

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140850.3 + phase: 0 /pseudo
(650 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
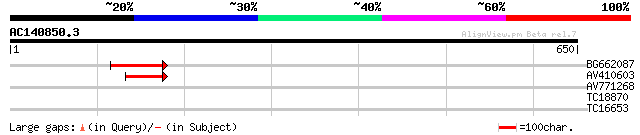
Score E
Sequences producing significant alignments: (bits) Value
BG662087 82 2e-16
AV410603 52 2e-07
AV771268 29 2.7
TC18870 similar to UP|Q9SGU7 (Q9SGU7) F1N19.25 (At1g64680), part... 28 4.6
TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfat... 27 7.9
>BG662087
Length = 373
Score = 82.4 bits (202), Expect = 2e-16
Identities = 37/65 (56%), Positives = 49/65 (74%)
Frame = +1
Query: 116 GKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREK 175
GK RM VD+ DLNKA PKD++PLP ID LVD + +++ S MD +SGY+ IKM P D +K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 176 TSFIT 180
T+F+T
Sbjct: 196 TAFMT 210
>AV410603
Length = 162
Score = 52.4 bits (124), Expect = 2e-07
Identities = 25/49 (51%), Positives = 33/49 (67%)
Frame = +1
Query: 133 KDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSFITH 181
KD+FP+P +D L+D S+ FS +D SGY+ I + PEDR KT F TH
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTH 156
>AV771268
Length = 514
Score = 28.9 bits (63), Expect = 2.7
Identities = 29/94 (30%), Positives = 42/94 (43%), Gaps = 8/94 (8%)
Frame = -3
Query: 306 ITSPDSYLT*PQPAGRSSSYSGRISLLYGTMNAKKLLIASRITCCNHLSLS---HPWKEG 362
IT SY T P G+SS +S ++ L+Y + L A I +L L + W E
Sbjct: 368 ITHVLSYPTMPPS*GKSSPHSNKLILMY--ILRSFLASAFPIHYAFNLRLFVSIYFWFER 195
Query: 363 L*LCIC-----QCLMNLWDAYLVNKMRLERKNML 391
L + +C L+N W Y+ + R NML
Sbjct: 194 LMIWLCPPNEENMLINHWQHYVARPLVTSRMNML 93
>TC18870 similar to UP|Q9SGU7 (Q9SGU7) F1N19.25 (At1g64680), partial (51%)
Length = 747
Score = 28.1 bits (61), Expect = 4.6
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +3
Query: 547 PRRGITSLLPPGFCSNVQTIWPNTK 571
P RG+ S+LPPG + + ++P TK
Sbjct: 12 PSRGLLSMLPPGAPAQFRKLFPPTK 86
>TC16653 similar to UP|Q8W1A1 (Q8W1A1) Adenosine 5'-phosphosulfate
reductase, partial (69%)
Length = 1242
Score = 27.3 bits (59), Expect = 7.9
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +2
Query: 74 GLIQIWLSRLRVRFKSRLMRVFLMTVEYPEW 104
G + I+L RLR +S L FLM + + +W
Sbjct: 485 GNLSIFLMRLRSTMESTLSTCFLMLLRFKDW 577
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.347 0.151 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,141,000
Number of Sequences: 28460
Number of extensions: 180784
Number of successful extensions: 1598
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1583
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1598
length of query: 650
length of database: 4,897,600
effective HSP length: 96
effective length of query: 554
effective length of database: 2,165,440
effective search space: 1199653760
effective search space used: 1199653760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140850.3


