

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140916.4 - phase: 0
(250 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
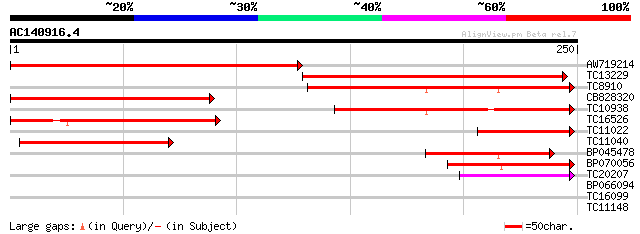
AW719214 137 2e-33
TC13229 weakly similar to UP|O81642 (O81642) IAA-ALA hydrolase (... 121 1e-28
TC8910 weakly similar to GB|AAB60293.1|887785|ATU23794 ILR1 {Ara... 104 2e-23
CB828320 100 3e-22
TC10938 weakly similar to GB|AAB60293.1|887785|ATU23794 ILR1 {Ar... 97 2e-21
TC16526 similar to UP|Q9SWX9 (Q9SWX9) Auxin CONJUGATE hydrolase ... 93 5e-20
TC11022 similar to UP|O81702 (O81702) Gr1-protein, partial (9%) 83 4e-17
TC11040 similar to UP|O81642 (O81642) IAA-ALA hydrolase (IAR3) (... 67 4e-12
BP045478 54 3e-08
BP070056 52 7e-08
TC20207 similar to UP|Q84XG9 (Q84XG9) IAA-amino acid hydrolase, ... 40 3e-04
BP066094 29 0.67
TC16099 similar to UP|Q9LT41 (Q9LT41) RCD1, partial (91%) 28 1.9
TC11148 homologue to UP|Q7X9B8 (Q7X9B8) Ferrous ion membrane tra... 26 7.4
>AW719214
Length = 439
Score = 137 bits (344), Expect = 2e-33
Identities = 70/129 (54%), Positives = 91/129 (70%)
Frame = +1
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ GALE+V AIF +HV P G + SR GPL+AG G F A+I+GK AA P + D
Sbjct: 52 IVESGALENVSAIFGLHVLPTLPVGEVASRSGPLMAGSGRFEAIINGKGGHAALPHTAID 231
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS VIS+Q +VSRE++PL+ QVV+V F GG + ++IPD V IGGTFRAFS S
Sbjct: 232 PILAASNVVISLQYLVSREADPLEPQVVTVAKFEGGGTFNVIPDYVKIGGTFRAFSTKSL 411
Query: 121 YQLLERIEQ 129
L +RIEQ
Sbjct: 412 EYLKQRIEQ 438
>TC13229 weakly similar to UP|O81642 (O81642) IAA-ALA hydrolase (IAR3)
(AT1G51760/F19C24_4), partial (23%)
Length = 558
Score = 121 bits (304), Expect = 1e-28
Identities = 55/117 (47%), Positives = 81/117 (69%)
Frame = +3
Query: 130 VIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGA 189
VI+ +A+V C A V F ++E +PPTVN+ ++EH + V+ LLG + P+MG+
Sbjct: 54 VIIGKAAVQRCNATVSFLDEEKPFFPPTVNNGDLHEHFQSVAKSLLGVNKVNGMEPLMGS 233
Query: 190 EDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIAERYL 246
ED+SFY QVIP FF +G+ N ++G + HSP+F ++EDALP GAA+HA++A RYL
Sbjct: 234 EDFSFYQQVIPGYFFMLGMENASIGHFESPHSPYFKVNEDALPYGAALHASLATRYL 404
>TC8910 weakly similar to GB|AAB60293.1|887785|ATU23794 ILR1 {Arabidopsis
thaliana;} , partial (6%)
Length = 587
Score = 104 bits (259), Expect = 2e-23
Identities = 49/120 (40%), Positives = 79/120 (65%), Gaps = 2/120 (1%)
Frame = +3
Query: 132 VQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNFRV-VPPMMGAE 190
+ QA+V+ C A V+FFE+ +YPPT+ND ++E + V+++LLG +PP+ +E
Sbjct: 3 IGQAAVHRCNATVNFFEEVSPLYPPTINDAGLHEQFRDVALNLLGADKVHFDLPPVTASE 182
Query: 191 DYSFYSQVIPSAFFYIGIRNETLG-STHTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
D+SFY +V+P FF++G+ + H HSPH I+E+ LP GAA+HA++A YL ++
Sbjct: 183 DFSFYQEVMPGYFFFLGMHKPSNDPRAHILHSPHLFINEEGLPYGAALHASLAVNYLQKY 362
>CB828320
Length = 539
Score = 100 bits (248), Expect = 3e-22
Identities = 50/90 (55%), Positives = 67/90 (73%)
Frame = +3
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++ GALE+V AIF +HV P G +GSR GP++AG G F A ISG+ AA P++S D
Sbjct: 264 ILDSGALENVSAIFGLHVLPTLPVGEVGSRSGPIMAGSGRFEAKISGRGGHAAIPQHSID 443
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSV 90
P+LAAS +IS+Q +VSRE++PLDSQVV+V
Sbjct: 444 PILAASNVIISLQHLVSREADPLDSQVVTV 533
>TC10938 weakly similar to GB|AAB60293.1|887785|ATU23794 ILR1 {Arabidopsis
thaliana;} , partial (6%)
Length = 518
Score = 97.4 bits (241), Expect = 2e-21
Identities = 47/107 (43%), Positives = 72/107 (66%), Gaps = 1/107 (0%)
Frame = +2
Query: 144 VDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNFRV-VPPMMGAEDYSFYSQVIPSA 202
V+FF + Y YPPT+ND ++E + V+I+LLG + +PPM AED+SFY +V+P
Sbjct: 5 VNFFGEVYPPYPPTINDGGLHEQFRYVAINLLGIDKAHIDMPPMTAAEDFSFYQKVMPGY 184
Query: 203 FFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
FF++G++ + H HSP+ IDE+ LP GAA+HA++A YL ++
Sbjct: 185 FFFLGMQKD--HRDHFLHSPYLMIDEEGLPYGAALHASLAINYLQKY 319
>TC16526 similar to UP|Q9SWX9 (Q9SWX9) Auxin CONJUGATE hydrolase (ILL5),
partial (54%)
Length = 853
Score = 92.8 bits (229), Expect = 5e-20
Identities = 49/98 (50%), Positives = 70/98 (71%), Gaps = 5/98 (5%)
Frame = +3
Query: 1 MIQDGALEDVEAIFAVHVSHEHPT-----GMIGSRPGPLLAGCGFFRAVISGKRASAANP 55
++ GALE+V AIF +H++ PT G SR GP++AG G F A+ISG+ AA P
Sbjct: 567 ILDSGALENVSAIFGLHMA---PTPLLQVGEAASRSGPVMAGSGHFEAIISGRGGHAAIP 737
Query: 56 RNSADPVLAASAAVISIQGIVSRESNPLDSQVVSVTSF 93
++ DPVLAAS+ +IS+Q +VSRE++PLDS+VV+V +F
Sbjct: 738 HSTIDPVLAASSVIISLQQLVSREADPLDSEVVTVATF 851
>TC11022 similar to UP|O81702 (O81702) Gr1-protein, partial (9%)
Length = 514
Score = 83.2 bits (204), Expect = 4e-17
Identities = 39/43 (90%), Positives = 40/43 (92%)
Frame = +3
Query: 207 GIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
GIRNETLGSTHTGHSP+F IDEDALP GAAVHATIAERYL EH
Sbjct: 3 GIRNETLGSTHTGHSPYFMIDEDALPTGAAVHATIAERYLIEH 131
>TC11040 similar to UP|O81642 (O81642) IAA-ALA hydrolase (IAR3)
(AT1G51760/F19C24_4), partial (49%)
Length = 787
Score = 66.6 bits (161), Expect = 4e-12
Identities = 32/68 (47%), Positives = 42/68 (61%)
Frame = +1
Query: 5 GALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSADPVLA 64
GALE+V AIF +HVSH P G + SR GP+ A FF A I G A P +S P+LA
Sbjct: 583 GALENVSAIFGLHVSHRVPXGEVASRSGPMFAXXAFFEAXIIGXGGHAXIPHHSIXPILA 762
Query: 65 ASAAVISI 72
AS ++++
Sbjct: 763 ASNVIVTL 786
>BP045478
Length = 444
Score = 53.5 bits (127), Expect = 3e-08
Identities = 25/58 (43%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Frame = -3
Query: 184 PPMMGAEDYSFYSQVIPSAFFYIGIRNETLG-STHTGHSPHFTIDEDALPIGAAVHAT 240
PP+ ED+SFY +VIP FF++G + + H HSP+ I E+ LP GA +HA+
Sbjct: 406 PPVTAFEDFSFYPKVIPGYFFFLGKQKASNDHRAHFVHSPYLVIHEEGLPYGAPLHAS 233
>BP070056
Length = 411
Score = 52.4 bits (124), Expect = 7e-08
Identities = 24/58 (41%), Positives = 36/58 (61%), Gaps = 2/58 (3%)
Frame = -1
Query: 194 FYSQVIPSAFFYIGIRNETLGS--THTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
FY +VIP FF +G+ + H+ HSP+ I+E+ LP GAA+HA++A YL +
Sbjct: 411 FYQKVIPGYFFLLGMHDNASNDIRAHSVHSPYLKINEEGLPYGAALHASLAINYLQTY 238
>TC20207 similar to UP|Q84XG9 (Q84XG9) IAA-amino acid hydrolase, partial
(6%)
Length = 486
Score = 40.4 bits (93), Expect = 3e-04
Identities = 19/51 (37%), Positives = 30/51 (58%)
Frame = +3
Query: 199 IPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
+ ++FF R H HSP+ I+E+ LP GAA+HA++A YL ++
Sbjct: 81 LATSFFLECRRPSNDHRAHFVHSPYLVINEEGLPYGAALHASLAVNYLEKY 233
Score = 35.0 bits (79), Expect = 0.012
Identities = 13/26 (50%), Positives = 19/26 (73%)
Frame = +2
Query: 184 PPMMGAEDYSFYSQVIPSAFFYIGIR 209
PP+ ED+SFY +VIP FF+ G++
Sbjct: 32 PPVPACEDFSFYLKVIPGYFFFFGMQ 109
>BP066094
Length = 532
Score = 29.3 bits (64), Expect = 0.67
Identities = 15/49 (30%), Positives = 24/49 (48%)
Frame = +2
Query: 130 VIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQK 178
+I+ Q +V S F E+ Y PP D++ +H+ K+ L G K
Sbjct: 137 IILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKKSLYGLK 283
>TC16099 similar to UP|Q9LT41 (Q9LT41) RCD1, partial (91%)
Length = 1182
Score = 27.7 bits (60), Expect = 1.9
Identities = 18/51 (35%), Positives = 25/51 (48%), Gaps = 4/51 (7%)
Frame = -1
Query: 23 PTGMIGSRPGPL--LAGCGFFRAVISGK--RASAANPRNSADPVLAASAAV 69
P GM G+RP P +GC V+S + R A+ S +L AS A+
Sbjct: 984 PAGMAGTRPDPTW*SSGCSHLLVVVSSRRQRLKVASLNMSGRQLLKASQAL 832
>TC11148 homologue to UP|Q7X9B8 (Q7X9B8) Ferrous ion membrane transport
protein DMT1, partial (33%)
Length = 622
Score = 25.8 bits (55), Expect = 7.4
Identities = 11/32 (34%), Positives = 15/32 (46%)
Frame = +3
Query: 7 LEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGC 38
+E+ A+ VSH H G G L +GC
Sbjct: 189 MEEAMAVHRARVSHVHSVSRSGEPGGGLTSGC 284
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,833,807
Number of Sequences: 28460
Number of extensions: 46547
Number of successful extensions: 194
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 187
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 187
length of query: 250
length of database: 4,897,600
effective HSP length: 88
effective length of query: 162
effective length of database: 2,393,120
effective search space: 387685440
effective search space used: 387685440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC140916.4


