

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
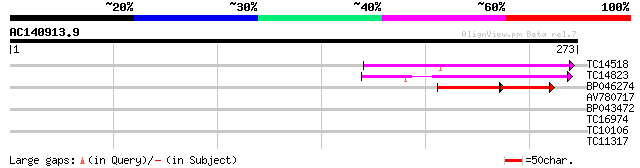
TC14518 similar to UP|Q93Y38 (Q93Y38) Transmembrane protein FT27... 67 4e-12
TC14823 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF... 64 3e-11
BP046274 32 4e-04
AV780717 28 2.2
BP043472 26 6.4
TC16974 UP|Q8GZD1 (Q8GZD1) Biogenesis protein, complete 26 6.4
TC10106 similar to GB|AAP12874.1|30017281|BT006225 At3g02700 {Ar... 26 6.4
TC11317 similar to UP|Q40363 (Q40363) NuM1 protein, partial (12%) 26 8.3
>TC14518 similar to UP|Q93Y38 (Q93Y38) Transmembrane protein FT27/PFT27-like
(At5g36290), partial (44%)
Length = 769
Score = 66.6 bits (161), Expect = 4e-12
Identities = 36/104 (34%), Positives = 59/104 (56%), Gaps = 2/104 (1%)
Frame = +1
Query: 171 ASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAAST--IVSTFLLVFVAEWGDKSFFSTI 228
+ S SQK + E+ E +L + T + +F+L F+AEWGD+S +TI
Sbjct: 16 SDSKTSQKKEMEEVEEKLEGGQGKTSFRRFFSRFCTPIFLESFILTFLAEWGDRSQIATI 195
Query: 229 ALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTSS 272
ALA + LGV G+ GH + T +AV+GGS+L + +S++ ++
Sbjct: 196 ALATHKNALGVAVGATIGHTICTSVAVVGGSMLASKISQRTVAT 327
>TC14823 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF|01169
Uncharacterized (transmembrane domain) protein family.
{Arabidopsis thaliana;}, complete
Length = 1231
Score = 63.9 bits (154), Expect = 3e-11
Identities = 36/103 (34%), Positives = 56/103 (53%), Gaps = 1/103 (0%)
Frame = +2
Query: 170 DASSSDSQKSDDEQKEAELA-VSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTI 228
+ +S DS K+DD++K+ + +S F + + F + F EWGDKS +TI
Sbjct: 461 NGASKDSNKADDDKKKNNRSFLSQFF---------SPIFLQAFSITFFGEWGDKSQLATI 613
Query: 229 ALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
LAA +P GV+ G + + T AV+GG L + +SEKV +
Sbjct: 614 GLAADENPFGVVLGGILAQTLCTTAAVMGGKSLASQISEKVVA 742
Score = 27.7 bits (60), Expect = 2.2
Identities = 15/46 (32%), Positives = 26/46 (55%)
Frame = +2
Query: 213 LVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGG 258
+ ++E GDK+FF+ LA V++G L+ V T+++ L G
Sbjct: 158 MTVLSEIGDKTFFAAAILAMRHPRRLVLSGCLSALIVMTILSALVG 295
>BP046274
Length = 487
Score = 31.6 bits (70), Expect(2) = 4e-04
Identities = 14/32 (43%), Positives = 20/32 (61%)
Frame = -2
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLG 238
I+ +F+L AEWGD+ +TIALA + G
Sbjct: 423 ILESFILTIPAEWGDRRPIATIALATPPNSAG 328
Score = 27.7 bits (60), Expect(2) = 4e-04
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -3
Query: 237 LGVIAGSLAGHGVATLIAVLGGSLLG 262
LG+ G+ GH + T +A++GGS G
Sbjct: 332 LGMAVGATIGHTLCTSVALVGGSTAG 255
>AV780717
Length = 444
Score = 27.7 bits (60), Expect = 2.2
Identities = 16/42 (38%), Positives = 25/42 (59%)
Frame = -2
Query: 191 SDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAA 232
SD + D + ++ AAST+V ++L++ S STIAL A
Sbjct: 428 SDNTDDMSSMMMAASTLVHSYLILHQKRLF*TSLISTIALLA 303
>BP043472
Length = 529
Score = 26.2 bits (56), Expect = 6.4
Identities = 13/34 (38%), Positives = 21/34 (61%), Gaps = 3/34 (8%)
Frame = -1
Query: 96 YFSLNWET---RLFSLQHC*QLEIQPVLFSLGHL 126
+F NW++ +F +Q *QL I P+L+ + HL
Sbjct: 394 FF*SNWKSFA*LVFRMQVL*QLAINPILYCIHHL 293
>TC16974 UP|Q8GZD1 (Q8GZD1) Biogenesis protein, complete
Length = 1406
Score = 26.2 bits (56), Expect = 6.4
Identities = 17/38 (44%), Positives = 24/38 (62%)
Frame = +2
Query: 227 TIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTF 264
T ALAA+ V+A +L G+ A+ V+GGSLL T+
Sbjct: 791 TFALAASPCSTPVLA-TLLGYVAASKDPVIGGSLLLTY 901
>TC10106 similar to GB|AAP12874.1|30017281|BT006225 At3g02700 {Arabidopsis
thaliana;}, partial (27%)
Length = 554
Score = 26.2 bits (56), Expect = 6.4
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +2
Query: 194 SGDGAGILAAASTIVSTFLLVFVAEWG 220
SG A LAAAST+VST L +G
Sbjct: 59 SGQAASYLAAASTVVSTPLRFMTNSFG 139
>TC11317 similar to UP|Q40363 (Q40363) NuM1 protein, partial (12%)
Length = 547
Score = 25.8 bits (55), Expect = 8.3
Identities = 34/136 (25%), Positives = 58/136 (42%), Gaps = 18/136 (13%)
Frame = -3
Query: 152 DDIAAVCLLVYFGVSTLLD----ASSSDSQKSDDEQKEAELAVSDFSGD----------- 196
++ ++ L +FG +TL+ SSS S++S E+ E ++ F+G
Sbjct: 512 EESSSELLSSFFGTTTLVGFFAITSSSSSEESSSEELELGAGLAFFAGVVPFFTEVFRGA 333
Query: 197 --GAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGH-GVATLI 253
GAG +++ S+ LL F+ SFF A +A L + S + + +
Sbjct: 332 TLGAG-FSSSDEESSSELLSFLTFCNFVSFFCFAATSAIFCFLALTCFSTSSSAALLAFL 156
Query: 254 AVLGGSLLGTFLSEKV 269
A LG + G S V
Sbjct: 155 ADLGAAATGAAASTLV 108
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,940,904
Number of Sequences: 28460
Number of extensions: 49847
Number of successful extensions: 314
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 310
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 313
length of query: 273
length of database: 4,897,600
effective HSP length: 89
effective length of query: 184
effective length of database: 2,364,660
effective search space: 435097440
effective search space used: 435097440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC140913.9


