

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140720.4 + phase: 0
(653 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
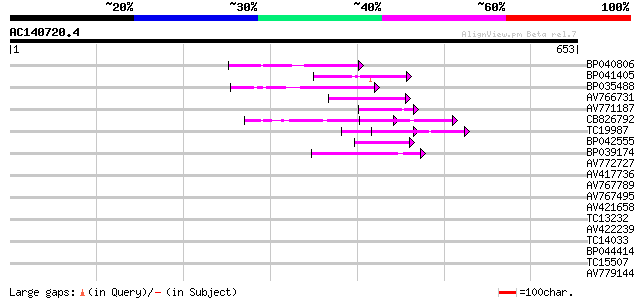
Score E
Sequences producing significant alignments: (bits) Value
BP040806 55 4e-08
BP041405 55 5e-08
BP035488 52 2e-07
AV766731 49 3e-06
AV771187 47 1e-05
CB826792 45 4e-05
TC19987 weakly similar to UP|Q7XJT3 (Q7XJT3) At2g17140 protein, ... 45 4e-05
BP042555 45 5e-05
BP039174 42 2e-04
AV772727 40 0.001
AV417736 40 0.001
AV767789 39 0.003
AV767495 37 0.008
AV421658 37 0.010
TC13232 34 0.085
AV422239 33 0.11
TC14033 similar to GB|AAO63396.1|28950945|BT005332 At5g13570 {Ar... 30 0.94
BP044414 30 0.94
TC15507 weakly similar to GB|AAP88326.1|32815835|BT009692 At1g61... 30 0.94
AV779144 30 1.6
>BP040806
Length = 534
Score = 55.1 bits (131), Expect = 4e-08
Identities = 36/156 (23%), Positives = 71/156 (45%), Gaps = 1/156 (0%)
Frame = +1
Query: 253 KDHRKFQSRF-VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAG 311
K+H ++ + Y KL +LGK+ +P A Q+F M+ P Y ++ ++
Sbjct: 61 KEHSFYEPKEGTYMKLXVLLGKSGQPHRARQLFTTMIEE-GCDPSPETYTALLAAYCRSN 237
Query: 312 LLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQ 371
LL E +I+ M+ P +PDV Y+ ++ CV + ++ V +++
Sbjct: 238 LLDEAFSILNEMKN-------------HPLCQPDVFTYSTLIKPCVDAFKFDLVELLYED 378
Query: 372 MRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKM 407
M S+ PN T + + ++G +D + ++ M
Sbjct: 379 MAARSIMPNTVTQNIVLSGYGKAGKFDQMEKVLSSM 486
Score = 33.1 bits (74), Expect = 0.14
Identities = 20/100 (20%), Positives = 47/100 (47%)
Frame = +1
Query: 513 MQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGH 572
+++ C P+ T +L Y ++++ A + E+K +PD +TY+ +++
Sbjct: 169 IEEGCDPSPETYTALLAAYCRSNLLDEAFSILNEMKNHPL-CQPDVFTYSTLIKPCVDAF 345
Query: 573 QWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNI 612
+++ E +Y++M + +L +AGK D +
Sbjct: 346 KFDLVELLYEDMAARSIMPNTVTQNIVLSGYGKAGKFDQM 465
>BP041405
Length = 546
Score = 54.7 bits (130), Expect = 5e-08
Identities = 38/148 (25%), Positives = 64/148 (42%), Gaps = 35/148 (23%)
Frame = +3
Query: 350 NAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR 409
N +LN V + + V +++Q+++ L PN TY + M+ + + G D+VH +F++M+
Sbjct: 24 NFLLNRLVGHGKVEMVLAIYEQLKRLGLSPNHYTYAIVMKALYRKG--DVVH-VFQEMEE 194
Query: 410 NGEVP-----------------------------------EALTYKVMVRTFWKEGKVDE 434
G P E Y ++ F E K+DE
Sbjct: 195 AGVTPDSYCNAVLIEGLCKNHRSDWGYQFLQEFRKVNAPIEVYAYTAVIHGFCNEMKLDE 374
Query: 435 AVKAVRDMERRGVMGTASVYYELACCLC 462
A V DMER+G++ ++Y L C C
Sbjct: 375 AESVVLDMERQGLVPDVNIYSALICGYC 458
>BP035488
Length = 534
Score = 52.4 bits (124), Expect = 2e-07
Identities = 46/175 (26%), Positives = 77/175 (43%), Gaps = 4/175 (2%)
Frame = +3
Query: 255 HRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLK 314
HR ++ V VLG A K + ++M R PD+ Y + L + G L
Sbjct: 60 HRDIKTWNVILNGWCVLGNAHEAKRVWK--DIMASKCR--PDLFTYATFIKALTKKGKLG 227
Query: 315 ELLNIVECMRQKPETLKYMYRKNWDP--TLEPDVVIYNAVLNACVPSKQWKGVSWVFQQM 372
L + +R W+ +PDVVI N +++A K+ VFQ M
Sbjct: 228 TALKL--------------FRGMWNEGCNCKPDVVICNCIIDALCFKKRVPEALEVFQDM 365
Query: 373 RKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR--NGEVPEALTYKVMVRT 425
++ +PN ATY ++ + + + V+EL E M+R +P A+TY ++ +
Sbjct: 366 KERGCEPNVATYNSLIKHLCKIRRMEKVYELVEDMERKKGSCMPNAVTYSCLLNS 530
Score = 27.7 bits (60), Expect = 6.1
Identities = 14/57 (24%), Positives = 26/57 (45%)
Frame = +3
Query: 511 EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEA 567
+ M C P++ T T +K ++ TA LF + + +PD N +++A
Sbjct: 144 DIMASKCRPDLFTYATFIKALTKKGKLGTALKLFRGMWNEGCNCKPDVVICNCIIDA 314
>AV766731
Length = 576
Score = 48.9 bits (115), Expect = 3e-06
Identities = 24/94 (25%), Positives = 47/94 (49%)
Frame = -3
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+F +++ ++PN Y + ++ + + G + E+F+ + G +TY VM+ +
Sbjct: 376 LFMKVKDPKIQPNIPPYPVIIDGLCKVGRLKIAQEIFQVLLSEGYNLNVMTYTVMINGYC 197
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
KEG +DEA + ME G + A + + C L
Sbjct: 196 KEGLLDEAQALLSKMEDNGCIPNAVNFQTIICAL 95
>AV771187
Length = 484
Score = 47.0 bits (110), Expect = 1e-05
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 2/71 (2%)
Frame = -3
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMER--RGVMGTASVYYELAC 459
ELFE M++ G P +TY ++R + ++D AV+ +RDM+R G+ ++S Y +
Sbjct: 221 ELFEDMKKKGCAPNRVTYDSLIRYYSATNEIDRAVEVLRDMQRLNHGIPSSSS-YTPIIH 45
Query: 460 CLCNCGRWQDA 470
LC GR +A
Sbjct: 44 ALCEAGRVAEA 12
>CB826792
Length = 485
Score = 45.1 bits (105), Expect = 4e-05
Identities = 41/178 (23%), Positives = 76/178 (42%)
Frame = +3
Query: 271 LGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL 330
L K R KEA+ + M N V PD+ Y+S L+K L C +K +
Sbjct: 3 LAKGRLVKEAVNLLQTMEKN-NVTPDVVTYNS---------LIKPL-----CKNRKIDEA 137
Query: 331 KYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEV 390
K ++ + P + ++A ++ V + +MR+ P TY + +
Sbjct: 138 KEVFNDMMKRNITPTIRTFHAFFRILRVEEE---VFELLDKMRELGCYPTIETYIMLIRK 308
Query: 391 MLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVM 448
+ D V +++ M+ +G + +Y V++ + GKV EA +M+R+G +
Sbjct: 309 FCRWRKLDEVFKIWNMMREDGVSHDRSSYIVLIHGLFLNGKVKEAHDYYIEMQRKGFL 482
Score = 40.8 bits (94), Expect = 7e-04
Identities = 26/113 (23%), Positives = 50/113 (44%)
Frame = +3
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
L + M++N P+ +TY +++ K K+DEA + DM +R + T ++ L
Sbjct: 39 LLQTMEKNNVTPDVVTYNSLIKPLCKNRKIDEAKEVFNDMMKRNITPTIRTFHAFFRIL- 215
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD 515
R ++ E+ R P T+ +IR +D+ I+ M++
Sbjct: 216 ---RVEEEVFELLDKMRELGCYPTIETYIMLIRKFCRWRKLDEVFKIWNMMRE 365
>TC19987 weakly similar to UP|Q7XJT3 (Q7XJT3) At2g17140 protein, partial
(12%)
Length = 350
Score = 45.1 bits (105), Expect = 4e-05
Identities = 29/114 (25%), Positives = 54/114 (46%), Gaps = 1/114 (0%)
Frame = +3
Query: 417 LTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEK 476
+TY + F KEGK+ A++ ++DMER G T Y L L + G+ + +++
Sbjct: 3 VTYDTFIWKFCKEGKISSALRVLKDMERNGCSKTLQTYNSLILGLGSKGQIFEMYGLMDE 182
Query: 477 IKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLK 529
++ P T+ +I +GG +D + M D +PN+ + ++K
Sbjct: 183 MRERGIC-PDICTYNNVISCLCEGGKTEDATSLLHEMLDKGISPNISSFKILIK 341
Score = 41.2 bits (95), Expect = 5e-04
Identities = 22/89 (24%), Positives = 38/89 (41%)
Frame = +3
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY + + G + + M+RNG TY ++ +G++ E + +M
Sbjct: 6 TYDTFIWKFCKEGKISSALRVLKDMERNGCSKTLQTYNSLILGLGSKGQIFEMYGLMDEM 185
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDAT 471
RG+ Y + CLC G+ +DAT
Sbjct: 186 RERGICPDICTYNNVISCLCEGGKTEDAT 272
Score = 28.9 bits (63), Expect = 2.7
Identities = 16/93 (17%), Positives = 43/93 (46%)
Frame = +3
Query: 297 MAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNAC 356
+ Y+S+ + LG G + E+ +++ MR++ + PD+ YN V++
Sbjct: 105 LQTYNSLILGLGSKGQIFEMYGLMDEMRERG--------------ICPDICTYNNVISCL 242
Query: 357 VPSKQWKGVSWVFQQMRKSSLKPNGATYGLAME 389
+ + + + +M + PN +++ + ++
Sbjct: 243 CEGGKTEDATSLLHEMLDKGISPNISSFKILIK 341
>BP042555
Length = 510
Score = 44.7 bits (104), Expect = 5e-05
Identities = 21/69 (30%), Positives = 31/69 (44%)
Frame = +3
Query: 398 DLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
D H + + M G P TY ++ + + + EAV +RDM +RG YY L
Sbjct: 234 DKAHNMLKYMIVRGFSPSVATYNELIHGYCRVDRFKEAVGILRDMAKRGXSPDVDTYYPL 413
Query: 458 ACCLCNCGR 466
C C G+
Sbjct: 414 ICGFCQTGK 440
>BP039174
Length = 564
Score = 42.4 bits (98), Expect = 2e-04
Identities = 33/132 (25%), Positives = 59/132 (44%)
Frame = -2
Query: 348 IYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKM 407
+Y A+L A +G VF ++ + + P+ GL ++ +G + FE M
Sbjct: 554 VYKALLRAYSRIGNAEGAQRVFDAIQLAGIIPDDKICGLVIKAYGMAGQSEKARIAFENM 375
Query: 408 QRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRW 467
+R G P ++ + KE K++ A++ + D+E+ G+M V E + L R
Sbjct: 374 KRAGIEPTDRCIGSVLVAYEKESKLNTALEFLIDLEKEGIM----VGEEASAILAGWFRK 207
Query: 468 QDATLEVEKIKR 479
EVE + R
Sbjct: 206 LGVVEEVELVLR 171
>AV772727
Length = 528
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/91 (23%), Positives = 46/91 (50%)
Frame = -2
Query: 346 VVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFE 405
V+ Y+++L+A + + ++++ ++P+ TY + ++ + +SG ELF+
Sbjct: 527 VITYHSLLDALCKNHHVDKAIALLERVKDKGIQPDMYTYNIIIDGLCKSGRVKDAQELFQ 348
Query: 406 KMQRNGEVPEALTYKVMVRTFWKEGKVDEAV 436
+ G +TY +M+ EG DEA+
Sbjct: 347 DLLIKGYRLNVVTYNIMINGLCIEGLSDEAL 255
Score = 38.1 bits (87), Expect = 0.004
Identities = 21/97 (21%), Positives = 49/97 (49%)
Frame = -2
Query: 523 TVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYK 582
T +++L +N A L E VK ++PD YTYN++++ + + + + +++
Sbjct: 521 TYHSLLDALCKNHHVDKAIALLERVK--DKGIQPDMYTYNIIIDGLCKSGRVKDAQELFQ 348
Query: 583 EMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRS 619
++++ GY L+ + ++ G D L + I++
Sbjct: 347 DLLIKGYRLNVVTYNIMINGLCIEGLSDEALALLIQN 237
>AV417736
Length = 428
Score = 40.4 bits (93), Expect = 0.001
Identities = 28/106 (26%), Positives = 46/106 (42%)
Frame = +1
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+EPD+ Y+ + V S Q + MRK L+PN + + +G D
Sbjct: 43 IEPDIHAYSILAKGYVRSGQPGKAEALLTSMRKYGLQPNVVIFTTIISGWCTAGKMDRAV 222
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGV 447
L EKM G TY+ ++ + + + +A + + ME RGV
Sbjct: 223 RLHEKMHEMGIPSNLKTYETLMWGYGEAKQPWKAEELLVTMEERGV 360
>AV767789
Length = 560
Score = 38.9 bits (89), Expect = 0.003
Identities = 24/81 (29%), Positives = 41/81 (49%)
Frame = -1
Query: 519 PNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFE 578
P V T N ++ Y++ S L +E+ A +L+PD+ TY+ M+ A R +
Sbjct: 503 PTVMTYNMLMNAYARGGQHSKLPQLLKEM--AALNLKPDSVTYSTMIYAFVRVRDFRRAF 330
Query: 579 HVYKEMILSGYHLDQNKHLPL 599
+K+MI SG +D + + L
Sbjct: 329 FYHKQMIKSGQVMDVDSYQKL 267
Score = 28.5 bits (62), Expect = 3.6
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = -1
Query: 551 KSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
K L P TYN+++ A +RG Q + KEM
Sbjct: 518 KIGLHPTVMTYNMLMNAYARGGQHSKLPQLLKEM 417
>AV767495
Length = 586
Score = 37.4 bits (85), Expect = 0.008
Identities = 16/57 (28%), Positives = 34/57 (59%)
Frame = +2
Query: 332 YMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAM 388
+++RK + ++P+VV ++A+LNAC K ++ S + ++R + G +GL +
Sbjct: 389 WLFRKMHEMEIKPNVVTFSAILNACSNCKSFEDASKLLDELRLFDNQVYGVAHGLLL 559
>AV421658
Length = 281
Score = 37.0 bits (84), Expect = 0.010
Identities = 24/85 (28%), Positives = 39/85 (45%), Gaps = 2/85 (2%)
Frame = +1
Query: 372 MRKSSLKPNGATYGLAMEVMLQSGNYDL--VHELFEKMQRNGEVPEALTYKVMVRTFWKE 429
M + L P+ TY + + DL E+ +M G +P+A TY+ ++RT +
Sbjct: 7 MTERDLSPDVVTYNTLISKFCKLKEPDLEKAFEMKAEMVHKGILPDADTYEPLIRTLCLQ 186
Query: 430 GKVDEAVKAVRDMERRGVMGTASVY 454
++ EA R+M R GV Y
Sbjct: 187 QRLSEAYDLFREMLRWGVSPNNETY 261
>TC13232
Length = 551
Score = 33.9 bits (76), Expect = 0.085
Identities = 15/53 (28%), Positives = 27/53 (50%)
Frame = +1
Query: 414 PEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGR 466
P TY +++R F E ++D A+ M RG++ V++ L LC+ +
Sbjct: 19 PTVSTYDIILRMFCDEERLDMAMAVWDQMRARGILPGMHVFFVLISALCHANK 177
>AV422239
Length = 387
Score = 33.5 bits (75), Expect = 0.11
Identities = 20/79 (25%), Positives = 39/79 (49%)
Frame = +2
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y L +E + + Y ++ ++ KM+ G + T+ +++R + + KVDEAV M+
Sbjct: 29 YHLMIESLARIRQYQIMWDIVTKMRNKGML-NVETFCIVMRKYARAHKVDEAVYTFNVMD 205
Query: 444 RRGVMGTASVYYELACCLC 462
+ V + + L LC
Sbjct: 206 KYEVPQNLAAFNGLLSALC 262
>TC14033 similar to GB|AAO63396.1|28950945|BT005332 At5g13570 {Arabidopsis
thaliana;}, partial (52%)
Length = 583
Score = 30.4 bits (67), Expect = 0.94
Identities = 22/92 (23%), Positives = 39/92 (41%)
Frame = +2
Query: 347 VIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEK 406
+I + C+ K WKG SW F + +KS + + A+ +L+ +D+ K
Sbjct: 218 IILDETYERCLLVKGWKGTSWSFPRGKKSK---DEEDHACAVREVLEETGFDV-----SK 373
Query: 407 MQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
+ E E + + VR + G D+ A
Sbjct: 374 LLNKDEYLEVIFGQQRVRLYIIAGVKDDTAFA 469
>BP044414
Length = 536
Score = 30.4 bits (67), Expect = 0.94
Identities = 18/76 (23%), Positives = 36/76 (46%)
Frame = -1
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
E ++M++ P+ + +++ F EG +DEA K ++E+ + + L L
Sbjct: 527 EFLQEMEKKDVKPDLFSLNAIIKGFVNEGNLDEAKKWFGEIEKSENDPDKNTFATLVPFL 348
Query: 462 CNCGRWQDATLEVEKI 477
C G + A V++I
Sbjct: 347 CEKGDLKTAIDVVKEI 300
>TC15507 weakly similar to GB|AAP88326.1|32815835|BT009692 At1g61870
{Arabidopsis thaliana;}, partial (24%)
Length = 746
Score = 30.4 bits (67), Expect = 0.94
Identities = 20/69 (28%), Positives = 34/69 (48%), Gaps = 3/69 (4%)
Frame = +3
Query: 429 EGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA---TLEVEKIKRLPHAKP 485
+G + EA K DM++RG + Y+ LA LC G ++ A T E +P+
Sbjct: 81 KGTLYEAKKVFADMKKRGYKPESPCYFTLAHFLCEGGDFESAFEITKESMDKGWVPNFTT 260
Query: 486 LEVTFTGMI 494
++ TG++
Sbjct: 261 MKKLVTGLV 287
>AV779144
Length = 544
Score = 29.6 bits (65), Expect = 1.6
Identities = 17/70 (24%), Positives = 32/70 (45%)
Frame = -3
Query: 349 YNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQ 408
Y +++ C+ G WV + + P Y A++ M G + ++++EKM
Sbjct: 536 YLSLIRCCI-----YGELWVRLEEILTHTTPYATLYNAAVQGMCLRGKINFANKVYEKML 372
Query: 409 RNGEVPEALT 418
+G P+A T
Sbjct: 371 ESGLQPDAKT 342
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,189,411
Number of Sequences: 28460
Number of extensions: 148924
Number of successful extensions: 920
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 886
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 917
length of query: 653
length of database: 4,897,600
effective HSP length: 96
effective length of query: 557
effective length of database: 2,165,440
effective search space: 1206150080
effective search space used: 1206150080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140720.4


