

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
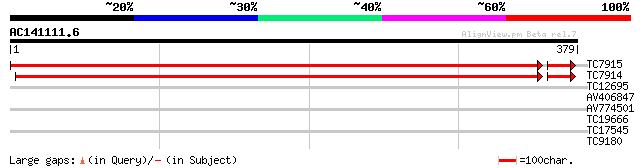
Query= AC141111.6 - phase: 0 /pseudo
(379 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7915 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete 668 0.0
TC7914 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete 647 0.0
TC12695 similar to UP|Q9SAK1 (Q9SAK1) T8K14.10, partial (17%) 28 1.9
AV406847 28 1.9
AV774501 27 4.3
TC19666 27 4.3
TC17545 27 5.6
TC9180 26 9.6
>TC7915 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete
Length = 1622
Score = 668 bits (1724), Expect(2) = 0.0
Identities = 321/356 (90%), Positives = 343/356 (96%)
Frame = +2
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
MDY N PGR+HLFVPGPVNIP+QVIRAM+RNNEDYR+PAIPALTKTLLEDVKKIFKTTTG
Sbjct: 143 MDYFNAPGRHHLFVPGPVNIPEQVIRAMNRNNEDYRAPAIPALTKTLLEDVKKIFKTTTG 322
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRG 120
TPFLIPTTGTGAWESALTNTLSPGDR VSF+IGQFSLLW+DQQQRL FNVDVVESEWGRG
Sbjct: 323 TPFLIPTTGTGAWESALTNTLSPGDRTVSFLIGQFSLLWIDQQQRLNFNVDVVESEWGRG 502
Query: 121 ADLDILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSI 180
ADLD+LESKLASDSAHTIKAICIVHNETATGVTN+L+KVR+LLDAY HPALL+VDGVSSI
Sbjct: 503 ADLDVLESKLASDSAHTIKAICIVHNETATGVTNDLSKVRKLLDAYNHPALLIVDGVSSI 682
Query: 181 CALDFRMDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKF 240
CALDFRMDEWGVDVAITGSQKALSLPTG+G V ASPKA+EASK AKSLRVFFDW DYLKF
Sbjct: 683 CALDFRMDEWGVDVAITGSQKALSLPTGMGIVCASPKALEASKYAKSLRVFFDWKDYLKF 862
Query: 241 YKMGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQE 300
Y+MGTYWPYTPSIQLLYGLR ALDL+FEEGLEN+ ARH RLG ATR+AVEAWGLKNCTQ+
Sbjct: 863 YEMGTYWPYTPSIQLLYGLRTALDLLFEEGLENVFARHKRLGKATRIAVEAWGLKNCTQK 1042
Query: 301 EEWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
EEWFSDTVTAVVVPPYIDGAE+V+R+WKRYN+SLGLGLNKVAGKVFRIGHLGNLNE
Sbjct: 1043EEWFSDTVTAVVVPPYIDGAEVVKRSWKRYNMSLGLGLNKVAGKVFRIGHLGNLNE 1210
Score = 30.0 bits (66), Expect(2) = 0.0
Identities = 15/19 (78%), Positives = 16/19 (83%)
Frame = +3
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLLPVHT RT+ L SL
Sbjct: 1281 EVELLLPVHTCRTLSL*SL 1337
>TC7914 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete
Length = 1532
Score = 647 bits (1669), Expect(2) = 0.0
Identities = 309/352 (87%), Positives = 336/352 (94%)
Frame = +2
Query: 5 NGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFL 64
N PGR+HLFVPGP NIP+QVIRAM+RNNEDYR+PAIPALTKTLLEDVKKIFKT+TG PFL
Sbjct: 149 NAPGRHHLFVPGPTNIPEQVIRAMNRNNEDYRAPAIPALTKTLLEDVKKIFKTSTGIPFL 328
Query: 65 IPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLD 124
PTTGTGAWESALTNTLSPGDRIVSF+IGQFSLLW+DQQQRL FNVDVVES+WG+GA+LD
Sbjct: 329 FPTTGTGAWESALTNTLSPGDRIVSFLIGQFSLLWIDQQQRLNFNVDVVESDWGQGANLD 508
Query: 125 ILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALD 184
+LESKLASD HTIKAICIVHNETATGVTNNLAKVR++LD YQHPALL+VDGVSSICALD
Sbjct: 509 VLESKLASDRTHTIKAICIVHNETATGVTNNLAKVRKILDDYQHPALLIVDGVSSICALD 688
Query: 185 FRMDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMG 244
FRMDEWGVDVA+TGSQKALSLPTG+G V A PKAIEASKSAKSLRVFFDW+DYLKFYK+G
Sbjct: 689 FRMDEWGVDVAVTGSQKALSLPTGLGIVCAGPKAIEASKSAKSLRVFFDWNDYLKFYKLG 868
Query: 245 TYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWF 304
TYWPYTPSIQLLYGLR ALDLIFEEGL+N+I RH RLG ATRLAVEAWGLKNCTQ+EEW+
Sbjct: 869 TYWPYTPSIQLLYGLREALDLIFEEGLDNVILRHKRLGHATRLAVEAWGLKNCTQKEEWY 1048
Query: 305 SDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
SDTVTAV+VPPYID AE+VRR+WKRYN+SLGLGLN+VAGKVFRIGHLGNLNE
Sbjct: 1049SDTVTAVLVPPYIDAAEVVRRSWKRYNMSLGLGLNQVAGKVFRIGHLGNLNE 1204
Score = 29.6 bits (65), Expect(2) = 0.0
Identities = 15/19 (78%), Positives = 16/19 (83%)
Frame = +3
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLL VHT+R IFL SL
Sbjct: 1275 EVELLLLVHTFRIIFL*SL 1331
>TC12695 similar to UP|Q9SAK1 (Q9SAK1) T8K14.10, partial (17%)
Length = 571
Score = 28.5 bits (62), Expect = 1.9
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Frame = -2
Query: 302 EWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSL-GLGLNKVAGKVFRIG 349
+W TVT++ VPP GA + K S+ +G + V K + IG
Sbjct: 195 QWLGSTVTSIAVPPKSQGASTILGVLKIAIKSIRSMGQHSVGLKAWAIG 49
>AV406847
Length = 415
Score = 28.5 bits (62), Expect = 1.9
Identities = 18/46 (39%), Positives = 21/46 (45%)
Frame = +3
Query: 38 PAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTGAWESALTNTLSP 83
P P+ T +L V F TT PF P T T L +TLSP
Sbjct: 213 PRAPSHTPSLRTSVPPSFA*TTVAPFPSPPTATPRASLKLYSTLSP 350
>AV774501
Length = 514
Score = 27.3 bits (59), Expect = 4.3
Identities = 8/27 (29%), Positives = 17/27 (62%)
Frame = -3
Query: 279 NRLGTATRLAVEAWGLKNCTQEEEWFS 305
+R + +V+ W L+ C++++ WFS
Sbjct: 311 SRTSLSLHFSVDPWELRLCSEDDSWFS 231
>TC19666
Length = 521
Score = 27.3 bits (59), Expect = 4.3
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +1
Query: 314 PPYIDGAEIVRRAWKRYNLSLGLGLNKVAGK 344
PP G++ R WKR+ L L LG +++ K
Sbjct: 94 PPLHLGSQHTRTIWKRFTLELLLGASEILQK 186
>TC17545
Length = 465
Score = 26.9 bits (58), Expect = 5.6
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = -1
Query: 247 WPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRL 281
WP+TPS Q+L+ R + E LE +H ++
Sbjct: 258 WPHTPSKQVLH-FRTITSIKMESKLEGSFGQHEKI 157
>TC9180
Length = 724
Score = 26.2 bits (56), Expect = 9.6
Identities = 16/50 (32%), Positives = 25/50 (50%)
Frame = -1
Query: 25 IRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTGAWE 74
+R + R+ + P +T T++ K F T T FL TTG G+W+
Sbjct: 679 LRKVERDTNQAQGPEKTRITITII--CKN*F--TKATSFLF*TTGVGSWD 542
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,446,641
Number of Sequences: 28460
Number of extensions: 81527
Number of successful extensions: 319
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 319
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 319
length of query: 379
length of database: 4,897,600
effective HSP length: 92
effective length of query: 287
effective length of database: 2,279,280
effective search space: 654153360
effective search space used: 654153360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC141111.6


