

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
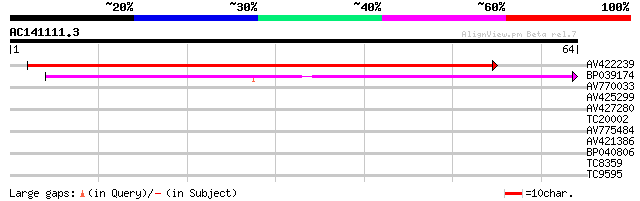
Score E
Sequences producing significant alignments: (bits) Value
AV422239 42 2e-05
BP039174 40 9e-05
AV770033 30 0.091
AV425299 26 1.3
AV427280 26 1.7
TC20002 similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding protein-l... 26 1.7
AV775484 25 2.9
AV421386 25 3.8
BP040806 24 5.0
TC8359 similar to UP|Q39436 (Q39436) SIEP1L protein precursor, p... 23 8.6
TC9595 similar to UP|Q946U9 (Q946U9) 5-enolpyruvylshikimate-3-ph... 23 8.6
>AV422239
Length = 387
Score = 42.0 bits (97), Expect = 2e-05
Identities = 17/53 (32%), Positives = 32/53 (60%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEK 55
+ +N +++ K+ N+ A+ +F ++ GR EPD +Y ++EGWG+ N K
Sbjct: 227 LAAFNGLLSALCKSRNVRKAQEIFDSMKGRFEPDLKTYSILLEGWGKDPNLPK 385
>BP039174
Length = 564
Score = 40.0 bits (92), Expect = 9e-05
Identities = 23/62 (37%), Positives = 35/62 (56%), Gaps = 2/62 (3%)
Frame = -2
Query: 5 VYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
VY ++ Y + N +GA+ VF + L G I PD+ +I+ +G AG EKAR +E
Sbjct: 554 VYKALLRAYSRIGNAEGAQRVFDAIQLAGII-PDDKICGLVIKAYGMAGQSEKARIAFEN 378
Query: 63 LK 64
+K
Sbjct: 377 MK 372
>AV770033
Length = 298
Score = 30.0 bits (66), Expect = 0.091
Identities = 15/50 (30%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Frame = -1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAG 51
++ YN ++ G K + A V + + P+ET+Y ++EG G AG
Sbjct: 268 VISYNIVLLGLSKVHRISDAIEVLAAMVDKGCRPNETTYTLLVEGIGYAG 119
>AV425299
Length = 323
Score = 26.2 bits (56), Expect = 1.3
Identities = 15/41 (36%), Positives = 22/41 (53%), Gaps = 1/41 (2%)
Frame = +3
Query: 13 YGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGN 52
Y K N++ A+ F L + +PD Y SMI G+ AG+
Sbjct: 57 YCKVGNLERAKEAFEVLTRKGFQPDIKVYNSMIMGYVNAGD 179
>AV427280
Length = 268
Score = 25.8 bits (55), Expect = 1.7
Identities = 15/60 (25%), Positives = 26/60 (43%), Gaps = 1/60 (1%)
Frame = +3
Query: 6 YNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN ++ Y ++ + GV + + PD SYR I +G + E EE++
Sbjct: 60 YNNIMCLYAQSDQHEKVPGVLAMMKEDGVSPDIFSYRVCINSYGARSDLENMEKLLEEIE 239
>TC20002 similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding protein-like,
partial (4%)
Length = 560
Score = 25.8 bits (55), Expect = 1.7
Identities = 15/49 (30%), Positives = 26/49 (52%)
Frame = -3
Query: 16 ASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
A +M+ A VF + +PD + +MI G+G EKA +Y+ ++
Sbjct: 210 AGDMNYAVSVFDRVD---KPDAFLWNTMIRGFGNTNQPEKAVLFYKRMQ 73
>AV775484
Length = 373
Score = 25.0 bits (53), Expect = 2.9
Identities = 8/12 (66%), Positives = 8/12 (66%)
Frame = +2
Query: 47 WGRAGNYEKARW 58
WG GNYE A W
Sbjct: 2 WGGQGNYENAHW 37
>AV421386
Length = 312
Score = 24.6 bits (52), Expect = 3.8
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = +1
Query: 5 VYNTMITGYGKASNMD 20
+YN +I G+GKA +D
Sbjct: 205 IYNILINGFGKAGKID 252
>BP040806
Length = 534
Score = 24.3 bits (51), Expect = 5.0
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +1
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL 29
V N +++GYGKA D E V ++
Sbjct: 409 VTQNIVLSGYGKAGKFDQMEKVLSSM 486
>TC8359 similar to UP|Q39436 (Q39436) SIEP1L protein precursor, partial
(50%)
Length = 767
Score = 23.5 bits (49), Expect = 8.6
Identities = 17/72 (23%), Positives = 25/72 (34%), Gaps = 12/72 (16%)
Frame = +1
Query: 1 MCIVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETS------------YRSMIEGWG 48
+C+ N G + A + T R+EP TS I W
Sbjct: 250 LCVWASNARSRSSGGSGRPTEATQLKKTPPSRLEPTVTSCLPTLTAESRGKQTPPIRAWW 429
Query: 49 RAGNYEKARWYY 60
R+G ++ A W Y
Sbjct: 430 RSGYFQMATWCY 465
>TC9595 similar to UP|Q946U9 (Q946U9) 5-enolpyruvylshikimate-3-phosphate
synthase B (3-phosphoshikimate
1-carboxyvinyltransferase) (EPSP synthase) (EPSPS) ,
partial (48%)
Length = 1099
Score = 23.5 bits (49), Expect = 8.6
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +2
Query: 24 GVFLTLGGRIEPDETSYRSMIEGWG 48
G TLG +E D+ + ++++EG G
Sbjct: 542 GALRTLGLHVEDDKATKQAIVEGCG 616
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,025,711
Number of Sequences: 28460
Number of extensions: 8301
Number of successful extensions: 43
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 43
length of query: 64
length of database: 4,897,600
effective HSP length: 40
effective length of query: 24
effective length of database: 3,759,200
effective search space: 90220800
effective search space used: 90220800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 49 (23.5 bits)
Medicago: description of AC141111.3


