

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
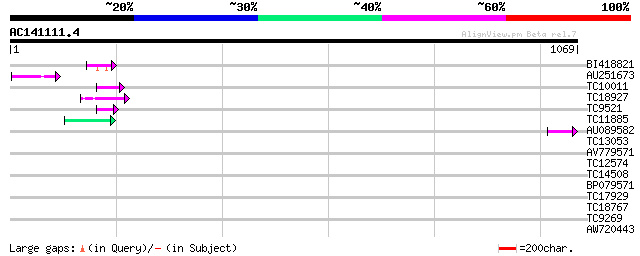
Query= AC141111.4 - phase: 0 /pseudo
(1069 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI418821 50 2e-06
AU251673 46 3e-05
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 44 1e-04
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 44 1e-04
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 44 2e-04
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 44 2e-04
AU089582 42 5e-04
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 35 0.082
AV779571 31 0.90
TC12574 30 2.6
TC14508 similar to AAP86659 (AAP86659) 26S proteasome subunit RP... 30 2.6
BP079571 24 3.1
TC17929 29 3.4
TC18767 29 4.5
TC9269 similar to UP|AAR92258 (AAR92258) At1g03260, partial (8%) 28 10.0
AW720443 28 10.0
>BI418821
Length = 614
Score = 49.7 bits (117), Expect = 2e-06
Identities = 22/71 (30%), Positives = 30/71 (41%), Gaps = 15/71 (21%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDEKK--------CFRCGKKGHTLADCKR-------GDIVCYNFN 189
C+ CG+ GH + C R C+ CG GH DC R G CYN
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGGGAGCYNCG 571
Query: 190 EEGHISLQCTQ 200
+ GH++ C +
Sbjct: 572 DTGHLARDCNR 604
Score = 40.0 bits (92), Expect = 0.002
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 7/45 (15%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDEKK-------CFRCGKKGHTLADCKRGD 182
C+ CG+ GH + C+R C+ CG GH DC R +
Sbjct: 476 CYNCGDAGHLARDCNRSNNNSGGGGAGCYNCGDTGHLARDCNRSN 610
Score = 39.7 bits (91), Expect = 0.003
Identities = 17/52 (32%), Positives = 22/52 (41%), Gaps = 8/52 (15%)
Frame = +2
Query: 165 CFRCGKKGHTLADCKR--------GDIVCYNFNEEGHISLQCTQPKKVRTGG 208
C+ CG GH DC R G CYN + GH++ C + GG
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGHLARDCNRSNNNSGGG 547
>AU251673
Length = 413
Score = 46.2 bits (108), Expect = 3e-05
Identities = 27/95 (28%), Positives = 52/95 (54%), Gaps = 1/95 (1%)
Frame = +2
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGD-MSVTEYAAKFVELAKFYPHYTAETAEFSKCIKI 61
EF+NR+ P+ VR +F L+Q + M+V+EY+A F L+++ P+ E + ++
Sbjct: 134 EFMNRFLPQSVRDGFVRDFERLEQAEGMTVSEYSAHFTHLSRYVPYPLLEEERVKRFVR- 310
Query: 62 ENGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTK 96
GL+ + +++ K SE++S + E+ K
Sbjct: 311 --GLKEYLFKSVVGSKSSTLSEVLSLALLVEQRQK 409
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 44.3 bits (103), Expect = 1e-04
Identities = 23/57 (40%), Positives = 30/57 (52%), Gaps = 5/57 (8%)
Frame = +2
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKK--VRTGGKVFALTG 215
+CF CG GH DCK GD CY + GH+ C PKK +T ++F+ TG
Sbjct: 368 RCFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCKNSPKKNEWQTWKELFSFTG 538
Score = 40.4 bits (93), Expect = 0.001
Identities = 16/37 (43%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG +GH +CK
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGHVERNCK 481
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 43.9 bits (102), Expect = 1e-04
Identities = 27/95 (28%), Positives = 45/95 (46%), Gaps = 1/95 (1%)
Frame = -2
Query: 133 RPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDI-VCYNFNEE 191
RP D ++IVC +C +KGH +N C + C+ C K GH+ DC + N
Sbjct: 362 RPTQSDT-SEIVCHRCSKKGHFANRC--PDLVCWNCQKTGHSGKDCTNPKVEAATNAIAA 192
Query: 192 GHISLQCTQPKKVRTGGKVFALTGTQTVNED*LIR 226
+ + K+ +V+ ++G ++ D LIR
Sbjct: 191 RRPAPAANKGKRPVASARVYTVSGAESHRADGLIR 87
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/44 (47%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Frame = +1
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKKV 204
+CF CG GH DCK GD CY E GHI C PKK+
Sbjct: 388 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKL 519
Score = 40.8 bits (94), Expect = 0.001
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Frame = +1
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG++GH +CK
Sbjct: 391 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 501
Score = 28.9 bits (63), Expect = 4.5
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = +1
Query: 145 CFKCGEKGHKSNVCDRDEKKCFR 167
C++CGE+GH C KK R
Sbjct: 457 CYRCGERGHIEKNCKNSPKKLSR 525
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 43.5 bits (101), Expect = 2e-04
Identities = 26/96 (27%), Positives = 38/96 (39%)
Frame = +2
Query: 103 SERRGKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDE 162
S R + + P D+ R RR R D +C C GH + C +
Sbjct: 218 SRSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRPGHFARECP-NV 394
Query: 163 KKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
C CG GH ++C + C+N E GH++ C
Sbjct: 395 AICHNCGLPGHIASECTTKSL-CWNCKEPGHMASSC 499
Score = 35.0 bits (79), Expect = 0.063
Identities = 22/76 (28%), Positives = 33/76 (42%), Gaps = 5/76 (6%)
Frame = +2
Query: 113 RPKPYSAPPDKGK*R---LKDERRPK--MRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFR 167
R PY +G R K+ +RP R+ P +C CG GH ++ C + C+
Sbjct: 290 RDAPYRRDSRRGFSRDNLCKNCKRPGHFARECPNVAICHNCGLPGHIASEC-TTKSLCWN 466
Query: 168 CGKKGHTLADCKRGDI 183
C + GH + C I
Sbjct: 467 CKEPGHMASSCPNDGI 514
Score = 29.6 bits (65), Expect = 2.6
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = +2
Query: 165 CFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCT 199
C C + GH +C I C+N GHI+ +CT
Sbjct: 344 CKNCKRPGHFARECPNVAI-CHNCGLPGHIASECT 445
>AU089582
Length = 383
Score = 42.0 bits (97), Expect = 5e-04
Identities = 22/56 (39%), Positives = 30/56 (53%)
Frame = +1
Query: 1014 DLYS*YCEVAWSSVEYCVG*RSEVHF*ILEEFARCIGFEVEVEFGLSSVDRWSVGE 1069
DL * C +AW + Y RS +H LE F+ C G ++ E+ SS + WSV E
Sbjct: 28 DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSVRE 195
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 34.7 bits (78), Expect = 0.082
Identities = 14/39 (35%), Positives = 18/39 (45%)
Frame = +3
Query: 143 IVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRG 181
+VC C + GH S C C CG +GH +C G
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYECPSG 119
Score = 32.3 bits (72), Expect = 0.41
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = +3
Query: 165 CFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
C C + GH DC ++C+N GH++ +C
Sbjct: 9 CRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 110
>AV779571
Length = 538
Score = 31.2 bits (69), Expect = 0.90
Identities = 21/55 (38%), Positives = 37/55 (67%)
Frame = -2
Query: 231 FNIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*KDKWLLKLQLRVQ*LLLLFV 285
++++L LLL +L+LLIV ++L + I V + + LLK++LR+Q LL+L +
Sbjct: 312 YHLLLSLLLLLLLLLLIVIMILSLTIE*VLML*V-----LLKMRLRLQQLLILLI 163
>TC12574
Length = 325
Score = 29.6 bits (65), Expect = 2.6
Identities = 12/23 (52%), Positives = 18/23 (78%)
Frame = +2
Query: 555 SMLDKFVVVFIDDILIYSETEEE 577
S + F++VFI+DIL Y+E +EE
Sbjct: 32 SFFEHFMIVFINDILSYTEDKEE 100
>TC14508 similar to AAP86659 (AAP86659) 26S proteasome subunit RPN5a,
partial (96%)
Length = 1957
Score = 29.6 bits (65), Expect = 2.6
Identities = 14/28 (50%), Positives = 16/28 (57%), Gaps = 1/28 (3%)
Frame = -3
Query: 398 QRGRLNFQLTLFQER-SRCRWHLTVCRR 424
Q GRL F Q R S CRW L+ CR+
Sbjct: 251 QSGRLGFSRLAHQSRASECRWRLSWCRK 168
>BP079571
Length = 414
Score = 24.3 bits (51), Expect(2) = 3.1
Identities = 8/12 (66%), Positives = 8/12 (66%)
Frame = -3
Query: 162 EKKCFRCGKKGH 173
E CF CGK GH
Sbjct: 271 ETHCFHCGKPGH 236
Score = 23.5 bits (49), Expect(2) = 3.1
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = -1
Query: 145 CFKCGEKGHKSNVCD 159
CF+ GE GH + CD
Sbjct: 357 CFRFGEVGHLARDCD 313
>TC17929
Length = 791
Score = 29.3 bits (64), Expect = 3.4
Identities = 10/35 (28%), Positives = 19/35 (53%)
Frame = +2
Query: 127 RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRD 161
++ +E + D+ C++CGE GHK C ++
Sbjct: 8 QIGEETSNRPNDSKFRQTCYRCGESGHKMRNCPKE 112
>TC18767
Length = 1004
Score = 28.9 bits (63), Expect = 4.5
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = +2
Query: 161 DEKKCFRCGKKGHTLADCKR 180
D +CF CG H+L +C R
Sbjct: 155 DASRCFNCGSYNHSLRECSR 214
>TC9269 similar to UP|AAR92258 (AAR92258) At1g03260, partial (8%)
Length = 586
Score = 27.7 bits (60), Expect = 10.0
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = +1
Query: 717 KRIILHMIWSWLRWFLF*KFGGIICTVPDLKCLVITKV*N 756
K ++ H+ WS+L IICTVP L+ +T+V N
Sbjct: 301 KAMLFHLTWSYLF---------IICTVPTLEI*FVTEVVN 393
>AW720443
Length = 406
Score = 27.7 bits (60), Expect = 10.0
Identities = 23/53 (43%), Positives = 29/53 (54%)
Frame = +3
Query: 233 IVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*KDKWLLKLQLRVQ*LLLLFV 285
++LL LL*+LVLLIVS LL I W + +L L+ Q LL L V
Sbjct: 216 MMLLSTLL*LLVLLIVSPLLSQSIQWTGF-----GRRMLLLEAGFQMLLSLMV 359
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.369 0.167 0.628
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,011,708
Number of Sequences: 28460
Number of extensions: 315143
Number of successful extensions: 4201
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2202
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 1726
Number of HSP's gapped (non-prelim): 2620
length of query: 1069
length of database: 4,897,600
effective HSP length: 100
effective length of query: 969
effective length of database: 2,051,600
effective search space: 1988000400
effective search space used: 1988000400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC141111.4


