

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141115.10 + phase: 0 /pseudo
(173 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
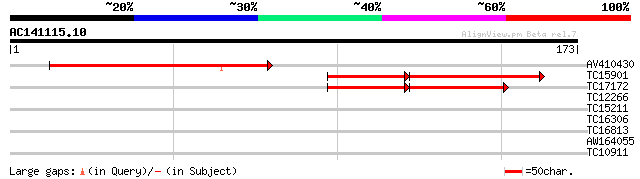
AV410430 87 1e-18
TC15901 homologue to UP|AAR20777 (AAR20777) At2g22125, partial (... 47 2e-08
TC17172 homologue to UP|AAR20777 (AAR20777) At2g22125, partial (... 42 2e-07
TC12266 weakly similar to GB|AAB68410.1|500677|YSCH9315 Yhr122wp... 27 2.5
TC15211 similar to UP|Q9ZRD9 (Q9ZRD9) GMFP4 (Fragment), partial ... 25 7.3
TC16306 25 7.3
TC16813 homologue to UP|BPHE_COMTE (Q46373) Biphenyl dioxygenase... 25 9.5
AW164055 25 9.5
TC10911 25 9.5
>AV410430
Length = 374
Score = 87.4 bits (215), Expect = 1e-18
Identities = 49/75 (65%), Positives = 54/75 (71%), Gaps = 7/75 (9%)
Frame = +3
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D L LL Q WSACP EVSR QS AAA AIPLLQ LIQ GP F EKAEF+ LV
Sbjct: 150 LDTLSLLRQAWSACPAEVSRAQSIAAADAIPLLQYLIQSGPPRFQEKAEFLLQCLPGTLV 329
Query: 66 MIVKRGNNMRQCVGN 80
+I+KRGNNM+Q VGN
Sbjct: 330 VIIKRGNNMKQSVGN 374
>TC15901 homologue to UP|AAR20777 (AAR20777) At2g22125, partial (24%)
Length = 606
Score = 47.4 bits (111), Expect(2) = 2e-08
Identities = 22/41 (53%), Positives = 28/41 (67%)
Frame = +3
Query: 123 ADEHTLLPTSKSGQPRNLEVELKWSNKPSYAYSIKLSIPSF 163
A E+TLLP SKSG PRNLE+E +WSNK + + + I F
Sbjct: 144 AGEYTLLPASKSGPPRNLEIEFQWSNKTNDTV*LGMEILDF 266
Score = 25.4 bits (54), Expect(2) = 2e-08
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = +2
Query: 98 YMVL*ECSSRTEASYLLQKQA*SGK 122
+MV+*E S R EA+ LQK SGK
Sbjct: 11 FMVI*ESSKRPEATDFLQK*EQSGK 85
>TC17172 homologue to UP|AAR20777 (AAR20777) At2g22125, partial (28%)
Length = 553
Score = 42.0 bits (97), Expect(2) = 2e-07
Identities = 19/30 (63%), Positives = 23/30 (76%)
Frame = +3
Query: 123 ADEHTLLPTSKSGQPRNLEVELKWSNKPSY 152
A E+TL P SKSG RNLE+E +WSNK S+
Sbjct: 183 AGEYTLSPESKSGPSRNLEIEFQWSNK*SW 272
Score = 28.1 bits (61), Expect(2) = 2e-07
Identities = 15/25 (60%), Positives = 18/25 (72%)
Frame = +2
Query: 98 YMVL*ECSSRTEASYLLQKQA*SGK 122
+MVL*E S R EAS+ LQK +GK
Sbjct: 50 FMVL*ESSKRPEASHFLQK*EQNGK 124
>TC12266 weakly similar to GB|AAB68410.1|500677|YSCH9315 Yhr122wp
{Saccharomyces cerevisiae;} , partial (27%)
Length = 715
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/48 (29%), Positives = 23/48 (47%)
Frame = -2
Query: 113 LLQKQA*SGKADEHTLLPTSKSGQPRNLEVELKWSNKPSYAYSIKLSI 160
++Q Q S K + PTS+ G + V L W K S+ + ++ I
Sbjct: 609 IIQDQCSSNKKHDK*S*PTSRKGTKKTPTVLLHWKEKSSFNHKMEQGI 466
>TC15211 similar to UP|Q9ZRD9 (Q9ZRD9) GMFP4 (Fragment), partial (94%)
Length = 531
Score = 25.0 bits (53), Expect = 7.3
Identities = 11/22 (50%), Positives = 14/22 (63%), Gaps = 1/22 (4%)
Frame = -3
Query: 4 QWHVKKLSWMDALFL-LIQGWS 24
QW L W+D LF+ +I GWS
Sbjct: 118 QWLPSLLPWIDELFITIIIGWS 53
>TC16306
Length = 598
Score = 25.0 bits (53), Expect = 7.3
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -3
Query: 148 NKPSYAYSIKLSIPSFKFDYLFILVL 173
N P +KL +PS KFD+L +L L
Sbjct: 200 NDPMADKLLKLLLPSLKFDFLDLLGL 123
>TC16813 homologue to UP|BPHE_COMTE (Q46373) Biphenyl dioxygenase beta
subunit (Biphenyl 2,3-dioxygenase) , partial (8%)
Length = 621
Score = 24.6 bits (52), Expect = 9.5
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +2
Query: 132 SKSGQPRNLEVELKWSNKPS 151
+ G R LE+E+ WSN+ S
Sbjct: 62 NNDGSSRTLEIEIVWSNRIS 121
>AW164055
Length = 361
Score = 24.6 bits (52), Expect = 9.5
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -2
Query: 62 FILVMIVKRGNNMRQCVGNQGKIT 85
F+L+ I+ RGN + NQG IT
Sbjct: 282 FLLLSIMYRGNKISSIKENQGTIT 211
>TC10911
Length = 498
Score = 24.6 bits (52), Expect = 9.5
Identities = 9/28 (32%), Positives = 18/28 (64%)
Frame = +1
Query: 144 LKWSNKPSYAYSIKLSIPSFKFDYLFIL 171
+ WS+ SYA+ I + P+ + Y++I+
Sbjct: 1 ITWSSPKSYAWHILEAYPTHRMPYVWIV 84
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,076,736
Number of Sequences: 28460
Number of extensions: 36868
Number of successful extensions: 229
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 228
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 228
length of query: 173
length of database: 4,897,600
effective HSP length: 84
effective length of query: 89
effective length of database: 2,506,960
effective search space: 223119440
effective search space used: 223119440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC141115.10


