

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.15 - phase: 0
(682 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
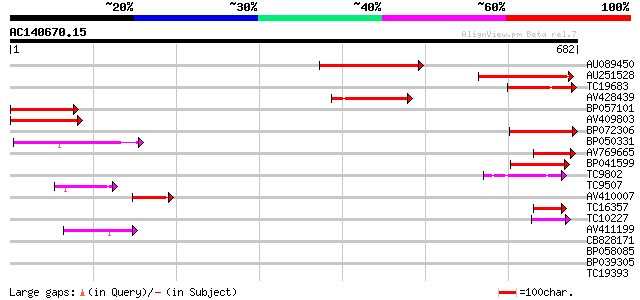
Sequences producing significant alignments: (bits) Value
AU089450 197 6e-51
AU251528 149 2e-36
TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein (AT2G052... 146 1e-35
AV428439 140 7e-34
BP057101 117 6e-27
AV409803 90 1e-18
BP072306 78 4e-15
BP050331 69 2e-12
AV769665 56 2e-08
BP041599 55 5e-08
TC9802 weakly similar to UP|AAS49059 (AAS49059) At5g53150, parti... 55 5e-08
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 54 1e-07
AV410007 50 1e-06
TC16357 weakly similar to UP|Q8DZP0 (Q8DZP0) Bacterial luciferas... 44 7e-05
TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%) 42 3e-04
AV411199 42 4e-04
CB828171 39 0.003
BP058085 39 0.003
BP039305 39 0.004
TC19393 36 0.023
>AU089450
Length = 383
Score = 197 bits (500), Expect = 6e-51
Identities = 98/125 (78%), Positives = 106/125 (84%)
Frame = +3
Query: 373 PAFDARKLLIEKARTEIRKKLEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVS 432
PAFDARKLLIEKARTEIRKKLEEMKLAS+T AAV E +KSQA+ GQVK T T L+VS
Sbjct: 9 PAFDARKLLIEKARTEIRKKLEEMKLASKTVAAVNEREKSQAEAGQVKSGSHTNTVLDVS 188
Query: 433 DNQLEHRKTVPVTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVS 492
N EH K PV++TVPDPDFHDFDKDRSE CF+PKQIWAL DEEDGMPRLYCLIREV+S
Sbjct: 189 GNHFEHGKIGPVSMTVPDPDFHDFDKDRSEECFRPKQIWAL*DEEDGMPRLYCLIREVIS 368
Query: 493 VNPFK 497
NPFK
Sbjct: 369 FNPFK 383
>AU251528
Length = 388
Score = 149 bits (375), Expect = 2e-36
Identities = 72/115 (62%), Positives = 85/115 (73%)
Frame = +3
Query: 564 GDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYKRNA 623
GD+WA+YRNWS DWN T DEV H++D+VEVL+D+SEE G+ + PL+K+ GFKTV+ +
Sbjct: 6 GDIWAIYRNWSPDWNELTADEVMHKFDVVEVLEDFSEEHGVTIIPLVKVAGFKTVFHHHI 185
Query: 624 DKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDELLHAAIE 678
D IR IPR EMLRFSHQVPS LL G EA N P C LDPAATP E LH IE
Sbjct: 186 DPREIRVIPREEMLRFSHQVPSCLLTGLEAQNAPKGCRVLDPAATPFE-LHQVIE 347
>TC19683 similar to UP|Q9S7L6 (Q9S7L6) AT2G05250 protein
(AT2G05250/F5G3.15), partial (9%)
Length = 555
Score = 146 bits (368), Expect = 1e-35
Identities = 72/84 (85%), Positives = 79/84 (93%), Gaps = 1/84 (1%)
Frame = +1
Query: 599 SEELGLCVSPLIKLDGFKTVYKRNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPD 658
SEELG+CV+PLIKL GFKTVYK NADKSAI++IPRREMLRFSHQVPSWLLKG EASNLP+
Sbjct: 16 SEELGVCVTPLIKLSGFKTVYKSNADKSAIKWIPRREMLRFSHQVPSWLLKG-EASNLPE 192
Query: 659 KCWDLDPAATPDELLH-AAIEANA 681
+CWDLDPAATPD+LLH AA EANA
Sbjct: 193 RCWDLDPAATPDDLLHAAATEANA 264
>AV428439
Length = 292
Score = 140 bits (353), Expect = 7e-34
Identities = 68/97 (70%), Positives = 80/97 (82%)
Frame = +3
Query: 388 EIRKKLEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVSDNQLEHRKTVPVTIT 447
EIRKKLEEMKL+SE AA + E +K+QADVGQVK + C + L+V +LEH K P++IT
Sbjct: 3 EIRKKLEEMKLSSE-AATLKEREKAQADVGQVKRETCRRAVLSVPGLELEHSKAGPISIT 179
Query: 448 VPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLY 484
VPD DFHDFDKDR+E CF+PKQIWALYDEEDGMPRLY
Sbjct: 180 VPDSDFHDFDKDRAEECFRPKQIWALYDEEDGMPRLY 290
>BP057101
Length = 546
Score = 117 bits (293), Expect = 6e-27
Identities = 56/82 (68%), Positives = 71/82 (86%)
Frame = +3
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
++A KEEAL+A+E AE+RF+ RDF GAKSYA+KAKTLCP +EGISQ+V TFEVYIAS++
Sbjct: 300 VQANKEEALQALEIAERRFAQRDFAGAKSYAVKAKTLCPEVEGISQMVATFEVYIASEIK 479
Query: 61 CNGELDWYSIMGLNPSTNIEAV 82
NGELD+Y+I+GL PS + EAV
Sbjct: 480 HNGELDYYAILGLKPSADKEAV 545
>AV409803
Length = 413
Score = 89.7 bits (221), Expect = 1e-18
Identities = 39/87 (44%), Positives = 62/87 (70%)
Frame = +1
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
M+ K+EA +A E AEK+FS R++ GAK +ALKA L P LEG+ Q ++ VYI+++
Sbjct: 148 MDCNKDEAARAKEIAEKKFSLREYAGAKKFALKALNLYPALEGVPQFLSILNVYISAENK 327
Query: 61 CNGELDWYSIMGLNPSTNIEAVKKQYK 87
NG++DWY I+G++P + E ++K+Y+
Sbjct: 328 INGQMDWYGILGVDPFADEETIRKKYR 408
>BP072306
Length = 399
Score = 78.2 bits (191), Expect = 4e-15
Identities = 44/82 (53%), Positives = 52/82 (62%), Gaps = 1/82 (1%)
Frame = -1
Query: 602 LGLCVSPLIKLDGFKTVYKRNADKSAIRYIPRRE-MLRFSHQVPSWLLKGEEASNLPDKC 660
LG+CVSPL KLDGF+TVYKRN D+ IR+IPR E F + +G+ +
Sbjct: 399 LGVCVSPLTKLDGFETVYKRNTDQCVIRWIPRHEKCCAFHTKYLHGC*RGKRLAICRKGI 220
Query: 661 WDLDPAATPDELLHAAIEANAS 682
LDPAATPDELL AA EAN S
Sbjct: 219 GTLDPAATPDELLQAATEANTS 154
>BP050331
Length = 519
Score = 69.3 bits (168), Expect = 2e-12
Identities = 53/169 (31%), Positives = 81/169 (47%), Gaps = 13/169 (7%)
Frame = +3
Query: 5 KEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQ------ 58
K E + +E AE++ RD G++ AL A+ P LEG Q++ +V A++
Sbjct: 54 KSEGERLLEIAEQQXQKRDLKGSRQLALIAQENEPLLEGSDQILAIIDVLEAAEKPLTTT 233
Query: 59 --VTCNGE--LDWYSIMGLN--PSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSE 112
T N LD+Y+I+ ++ S ++ +K+QY+++A LLHPD N + F LVS
Sbjct: 234 ASTTTNNRNNLDFYAILQVDHRDSQDLNLIKRQYRRLALLLHPDKNLFSFSHHTFDLVSH 413
Query: 113 AWARLSGSYDMK-RNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSC 160
AWA LS + +A LG G +FWT C C
Sbjct: 414 AWALLSDPAQKEIYDAGLGCAPG-----------------SFWTACPYC 509
>AV769665
Length = 553
Score = 55.8 bits (133), Expect = 2e-08
Identities = 28/50 (56%), Positives = 35/50 (70%)
Frame = -3
Query: 631 IPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDELLHAAIEAN 680
IP+ EM FS QVPS+ L G+E+ N P C +LDPAATP +LL A EA+
Sbjct: 551 IPKVEMFCFSLQVPSYSLTGQESHNAPKGCRELDPAATPLDLLQATTEAD 402
>BP041599
Length = 540
Score = 54.7 bits (130), Expect = 5e-08
Identities = 31/73 (42%), Positives = 45/73 (61%), Gaps = 2/73 (2%)
Frame = -3
Query: 603 GLCVSPLIKLDGFKTVYKRNADKS-AIRYIPRREMLRFSHQVPSWLLKG-EEASNLPDKC 660
GL ++ L K+DG+KTV+KR S AIR++ + M SHQ+P+ +E L + C
Sbjct: 529 GLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDC 350
Query: 661 WDLDPAATPDELL 673
W+LDPA+ P LL
Sbjct: 349 WELDPASLPSYLL 311
>TC9802 weakly similar to UP|AAS49059 (AAS49059) At5g53150, partial (3%)
Length = 593
Score = 54.7 bits (130), Expect = 5e-08
Identities = 36/102 (35%), Positives = 54/102 (52%), Gaps = 2/102 (1%)
Frame = +3
Query: 570 YRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYK--RNADKSA 627
YR W+ S D +YD+V++LD +L + V L + GF +V+K +N +
Sbjct: 3 YRKWTAKIKPS--DLQTWEYDIVQILD--VNDLRIAVVHLELVKGFPSVFKVQKNGGSAV 170
Query: 628 IRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATP 669
IP+ E+LRFSHQ+P+ +K E C +LDP A P
Sbjct: 171 TMKIPKAELLRFSHQIPA--IKLTEKHGSVKGCCELDPGALP 290
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 53.5 bits (127), Expect = 1e-07
Identities = 31/80 (38%), Positives = 47/80 (58%), Gaps = 4/80 (5%)
Frame = +1
Query: 54 YIASQVTCNGEL----DWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHL 109
Y QV+ E+ ++Y I+GL S +E V+K Y+K++ +HPD NK GA+ AF
Sbjct: 469 YTTEQVSIIREIKRKKNYYEILGLEKSCTVEDVRKSYRKLSLKVHPDKNKAPGAEEAFKA 648
Query: 110 VSEAWARLSGSYDMKRNAQL 129
VS+A+ L G+ + KR L
Sbjct: 649 VSKAFQCL-GNEESKRKYDL 705
>AV410007
Length = 425
Score = 50.1 bits (118), Expect = 1e-06
Identities = 22/50 (44%), Positives = 33/50 (66%)
Frame = +3
Query: 148 GNQDTFWTICTSCKVQYEYLRKYVNKKLSCKNCRGIFIALETAPANGSFP 197
G++ TFWT C C V+Y+Y R+ +NK L C++C+ FIA + A G+ P
Sbjct: 276 GSRPTFWTHCPFCSVKYQYYREVLNKPLRCQHCQKPFIACD-ANVQGTSP 422
>TC16357 weakly similar to UP|Q8DZP0 (Q8DZP0) Bacterial luciferase family
protein, partial (6%)
Length = 572
Score = 44.3 bits (103), Expect = 7e-05
Identities = 18/39 (46%), Positives = 26/39 (66%)
Frame = +1
Query: 631 IPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATP 669
IP E+ RFSHQ+PS+ + G+E +P ++LDPA P
Sbjct: 22 IPPNELYRFSHQIPSYKMTGDEREGVPRGSFELDPAGLP 138
>TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%)
Length = 624
Score = 42.4 bits (98), Expect = 3e-04
Identities = 21/47 (44%), Positives = 28/47 (58%)
Frame = +2
Query: 628 IRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDELLH 674
+R I +E+LRFSH++P + L EE NL W+LDP A P H
Sbjct: 89 LRKISTKELLRFSHRIPGFRLTEEEHGNLRG-FWELDPGALPVHYYH 226
>AV411199
Length = 423
Score = 41.6 bits (96), Expect = 4e-04
Identities = 29/96 (30%), Positives = 45/96 (46%), Gaps = 7/96 (7%)
Frame = +2
Query: 65 LDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNN--KCVGADGAFHLVSEAWARLS---- 118
+D+Y I+ ++ E +KK Y+++A HPD N A+ F L+SE++ LS
Sbjct: 65 VDYYEILEVDHHATDEELKKAYRRLAMKWHPDKNPDNKNDAETKFKLISESYEVLSDPQK 244
Query: 119 -GSYDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTF 153
YD +L G G+ S +GG TF
Sbjct: 245 RAIYDRYGEDRLKGGMPSPDAGMDSFFRTGGGPTTF 352
>CB828171
Length = 570
Score = 38.9 bits (89), Expect = 0.003
Identities = 15/39 (38%), Positives = 24/39 (61%)
Frame = +3
Query: 454 HDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVS 492
+DF K++S + QIWA+Y + D MP Y I+++ S
Sbjct: 294 YDFKKEKSREMLQCDQIWAIYGDRDNMPHYYAQIKKIES 410
>BP058085
Length = 288
Score = 38.9 bits (89), Expect = 0.003
Identities = 19/35 (54%), Positives = 24/35 (68%)
Frame = +3
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAK 35
ME KEEAL+A AEK+ +DF GA+ ALKA+
Sbjct: 180 MECNKEEALRAKVIAEKKMESKDFTGARKIALKAQ 284
>BP039305
Length = 555
Score = 38.5 bits (88), Expect = 0.004
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = +3
Query: 66 DWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGSYDMKR 125
D+Y+ +G+ S ++ +K Y+++A HPD K GA F +S A+ LS D KR
Sbjct: 381 DYYNTLGVPKSATVKEIKAAYRRLARQYHPDVYKEPGATDKFKEISNAYEVLSD--DKKR 554
>TC19393
Length = 603
Score = 35.8 bits (81), Expect = 0.023
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Frame = +1
Query: 152 TFWTICTSCKVQYEYLRKYVNKKLSCKNCRGIFIALET-APA 192
+FWT+C C +EY +KY + C NC F +E APA
Sbjct: 4 SFWTMCPYCWYLHEYEKKYEDCTFRCGNCLRTFHGVEVKAPA 129
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.132 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,204,151
Number of Sequences: 28460
Number of extensions: 181673
Number of successful extensions: 714
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 704
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 708
length of query: 682
length of database: 4,897,600
effective HSP length: 96
effective length of query: 586
effective length of database: 2,165,440
effective search space: 1268947840
effective search space used: 1268947840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140670.15


