

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
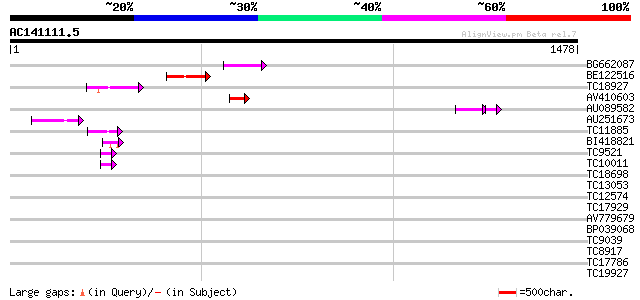
Query= AC141111.5 - phase: 0 /pseudo
(1478 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662087 92 6e-19
BE122516 78 9e-15
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 74 2e-13
AV410603 72 8e-13
AU089582 50 6e-08
AU251673 55 1e-07
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 53 3e-07
BI418821 46 5e-05
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 44 2e-04
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 42 5e-04
TC18698 39 0.006
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 37 0.023
TC12574 36 0.051
TC17929 34 0.19
AV779679 34 0.19
BP039068 31 1.2
TC9039 30 2.1
TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, parti... 29 6.2
TC17786 similar to PIR|T46177|T46177 villin 3 homolog T8H10.10 -... 28 8.1
TC19927 weakly similar to UP|O80689 (O80689) F8K4.2 protein, par... 28 8.1
>BG662087
Length = 373
Score = 92.0 bits (227), Expect = 6e-19
Identities = 45/113 (39%), Positives = 68/113 (59%)
Frame = +1
Query: 557 GSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQK 616
G R+ +DY LNK K+ YPLP ID L+D ++ S +D SGYHQIK+ D K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 617 TAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
TAF T +Y Y+ +PFG+ NA + M+R+F + R + V++D++++ S
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKS 354
>BE122516
Length = 364
Score = 78.2 bits (191), Expect = 9e-15
Identities = 46/116 (39%), Positives = 71/116 (60%)
Frame = +2
Query: 408 LGMNWLEYNHVHINYFTKSVYFSSVEEESGAEFLSTKQLKQMERDGILMYPLMASLSFEN 467
+GMNWL N +N K+V F + E ++ + K K E + ++ L A + ++
Sbjct: 14 VGMNWLTANDATLNCRKKTVTFGTSEGDAKRVKRTDKVGKASECESDVL--LGALETDKS 187
Query: 468 QAVIDKLQVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRMSASELA 523
++ + VV +F +VFP+E+ ++P EREVEFSID + GT P+S+APYRMS ELA
Sbjct: 188 DTGVEGIPVVREFSDVFPEEVSELPPEREVEFSID*VPGTGPISIAPYRMSLVELA 355
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 73.6 bits (179), Expect = 2e-13
Identities = 45/156 (28%), Positives = 68/156 (42%), Gaps = 7/156 (4%)
Frame = -2
Query: 201 RKTKGQQSRPKPYSAPADKGKQRMVDDR------RPKKKDAPAEITCFNCGEKGHKSNVC 254
R + + + KP+ P ++G RP + D +EI C C +KGH +N C
Sbjct: 464 RFDRNKSFQKKPFQRPQNRGTSSGYSHSFGNFVPRPTQSDT-SEIVCHRCSKKGHFANRC 288
Query: 255 PEEIKKCVRCGKKGHIVADCKRNDI-VCFNFNEEGHIGSQCKQPKKSPTTGRVFALDGTQ 313
P+ + C C K GH DC + N + K+ + RV+ + G +
Sbjct: 287 PDLV--CWNCQKTGHSGKDCTNPKVEAATNAIAARRPAPAANKGKRPVASARVYTVSGAE 114
Query: 314 TENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDC 349
+ D LIR +N PL + D+GATH FI C
Sbjct: 113 SHRADGLIRSVGSVNCKPLTILFDSGATHSFIDLAC 6
>AV410603
Length = 162
Score = 71.6 bits (174), Expect = 8e-13
Identities = 30/53 (56%), Positives = 42/53 (78%)
Frame = +1
Query: 572 TIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYG 624
T+K+ +P+P +D+L+D+L G++ FSK+DLRSGYHQI VK ED KT F T +G
Sbjct: 4 TVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>AU089582
Length = 383
Score = 50.4 bits (119), Expect(2) = 6e-08
Identities = 30/84 (35%), Positives = 42/84 (49%)
Frame = +1
Query: 1161 SCCPVGRDLYQGDREVTWCSFEHCIR*RSKIYF*VLEKLARGFGFKVEIEFGLSSADRWS 1220
SC + DL D + WC+ + R RS I+ LE + FG ++ E+ SS++ WS
Sbjct: 7 SCVSIC*DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WS 186
Query: 1221 VGEDNSVARGFVESLCS*ARRNLG 1244
V ED S RG+ LC R G
Sbjct: 187 VREDYSDLRGYASCLCVXPERXAG 258
Score = 24.6 bits (52), Expect(2) = 6e-08
Identities = 19/43 (44%), Positives = 23/43 (53%)
Frame = +2
Query: 1240 RRNLG*SSSVDRVHIQ**LSF*YWNDTF*GFVWSEMQNSVVLV 1282
R LG D + I **LS *+ + T * VW EMQ + LV
Sbjct: 245 RGXLGSVFVFDGIRI***LSV*HLDGTI*SLVW*EMQVTYRLV 373
>AU251673
Length = 413
Score = 54.7 bits (130), Expect = 1e-07
Identities = 37/138 (26%), Positives = 65/138 (46%), Gaps = 1/138 (0%)
Frame = +2
Query: 56 CSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRYFPEDVRRKKEI 115
CS+ + V + QL A DW+ L TWA F EFM R+ P+ VR
Sbjct: 5 CSDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVR 184
Query: 116 EFLELKQG-NMYVTEYAAKFVELAEFYPHYAAETVEFSKCIKFENGLRPDIKRAIEYQQI 174
+F L+Q M V+EY+A F L+ + P+ +E + +F GL+ + +++ +
Sbjct: 185 DFERLEQAEGMTVSEYSAHFTHLSRYVPY---PLLEEERVKRFVRGLKEYLFKSVVGSKS 355
Query: 175 RVFPDLVNSCRIYEEDTK 192
++++ + E+ K
Sbjct: 356 STLSEVLSLALLVEQRQK 409
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 53.1 bits (126), Expect = 3e-07
Identities = 26/93 (27%), Positives = 42/93 (44%)
Frame = +2
Query: 202 KTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCPEEIKKC 261
+++ + P +D+ R RR ++ + C NC GH + CP + C
Sbjct: 227 RSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRPGHFARECP-NVAIC 403
Query: 262 VRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQC 294
CG GHI ++C + C+N E GH+ S C
Sbjct: 404 HNCGLPGHIASECTTKSL-CWNCKEPGHMASSC 499
>BI418821
Length = 614
Score = 45.8 bits (107), Expect = 5e-05
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 15/71 (21%)
Frame = +2
Query: 241 CFNCGEKGHKSNVCPEEIKK--------CVRCGKKGHIVADCKRND-------IVCFNFN 285
C+NCG+ GH + C C CG GH+ DC R++ C+N
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGGGAGCYNCG 571
Query: 286 EEGHIGSQCKQ 296
+ GH+ C +
Sbjct: 572 DTGHLARDCNR 604
Score = 38.5 bits (88), Expect = 0.008
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 7/45 (15%)
Frame = +2
Query: 241 CFNCGEKGHKSNVCPEEIKK-------CVRCGKKGHIVADCKRND 278
C+NCG+ GH + C C CG GH+ DC R++
Sbjct: 476 CYNCGDAGHLARDCNRSNNNSGGGGAGCYNCGDTGHLARDCNRSN 610
Score = 35.4 bits (80), Expect = 0.066
Identities = 14/52 (26%), Positives = 22/52 (41%), Gaps = 8/52 (15%)
Frame = +2
Query: 261 CVRCGKKGHIVADCKRND--------IVCFNFNEEGHIGSQCKQPKKSPTTG 304
C CG GH+ DC R++ C+N + GH+ C + + G
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGG 547
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 43.5 bits (101), Expect = 2e-04
Identities = 19/44 (43%), Positives = 25/44 (56%), Gaps = 2/44 (4%)
Frame = +1
Query: 236 PAEITCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRN 277
P CFNCG GH + C + KC RCG++GHI +CK +
Sbjct: 376 PGSGRCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNS 507
Score = 38.5 bits (88), Expect = 0.008
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Frame = +1
Query: 260 KCVRCGKKGHIVADCKRND--IVCFNFNEEGHIGSQCK-QPKK 299
+C CG GH DCK D C+ E GHI CK PKK
Sbjct: 388 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKK 516
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 42.4 bits (98), Expect = 5e-04
Identities = 18/44 (40%), Positives = 24/44 (53%), Gaps = 2/44 (4%)
Frame = +2
Query: 236 PAEITCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRN 277
P CFNCG GH + C + KC RCG +GH+ +CK +
Sbjct: 356 PGSGRCFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCKNS 487
Score = 37.4 bits (85), Expect = 0.017
Identities = 17/44 (38%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Frame = +2
Query: 260 KCVRCGKKGHIVADCKRND--IVCFNFNEEGHIGSQCK-QPKKS 300
+C CG GH DCK D C+ + GH+ CK PKK+
Sbjct: 368 RCFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCKNSPKKN 499
>TC18698
Length = 808
Score = 38.9 bits (89), Expect = 0.006
Identities = 17/55 (30%), Positives = 32/55 (57%)
Frame = -2
Query: 615 QKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
+KT +Y Y+VMP G+ N + M++IFH + + V V+++D+++ S
Sbjct: 804 KKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKS 640
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 37.0 bits (84), Expect = 0.023
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +3
Query: 239 ITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADC 274
+ C NC + GH S C + C CG +GH+ +C
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
Score = 31.2 bits (69), Expect = 1.2
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = +3
Query: 261 CVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQC 294
C C + GH+ DC ++C N GH+ +C
Sbjct: 9 CRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
>TC12574
Length = 325
Score = 35.8 bits (81), Expect = 0.051
Identities = 14/30 (46%), Positives = 22/30 (72%)
Frame = +2
Query: 642 FMEYMNRIFHAFLDRFVVVFIDDILIYSKD 671
F +N IF +F + F++VFI+DIL Y++D
Sbjct: 2 FKNSVNHIFESFFEHFMIVFINDILSYTED 91
>TC17929
Length = 791
Score = 33.9 bits (76), Expect = 0.19
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +2
Query: 231 KKKDAPAEITCFNCGEKGHKSNVCPEE 257
+ D+ TC+ CGE GHK CP+E
Sbjct: 32 RPNDSKFRQTCYRCGESGHKMRNCPKE 112
>AV779679
Length = 440
Score = 33.9 bits (76), Expect = 0.19
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +3
Query: 558 SMRLCIDYRQLNKVTIKNRYPLPRIDDLMD 587
+M+LC DY QL+ VTI N+ LP +D+ D
Sbjct: 345 TMQLCDDYMQLDYVTIPNKSLLPHLDEWSD 434
>BP039068
Length = 467
Score = 31.2 bits (69), Expect = 1.2
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = +3
Query: 236 PAEITCFNCGEKGHKSNVCP 255
P + C NC E GH SN CP
Sbjct: 297 PRQTVCMNCQETGHASNDCP 356
>TC9039
Length = 1218
Score = 30.4 bits (67), Expect = 2.1
Identities = 18/65 (27%), Positives = 29/65 (43%), Gaps = 7/65 (10%)
Frame = +2
Query: 197 VVNERKTKGQQSRPKPY-SAPADKGKQRMVDDRRPKKKDAPA------EITCFNCGEKGH 249
V E + K ++ + S DKGK++ + + + DAPA + TC+ C GH
Sbjct: 692 VQEEERLKQERKESAHFVSTSKDKGKRKKTVEPKNEAADAPAPKKQKEDDTCYFCNVSGH 871
Query: 250 KSNVC 254
C
Sbjct: 872 MKKKC 886
>TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, partial (3%)
Length = 676
Score = 28.9 bits (63), Expect = 6.2
Identities = 16/58 (27%), Positives = 29/58 (49%), Gaps = 4/58 (6%)
Frame = +2
Query: 188 EEDTKAHYK----VVNERKTKGQQSRPKPYSAPADKGKQRMVDDRRPKKKDAPAEITC 241
EE+ +A K E K + ++ PKP P ++ + V + +PK ++A + TC
Sbjct: 323 EEEVQAEPKEEKATTEEVKVETKEVNPKPEEEPKEEEPKAQVQEEKPKTEEA*S*ETC 496
>TC17786 similar to PIR|T46177|T46177 villin 3 homolog T8H10.10 - Arabidopsis
thaliana (fragment) {Arabidopsis thaliana;}, partial
(42%)
Length = 1228
Score = 28.5 bits (62), Expect = 8.1
Identities = 8/18 (44%), Positives = 11/18 (60%)
Frame = -1
Query: 1331 RRRPRVFESHSCDWCWMC 1348
R R E++ C+WCW C
Sbjct: 106 REREEKSENYCCEWCWCC 53
>TC19927 weakly similar to UP|O80689 (O80689) F8K4.2 protein, partial (14%)
Length = 529
Score = 28.5 bits (62), Expect = 8.1
Identities = 13/44 (29%), Positives = 26/44 (58%)
Frame = -1
Query: 1329 VSRRRPRVFESHSCDWCWMCFEVKEVDSEIFRSVSDIEKIWNGG 1372
V ++RP S+ C + + + +V +++F S+ IE I++GG
Sbjct: 139 VIKQRPHKVPSNICSFSYGTGQRIQVTTQVFHSIGIIEYIFSGG 8
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.349 0.155 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,666,281
Number of Sequences: 28460
Number of extensions: 338562
Number of successful extensions: 3030
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 2921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3006
length of query: 1478
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1376
effective length of database: 1,994,680
effective search space: 2744679680
effective search space used: 2744679680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141111.5


