPPP AP R
Sbjct: 151 KPHHNLPLPLPPPFPNL-TPSSAPSFKSLPIVTAAFTGSPTSSAPPPPSSSTVPSAPARF 327
Query: 66 RRRGPPRNGPIPTPFIWAT 84
++ PP + P AT
Sbjct: 328 SQQTPP*SSTFSRPTSPAT 384
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 37.0 bits (84), Expect = 0.003
Identities = 28/75 (37%), Positives = 32/75 (42%), Gaps = 5/75 (6%)
Frame = -1
Query: 10 SLPLPMLT-----PFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVR 64
SL LP L P R+ P PP ++ P P P P PPPPPP + P
Sbjct: 819 SLQLPPLE*RDPPPPSRLPP*PPPPPSRRPPYPPP---PLPP*PPPPPPL*RLPPPPPYP 649
Query: 65 SRRRGPPRNGPIPTP 79
RR PP P P P
Sbjct: 648 PSRRPPPYPPPPPPP 604
Score = 27.7 bits (60), Expect = 1.8
Identities = 20/64 (31%), Positives = 27/64 (41%), Gaps = 12/64 (18%)
Frame = -1
Query: 3 DSEQPTQSLPLPMLTPFQRM-FAHPNIPPLQQQPLPLTLNIPQPSHP-----------PP 50
D P++ P P P +R + P +PP P PL P P +P PP
Sbjct: 789 DPPPPSRLPP*PPPPPSRRPPYPPPPLPP*PPPPPPL*RLPPPPPYPPSRRPPPYPPPPP 610
Query: 51 PPPQ 54
PPP+
Sbjct: 609 PPPR 598
>TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial (28%)
Length = 606
Score = 36.6 bits (83), Expect = 0.004
Identities = 20/63 (31%), Positives = 23/63 (35%), Gaps = 13/63 (20%)
Frame = +1
Query: 30 PLQQQPLPLTLNIPQPSHPPPPPP-------------QDVVDNHAPVRSRRRGPPRNGPI 76
PL P P TL P+ PPPPP Q H PV + PP +
Sbjct: 55 PLSHSPSP*TLKTSTPASSPPPPPPPISTKPVHSLSQQPPTQTHFPVSTHTLPPPSESHL 234
Query: 77 PTP 79
P P
Sbjct: 235 PNP 243
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 36.6 bits (83), Expect = 0.004
Identities = 25/66 (37%), Positives = 30/66 (44%), Gaps = 1/66 (1%)
Frame = +1
Query: 7 PTQSLPLPMLT-PFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRS 65
P Q P P+L PFQ P P Q P P L IP P PPP PP + + P S
Sbjct: 511 PFQPSPPPLLPDPFQPPSPPPLFPNPFQPPSPPPL-IPNPFQPPPSPPPFIPNPFQPPPS 687
Query: 66 RRRGPP 71
++ P
Sbjct: 688 KQPPSP 705
Score = 33.1 bits (74), Expect = 0.042
Identities = 22/70 (31%), Positives = 28/70 (39%), Gaps = 1/70 (1%)
Frame = +1
Query: 12 PLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPP 71
P+P P + +P PPL P IP P PPP P + P+ PP
Sbjct: 388 PIPFFPP--PLIPNPFQPPLVPNPFQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPP 561
Query: 72 RNGPI-PTPF 80
P+ P PF
Sbjct: 562 SPPPLFPNPF 591
Score = 31.6 bits (70), Expect = 0.12
Identities = 28/81 (34%), Positives = 33/81 (40%), Gaps = 7/81 (8%)
Frame = +1
Query: 7 PTQSLPLPML-TPFQ-----RMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNH 60
P P P++ PFQ F P + P QP PL N QPS PP P +
Sbjct: 388 PIPFFPPPLIPNPFQPPLVPNPFQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSP 567
Query: 61 APVRSRRRGPPRNGP-IPTPF 80
P+ PP P IP PF
Sbjct: 568 PPLFPNPFQPPSPPPLIPNPF 630
Score = 29.6 bits (65), Expect = 0.47
Identities = 22/55 (40%), Positives = 25/55 (45%), Gaps = 7/55 (12%)
Frame = +1
Query: 6 QPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPL----TLNIP---QPSHPPPPPP 53
QP S P + PFQ PP +Q P PL + IP P PPPPPP
Sbjct: 631 QPPPSPPPFIPNPFQP-------PPSKQPPSPLFPFPPIVIPGLTPPPPPPPPPP 774
>BI418540
Length = 518
Score = 36.2 bits (82), Expect = 0.005
Identities = 24/79 (30%), Positives = 30/79 (37%), Gaps = 12/79 (15%)
Frame = +1
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPS------------HPPPP 51
S P S+P P + + PN P P PL+L IP P+ H PPP
Sbjct: 133 SPNPPPSVPSPHSPSSLKNPSSPNPPLTPTPPPPLSLPIPPPAALFTARIGGTPIHAPPP 312
Query: 52 PPQDVVDNHAPVRSRRRGP 70
PP + P R P
Sbjct: 313 PPAPSPPSATPAAPHCRNP 369
Score = 26.2 bits (56), Expect = 5.2
Identities = 19/61 (31%), Positives = 23/61 (37%), Gaps = 7/61 (11%)
Frame = +1
Query: 26 PNIPPLQQQPL-------PLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPT 78
PN PP P P + N P PPPP + A +R G P + P P
Sbjct: 136 PNPPPSVPSPHSPSSLKNPSSPNPPLTPTPPPPLSLPIPPPAALFTARIGGTPIHAPPPP 315
Query: 79 P 79
P
Sbjct: 316 P 318
>CN825556
Length = 655
Score = 36.2 bits (82), Expect = 0.005
Identities = 23/66 (34%), Positives = 30/66 (44%)
Frame = +1
Query: 14 PMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRN 73
P+L P Q + P++P Q QP P PQP P P PQ P S + P
Sbjct: 418 PVLPPPQPVPPPPSLPQPQPQPQPQPQPQPQPQPQPQPQPQPQPQPVPPPASLPQ--PLL 591
Query: 74 GPIPTP 79
P+P+P
Sbjct: 592 APLPSP 609
Score = 28.5 bits (62), Expect = 1.0
Identities = 16/46 (34%), Positives = 19/46 (40%)
Frame = +1
Query: 9 QSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQ 54
Q P P P + P +PP P PL +P P P PPQ
Sbjct: 502 QPQPQPQPQPQPQPQPQP-VPPPASLPQPLLAPLPSPGLPEQKPPQ 636
Score = 26.9 bits (58), Expect = 3.0
Identities = 17/58 (29%), Positives = 24/58 (41%), Gaps = 2/58 (3%)
Frame = +1
Query: 26 PNIPPLQQQPLPLTLNIPQPSHP--PPPPPQDVVDNHAPVRSRRRGPPRNGPIPTPFI 81
P +PP Q P P +L PQP P P PQ + + + P +P P +
Sbjct: 418 PVLPPPQPVPPPPSLPQPQPQPQPQPQPQPQPQPQPQPQPQPQPQPVPPPASLPQPLL 591
>AV419982
Length = 415
Score = 35.8 bits (81), Expect = 0.007
Identities = 27/78 (34%), Positives = 32/78 (40%), Gaps = 5/78 (6%)
Frame = +1
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTL----NIPQP-SHPPPPPPQDVVDNHA 61
PT + P TP F P P P T +IP S PPPPPP +
Sbjct: 166 PTPTSPPISTTPT*AQFPSPGHAPSWVAPSTFTSTTTPSIPLSLSPPPPPPPPPSASTSS 345
Query: 62 PVRSRRRGPPRNGPIPTP 79
P S + PR+ P PTP
Sbjct: 346 PSSSGKSTAPRSSP-PTP 396
>TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28040
{Arabidopsis thaliana;}, partial (8%)
Length = 937
Score = 35.8 bits (81), Expect = 0.007
Identities = 25/79 (31%), Positives = 37/79 (46%), Gaps = 2/79 (2%)
Frame = -1
Query: 2 NDSEQPTQSLPLPMLTPFQRMFAHPNIPPLQ--QQPLPLTLNIPQPSHPPPPPPQDVVDN 59
N +Q ++ P P +R PPLQ +QP P T N+ P+ P PPP ++
Sbjct: 787 NTQQQKKKNFPQKKKVPLRRRRRMK--PPLQPLRQPQPPTPNLLNPNPNPIPPPNPIL-- 620
Query: 60 HAPVRSRRRGPPRNGPIPT 78
+P + R PP P+ T
Sbjct: 619 -SPTPTPTRPPPHPLPLLT 566
Score = 32.0 bits (71), Expect = 0.094
Identities = 24/78 (30%), Positives = 35/78 (44%), Gaps = 4/78 (5%)
Frame = -1
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLT--LNIPQ--PSHPPPPPPQDVVDN 59
S PT + P P P + ++P+ P +P P + LN P+ P+ P PP+ +
Sbjct: 619 SPTPTPTRPPPHPLPLLTLKSNPSPPSPWTRPRPTSPRLNPPRLRPNRQPSAPPRAMPTP 440
Query: 60 HAPVRSRRRGPPRNGPIP 77
P RRR R P P
Sbjct: 439 ETPNGLRRRQLTRPPPPP 386
Score = 27.7 bits (60), Expect = 1.8
Identities = 27/88 (30%), Positives = 33/88 (36%), Gaps = 11/88 (12%)
Frame = -1
Query: 1 MNDSEQPTQSL------PLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPP---P 51
+ + PT +L P+P P P PP PLPL PS P P P
Sbjct: 697 LRQPQPPTPNLLNPNPNPIPPPNPILSPTPTPTRPPP--HPLPLLTLKSNPSPPSPWTRP 524
Query: 52 PPQDVVDNHAPVRSRRR--GPPRNGPIP 77
P N +R R+ PPR P P
Sbjct: 523 RPTSPRLNPPRLRPNRQPSAPPRAMPTP 440
>AW720113
Length = 512
Score = 35.8 bits (81), Expect = 0.007
Identities = 20/43 (46%), Positives = 22/43 (50%)
Frame = +1
Query: 17 TPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDN 59
TP F HP+ PL P PL PQP PPPPPP +N
Sbjct: 73 TPTTTAF-HPS--PLDPPPPPLPPPQPQPQPPPPPPPPQFHEN 192
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 35.0 bits (79), Expect = 0.011
Identities = 19/61 (31%), Positives = 24/61 (39%)
Frame = +1
Query: 17 TPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPI 76
+PF + P+ P LP N + P PPP + P RR PP N P
Sbjct: 166 SPFHSPPSSPSSNPNSHHLLPHNNNPTRSKSQPKPPPSPTSHSPPPPHPRRPNPPSNPPS 345
Query: 77 P 77
P
Sbjct: 346 P 348
Score = 28.1 bits (61), Expect = 1.4
Identities = 22/65 (33%), Positives = 27/65 (40%), Gaps = 4/65 (6%)
Frame = +2
Query: 12 PLPMLTPFQRMFAHPNIPPLQQ---QPLPLTLNIPQPSH-PPPPPPQDVVDNHAPVRSRR 67
P P L P + H ++ Q Q PL P+PS PPPPP+ P R R
Sbjct: 257 PNPSLHPHPPLTLHHHLTLAAQILLQIHPLRQPPPRPSLLRPPPPPRPFQRRRTPPRLHR 436
Query: 68 RGPPR 72
P R
Sbjct: 437 LVPQR 451
Score = 25.8 bits (55), Expect = 6.8
Identities = 22/73 (30%), Positives = 25/73 (34%), Gaps = 8/73 (10%)
Frame = +1
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQP-LPLT-------LNIPQPSHPPPPPPQDVVD 58
PT P P P R P+ PP P PLT +P P H PP
Sbjct: 280 PTSHSPPP---PHPRRPNPPSNPPSPPTPSAPLTP*APTAPSTLPTPPHSAAPPSSGSAT 450
Query: 59 NHAPVRSRRRGPP 71
A +R PP
Sbjct: 451 TSASATTRLSTPP 489
>AV429304
Length = 355
Score = 32.7 bits (73), Expect = 0.055
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 4/59 (6%)
Frame = +3
Query: 8 TQSLPLPMLTPFQRMFAHPNIPPLQQQP----LPLTLNIPQPSHPPPPPPQDVVDNHAP 62
T +P+P TP +++ PN+P Q P +P P P + PP P HAP
Sbjct: 60 TPYVPIPPKTP-SPVYSPPNVPSPPQTPHAPYVPTPPKTPSPVYSPPNVPSPPQTPHAP 233
Score = 28.9 bits (63), Expect = 0.80
Identities = 15/49 (30%), Positives = 21/49 (42%), Gaps = 4/49 (8%)
Frame = +3
Query: 18 PFQRMFAHPNIPPLQQQP----LPLTLNIPQPSHPPPPPPQDVVDNHAP 62
P ++ PN+P Q P +P+ P P + PP P HAP
Sbjct: 3 PTPPTYSPPNVPSPPQTPHTPYVPIPPKTPSPVYSPPNVPSPPQTPHAP 149
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,997,582
Number of Sequences: 28460
Number of extensions: 147448
Number of successful extensions: 4681
Number of sequences better than 10.0: 644
Number of HSP's better than 10.0 without gapping: 2607
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3488
length of query: 233
length of database: 4,897,600
effective HSP length: 87
effective length of query: 146
effective length of database: 2,421,580
effective search space: 353550680
effective search space used: 353550680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC141322.2


