

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141322.10 + phase: 0 /pseudo
(409 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
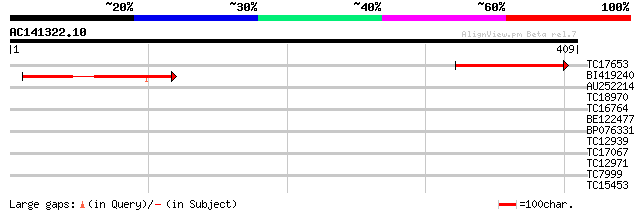
Score E
Sequences producing significant alignments: (bits) Value
TC17653 weakly similar to UP|Q7XAL4 (Q7XAL4) Mitochondrial inner... 134 2e-32
BI419240 124 2e-29
AU252214 30 0.96
TC18970 weakly similar to UP|Q8I8F0 (Q8I8F0) Chromosome scaffold... 29 1.2
TC16764 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp prot... 29 1.2
BE122477 29 1.6
BP076331 29 1.6
TC12939 28 2.1
TC17067 28 2.8
TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14... 27 4.7
TC7999 similar to UP|BAD01556 (BAD01556) ERF-like protein, parti... 27 4.7
TC15453 similar to UP|Q93ZE7 (Q93ZE7) AT3g52500/F22O6_120, parti... 27 8.1
>TC17653 weakly similar to UP|Q7XAL4 (Q7XAL4) Mitochondrial inner membrane
translocating protein-like protein, partial (16%)
Length = 587
Score = 134 bits (338), Expect = 2e-32
Identities = 59/82 (71%), Positives = 71/82 (85%)
Frame = +2
Query: 322 RCKAEHGAYKDRGIFYDNKILHISDADVREVKILESSPFIIVVFQTQQIHCVRDRNGEIT 381
RCKAEH AY+ GIF+DNKILH+SD DVRE K+LESSP I+V+F TQ IHCVRDR+GEIT
Sbjct: 2 RCKAEHNAYQSHGIFFDNKILHVSDVDVRETKMLESSPVIVVMFHTQPIHCVRDRHGEIT 181
Query: 382 EGGKDTIHSVYYLWALQMDSED 403
EGGKDTIH+++YLWAL ++D
Sbjct: 182 EGGKDTIHNIFYLWALPQVNQD 247
>BI419240
Length = 561
Score = 124 bits (312), Expect = 2e-29
Identities = 72/129 (55%), Positives = 81/129 (61%), Gaps = 18/129 (13%)
Frame = +2
Query: 10 SVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLY 69
SVDRR YSVF EFS+K+K E NPEF+KSVKELKE+ TKQTTEQLY
Sbjct: 215 SVDRRRYSVFKEFSEKIKGEVKSNPEFEKSVKELKER--------------TKQTTEQLY 352
Query: 70 RQFDSVWKEAEAAAKKVSHNVKEKISAA------------------TDFSTKQNADAKQG 111
+Q D VW EAEA AKKVSHNVKEK+SAA TD+S KQNADA QG
Sbjct: 353 KQVDEVWTEAEATAKKVSHNVKEKLSAANEEVKETFWIGKQDSSGSTDYSKKQNADANQG 532
Query: 112 SQKSPEEEK 120
S S E++
Sbjct: 533 STTSSGEKQ 559
>AU252214
Length = 350
Score = 29.6 bits (65), Expect = 0.96
Identities = 27/85 (31%), Positives = 43/85 (49%), Gaps = 6/85 (7%)
Frame = +1
Query: 43 LKEKAEELKGIKEGLK--EKTKQTTEQLYRQFDSVWKEAEA----AAKKVSHNVKEKISA 96
+ KAEE + K + +K K+T E+ D WKEAE AAKK ++K A
Sbjct: 88 VNSKAEEARARKASAESEKKNKETKEKE----DQFWKEAEGSRSRAAKKKEEEAEKKAEA 255
Query: 97 ATDFSTKQNADAKQGSQKSPEEEKN 121
A + A+A++ ++ +EEK+
Sbjct: 256 AA-----KRAEARRLAE---QEEKD 306
>TC18970 weakly similar to UP|Q8I8F0 (Q8I8F0) Chromosome scaffold protein
p85, partial (4%)
Length = 562
Score = 29.3 bits (64), Expect = 1.2
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +3
Query: 31 VKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQ 67
VK + VKE+K+K E +K KE +KEK ++ E+
Sbjct: 108 VKPKAGKPKVKEVKDKKEVVKDKKEVVKEKKEEAKEE 218
>TC16764 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (24%)
Length = 653
Score = 29.3 bits (64), Expect = 1.2
Identities = 16/42 (38%), Positives = 27/42 (64%)
Frame = +3
Query: 25 KVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTE 66
K KDE V+N +F+K+ EL+++ +LK L EK K+ ++
Sbjct: 321 KEKDEAVRNQDFEKA-GELRDREMDLKTQISALIEKGKEMSK 443
>BE122477
Length = 420
Score = 28.9 bits (63), Expect = 1.6
Identities = 23/92 (25%), Positives = 44/92 (47%)
Frame = +3
Query: 9 NSVDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQL 68
+S+D+ S +E SKKV + + + ++ K +++ + EGL E ++ + L
Sbjct: 123 SSLDQHASSPSSESSKKVAK--LIDSYIAEIASDVNLKPGKIRALAEGLPESSRLLHDGL 296
Query: 69 YRQFDSVWKEAEAAAKKVSHNVKEKISAATDF 100
YR D +K A +S KE++ D+
Sbjct: 297 YRALDIYFK----AHPWLSDKEKEELCNIIDY 380
>BP076331
Length = 358
Score = 28.9 bits (63), Expect = 1.6
Identities = 17/52 (32%), Positives = 27/52 (51%)
Frame = +2
Query: 330 YKDRGIFYDNKILHISDADVREVKILESSPFIIVVFQTQQIHCVRDRNGEIT 381
YK+ +DNK H S + + K S+ + ++FQ QQ R+R +IT
Sbjct: 71 YKNLEQVFDNKTPHASKSRR*DTKHFSSNRKLNIIFQRQQPKSCRNRATKIT 226
>TC12939
Length = 551
Score = 28.5 bits (62), Expect = 2.1
Identities = 28/113 (24%), Positives = 41/113 (35%)
Frame = -2
Query: 23 SKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAA 82
S+K D VK+ EFQ + KAE KG+ EK +A +
Sbjct: 352 SEKNSDNNVKDSEFQANKDLSNSKAEPDKGLPAAESEKPASD------------GDANSG 209
Query: 83 AKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESLFG 135
N E++ AA + DA G+ + E EE P+ + G
Sbjct: 208 TPNHELNNTEELPAAETETPVSGGDANAGT-SNHELNNTEELPAAESEAPASG 53
>TC17067
Length = 525
Score = 28.1 bits (61), Expect = 2.8
Identities = 16/54 (29%), Positives = 27/54 (49%), Gaps = 6/54 (11%)
Frame = +2
Query: 92 EKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNAS------ESLFGKFKS 139
EK+S TDF T +++ ++K ++N E +S S+FGK K+
Sbjct: 98 EKVSKVTDFLTGESSHRSAATEKQSRNKQNRELRVKGSSIRFPLKSSIFGKEKT 259
>TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (7%)
Length = 721
Score = 27.3 bits (59), Expect = 4.7
Identities = 30/106 (28%), Positives = 45/106 (42%), Gaps = 5/106 (4%)
Frame = +3
Query: 33 NPEFQKSVKELKEKAEELKGIKEGL-----KEKTKQTTEQLYRQFDSVWKEAEAAAKKVS 87
N E ++ + K K EELKGI L +EK +Q D K++E +
Sbjct: 285 NREEEEGDNKSKNKKEELKGILTSLLLLDEQEKVEQ---------DHQIKDSEDEKFSLE 437
Query: 88 HNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASESL 133
N K+K A + T N D G + E E+ + S N + S+
Sbjct: 438 TNHKKKTKAMIQYYT--NLDDYYGQVE--ESERVQRKKSRNMANSV 563
>TC7999 similar to UP|BAD01556 (BAD01556) ERF-like protein, partial (11%)
Length = 690
Score = 27.3 bits (59), Expect = 4.7
Identities = 26/88 (29%), Positives = 38/88 (42%)
Frame = +2
Query: 56 GLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKS 115
GLKEK K+ + Q V +EAEAAAK + +++ A ++ G Q
Sbjct: 167 GLKEKEKEKETE---QKTEVVEEAEAAAKNEVQELSDELMAYENYMKFYQIPYYDG-QSV 334
Query: 116 PEEEKNEESPSGNASESLFGKFKSTFSS 143
E +ES G+ +F FSS
Sbjct: 335 AENNSVQESEVGDLWSFDE*EFSRIFSS 418
>TC15453 similar to UP|Q93ZE7 (Q93ZE7) AT3g52500/F22O6_120, partial (8%)
Length = 639
Score = 26.6 bits (57), Expect = 8.1
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -1
Query: 105 NADAKQGSQKSPEEEKNEESPSGNASESLFGKFKSTFSS 143
N ++ G+ PE+ K +P G ++ KF+S FS+
Sbjct: 159 NINSGIGTVSVPEKSKQRPNPDGASTSFALEKFESCFSN 43
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,390,907
Number of Sequences: 28460
Number of extensions: 57164
Number of successful extensions: 323
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 318
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 321
length of query: 409
length of database: 4,897,600
effective HSP length: 92
effective length of query: 317
effective length of database: 2,279,280
effective search space: 722531760
effective search space used: 722531760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC141322.10


