

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.14 + phase: 0
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
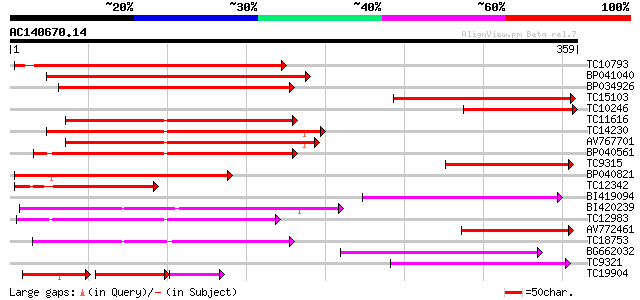
Score E
Sequences producing significant alignments: (bits) Value
TC10793 279 6e-76
BP041040 253 3e-68
BP034926 173 5e-44
TC15103 weakly similar to UP|Q9LH73 (Q9LH73) GDSL-motif lipase/h... 149 9e-37
TC10246 145 1e-35
TC11616 similar to UP|Q9SVU5 (Q9SVU5) Proline-rich APG-like prot... 140 3e-34
TC14230 140 3e-34
AV767701 139 7e-34
BP040561 136 6e-33
TC9315 130 3e-31
BP040821 127 3e-30
TC12342 122 9e-29
BI419094 118 1e-27
BI420239 118 2e-27
TC12983 115 1e-26
AV772461 106 7e-24
TC18753 106 7e-24
BG662032 103 4e-23
TC9321 similar to UP|O04320 (O04320) Proline-rich protein APG is... 96 1e-20
TC19904 weakly similar to UP|Q94CH8 (Q94CH8) Family II lipase EX... 68 9e-20
>TC10793
Length = 556
Score = 279 bits (713), Expect = 6e-76
Identities = 142/172 (82%), Positives = 150/172 (86%)
Frame = +1
Query: 4 GQRKNLLYTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQP 63
GQRK LL LC LL +L +V KVPAIIVFGDSSVDAGNNNFI TVARSNFQP
Sbjct: 55 GQRKKLL-----LCTTLLLQLLCLVVAGGKVPAIIVFGDSSVDAGNNNFIETVARSNFQP 219
Query: 64 YGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGY 123
YGRDF GGKPTGRFSNGRIATDFISEAFGIKPY+PAYLDPS+NIS FATGV+FASAATGY
Sbjct: 220 YGRDFQGGKPTGRFSNGRIATDFISEAFGIKPYVPAYLDPSYNISHFATGVAFASAATGY 399
Query: 124 DNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTN 175
DNATSDVLSVIPLWKQLEYYK YQKKL YLGEKKA +TITK+L+IISLGTN
Sbjct: 400 DNATSDVLSVIPLWKQLEYYKAYQKKLSTYLGEKKAHDTITKSLHIISLGTN 555
>BP041040
Length = 550
Score = 253 bits (646), Expect = 3e-68
Identities = 119/167 (71%), Positives = 143/167 (85%)
Frame = +3
Query: 24 LDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIA 83
L +S S KVPAIIVFGDSSVD+GNNNFI T+A+SNF+PYGRDF G PTGRFSNGRIA
Sbjct: 48 LHISTSSSGKVPAIIVFGDSSVDSGNNNFIPTMAKSNFEPYGRDFPDGNPTGRFSNGRIA 227
Query: 84 TDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYY 143
DFISEAFG+KP IPAYLDP+++IS FA+GV FASA TGYDN+TS+V VIPLWK++EYY
Sbjct: 228 PDFISEAFGLKPTIPAYLDPAYSISDFASGVCFASAGTGYDNSTSNVADVIPLWKEVEYY 407
Query: 144 KEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQ 190
K+Y++KL AYLG++KA E + +ALY++S+GTNDFLENYYT P R Q
Sbjct: 408 KDYRQKLVAYLGDEKANEIVKEALYLVSIGTNDFLENYYTFPERRCQ 548
>BP034926
Length = 607
Score = 173 bits (438), Expect = 5e-44
Identities = 80/149 (53%), Positives = 111/149 (73%)
Frame = +2
Query: 32 AKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAF 91
+ V I+VFGDSSVD GNNN + T +SNF PYG+DF PTGRF NGR+ATDFI+EA
Sbjct: 161 SNVSCILVFGDSSVDPGNNNVLRTSMKSNFPPYGKDFFNSLPTGRFCNGRLATDFIAEAL 340
Query: 92 GIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLG 151
G + +PA+LDP+ + GVSFASAATG+D+ T++V++V+P+ KQ++Y+ Y+ L
Sbjct: 341 GYRQMLPAFLDPNLKVEDLPYGVSFASAATGFDDYTANVVNVLPVSKQIQYFMHYKIHLR 520
Query: 152 AYLGEKKAKETITKALYIISLGTNDFLEN 180
LGE++A+ I AL+I+S+GTNDFL+N
Sbjct: 521 KLLGEERAEFIIRNALFIVSMGTNDFLQN 607
>TC15103 weakly similar to UP|Q9LH73 (Q9LH73) GDSL-motif
lipase/hydrolase-like protein, partial (9%)
Length = 544
Score = 149 bits (375), Expect = 9e-37
Identities = 64/115 (55%), Positives = 88/115 (75%)
Frame = +1
Query: 244 CVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMAC 303
C YN++ALEFNGKL L K+ K+LPG++LV +N YD+LL +V +P +GF+VA + C
Sbjct: 7 CSEEYNHVALEFNGKLGLLVKKMNKELPGLQLVDANAYDMLLQIVTQPSYFGFEVAGVGC 186
Query: 304 CATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQFL 358
C TG FEMGY C S F+C DA++YVFWD+FHP++KT+ IV+NYL++ LA+FL
Sbjct: 187 CGTGRFEMGYMCDPKSPFTCTDANKYVFWDAFHPSQKTSQIVSNYLIEKHLAKFL 351
>TC10246
Length = 525
Score = 145 bits (365), Expect = 1e-35
Identities = 67/72 (93%), Positives = 69/72 (95%)
Frame = +3
Query: 288 VKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVAN 347
VKKP QYGFQVASMACCATGMFEMGYACSRASLFSCMDAS+YVFWDSFH TEKT GI+AN
Sbjct: 18 VKKPAQYGFQVASMACCATGMFEMGYACSRASLFSCMDASKYVFWDSFHTTEKTYGIIAN 197
Query: 348 YLVKNALAQFLH 359
YLVKNALAQFLH
Sbjct: 198 YLVKNALAQFLH 233
>TC11616 similar to UP|Q9SVU5 (Q9SVU5) Proline-rich APG-like protein,
partial (41%)
Length = 569
Score = 140 bits (353), Expect = 3e-34
Identities = 70/148 (47%), Positives = 97/148 (65%), Gaps = 1/148 (0%)
Frame = +2
Query: 36 AIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKP 95
A VFGDS VD GNNN+++T AR++ PYG D G+PTGRFSNGR DFISE G +P
Sbjct: 128 AFFVFGDSLVDNGNNNYLATTARADSPPYGIDTPNGRPTGRFSNGRNIPDFISEQLGAEP 307
Query: 96 YIPAYLDPSFNISQFATGVSFASAATGYDNATS-DVLSVIPLWKQLEYYKEYQKKLGAYL 154
+P YL P N G +FASA G N T +++I +++QLEY+ EYQ+++ A +
Sbjct: 308 TLP-YLSPELNGDNLLVGANFASAGIGILNDTGIQFINIIRIFRQLEYFDEYQQRVSALI 484
Query: 155 GEKKAKETITKALYIISLGTNDFLENYY 182
G ++ + + AL +I+LG NDF+ NYY
Sbjct: 485 GPEETQRLVNGALVLITLGGNDFVNNYY 568
>TC14230
Length = 740
Score = 140 bits (353), Expect = 3e-34
Identities = 75/180 (41%), Positives = 112/180 (61%), Gaps = 3/180 (1%)
Frame = +1
Query: 24 LDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIA 83
L LS V + A VFGDS VD+GNN+F++T AR++ PYG DF +PTGRFSNG
Sbjct: 100 LFLSSVSAQHTRAFFVFGDSLVDSGNNDFLATTARADAPPYGIDFPTHRPTGRFSNGLNI 279
Query: 84 TDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATS-DVLSVIPLWKQLEY 142
D IS G++P +P YL P + G +FASA G N T L +I + KQL+
Sbjct: 280 PDLISLELGLEPTLP-YLSPLLVGEKLLIGANFASAGIGILNDTGIQFLHIIHIEKQLKL 456
Query: 143 YKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIP--GRASQYTPSEYQNFL 200
+++YQK++ A++G + + + +AL++I+LG NDF+ NYY +P R+ QY+ +Y ++
Sbjct: 457 FQQYQKRVSAHIGREGTRNLVNRALFLITLGGNDFVNNYYLVPYSARSRQYSLPDYVRYI 636
>AV767701
Length = 622
Score = 139 bits (350), Expect = 7e-34
Identities = 71/164 (43%), Positives = 106/164 (64%), Gaps = 3/164 (1%)
Frame = +3
Query: 36 AIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKP 95
A VFGDS VD+GNNN+++T AR++ PYG D+ +PTGRFSNG D IS+ G +
Sbjct: 132 AFFVFGDSLVDSGNNNYLATTARADSPPYGIDYPTRRPTGRFSNGLNIPDLISQQLGAES 311
Query: 96 YIPAYLDPSFNISQFATGVSFASAATGYDNAT-SDVLSVIPLWKQLEYYKEYQKKLGAYL 154
+P YL P ++ G +FASA G N T + L++I +++QL+Y++EYQ +L + +
Sbjct: 312 VLP-YLSPQLRGNKLLLGANFASAGIGILNDTGTQFLNIIRMYRQLDYFEEYQHRLASQI 488
Query: 155 GEKKAKETITKALYIISLGTNDFLENYYTIP--GRASQYTPSEY 196
G K K + KAL +I++G NDF+ NYY +P R+ QY+ +Y
Sbjct: 489 GVTKTKALVDKALVLITVGGNDFVNNYYLVPYSARSRQYSLPDY 620
>BP040561
Length = 515
Score = 136 bits (342), Expect = 6e-33
Identities = 74/168 (44%), Positives = 104/168 (61%), Gaps = 1/168 (0%)
Frame = +1
Query: 16 LCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTG 75
LC+++ V S + A VFGDS VD GNN+F++T AR++ PYG DF KPTG
Sbjct: 16 LCVIVTSLF--MAVDSTQTRAFFVFGDSLVDNGNNDFLATTARADAPPYGIDFPTHKPTG 189
Query: 76 RFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATS-DVLSVI 134
RFSNG D ISE G++P +P YL P + G +FASA G N T L++I
Sbjct: 190 RFSNGLNIPDLISEQLGLEPTMP-YLSPLLMGEKLLVGANFASAGIGILNDTGFQFLNII 366
Query: 135 PLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYY 182
++KQL+ ++ YQ++L A +G + K+ + KAL +I+LG NDF+ NYY
Sbjct: 367 HMYKQLKLFQHYQQRLSAQIGPEGTKKLVNKALVLITLGGNDFVNNYY 510
>TC9315
Length = 632
Score = 130 bits (328), Expect = 3e-31
Identities = 54/81 (66%), Positives = 70/81 (85%)
Frame = +3
Query: 277 FSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFH 336
FSNPY +++ +++KP YGF+ S+ACCATGMFEMGYACSR S+FSC DAS++VFWDSFH
Sbjct: 3 FSNPYYIMMQIIRKPDLYGFESTSLACCATGMFEMGYACSRGSMFSCTDASKFVFWDSFH 182
Query: 337 PTEKTNGIVANYLVKNALAQF 357
PTEKTNGIVA Y++++ L+ F
Sbjct: 183 PTEKTNGIVAKYVMQHVLSHF 245
>BP040821
Length = 478
Score = 127 bits (319), Expect = 3e-30
Identities = 66/140 (47%), Positives = 87/140 (62%), Gaps = 2/140 (1%)
Frame = +2
Query: 4 GQRKNLLYTNNILCILLLQRLD--LSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNF 61
G L + IL ++L R + L + +VPA++ FGDS VD GNNN I T+ + NF
Sbjct: 56 GVSPTTLILHFILLLVLTSRTKAVVKLPPNVEVPAVMAFGDSIVDPGNNNKIKTLVKCNF 235
Query: 62 QPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAAT 121
PYG+DF GG PTGRF NG+I +D ++E GIK +PAY P+ S TGVSFAS A+
Sbjct: 236 PPYGKDFEGGNPTGRFCNGKIPSDLLAEELGIKELLPAYKQPNLKPSDLLTGVSFASGAS 415
Query: 122 GYDNATSDVLSVIPLWKQLE 141
GYD T + SVI + QL+
Sbjct: 416 GYDPLTPKIASVISMSDQLD 475
>TC12342
Length = 541
Score = 122 bits (306), Expect = 9e-29
Identities = 69/92 (75%), Positives = 73/92 (79%), Gaps = 1/92 (1%)
Frame = +3
Query: 4 GQRKNLLYTNNILCILLLQRLDLSLVKS-AKVPAIIVFGDSSVDAGNNNFISTVARSNFQ 62
G RK LL ILCIL+LQ L K+ AKVPAIIVFGDS+VDAGNNNFI TVARSNFQ
Sbjct: 282 GHRKTLLCFP-ILCILVLQ-----LAKTNAKVPAIIVFGDSTVDAGNNNFIQTVARSNFQ 443
Query: 63 PYGRDFLGGKPTGRFSNGRIATDFISEAFGIK 94
PYGRDF GGK TGRF NGRI TDFI+E FGIK
Sbjct: 444 PYGRDFEGGKATGRFCNGRIPTDFIAEDFGIK 539
>BI419094
Length = 585
Score = 118 bits (296), Expect = 1e-27
Identities = 56/127 (44%), Positives = 76/127 (59%)
Frame = +3
Query: 224 LPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDV 283
L P+GCLP T N CV+ NN A+ FN KLN + L+K LPG++LV + Y
Sbjct: 9 LAPVGCLPAAITLFGHDSNQCVARLNNDAVNFNRKLNTTSQSLQKSLPGLKLVLLDIYQP 188
Query: 284 LLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNG 343
L +V KP + GF A ACC TG+ E C++ S+ +C +AS YVFWD FHP+E N
Sbjct: 189 LYDLVTKPSENGFAEARRACCGTGLLETSILCNQKSIGTCANASEYVFWDGFHPSEAANQ 368
Query: 344 IVANYLV 350
++A L+
Sbjct: 369 VLAGDLI 389
>BI420239
Length = 621
Score = 118 bits (295), Expect = 2e-27
Identities = 69/208 (33%), Positives = 118/208 (56%), Gaps = 3/208 (1%)
Frame = +1
Query: 7 KNLLYTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGR 66
+ LL ++ ++ + L ++++VP + +FGDS D+GNNN + T A+ ++ PYG
Sbjct: 1 RTLLVLALVILLVAIAIRQYCLQRNSQVPCLFIFGDSLSDSGNNNNLETDAKVDYLPYGI 180
Query: 67 DFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNA 126
DF G PTGR++NGR A D I+E G++ +IP + + S S GV++AS + G
Sbjct: 181 DFPTG-PTGRYTNGRNAIDKITELLGLEDFIPPFANLSG--SDILKGVNYASGSAGIRKE 351
Query: 127 TSDVLSV-IPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYY--T 183
+ L I + Q+ ++ ++ A LG KAK ++K LY +++GTND+ +NY+
Sbjct: 352 SGTNLGTNINMGLQIYFHMAIVSQISARLGFHKAKRHLSKCLYYVNIGTNDYEQNYFLPD 531
Query: 184 IPGRASQYTPSEYQNFLAGIAQNFIHKL 211
+ +S+YTP EY L +++ L
Sbjct: 532 LFDTSSKYTPEEYAKDLINRLSHYLQIL 615
>TC12983
Length = 515
Score = 115 bits (287), Expect = 1e-26
Identities = 62/168 (36%), Positives = 100/168 (58%), Gaps = 1/168 (0%)
Frame = +3
Query: 5 QRKNLLYTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPY 64
Q K L+ + + C+L++ ++ A+ A V GDS VD GNNN+++T AR++ PY
Sbjct: 18 QPKGCLFMSILACLLIITTWS-NVAPKAEARAFFVLGDSLVDNGNNNYLATTARADAYPY 194
Query: 65 GRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYD 124
G D + +GRFSNG D ISE G +P +P YL P + + G +FASA G
Sbjct: 195 GIDSASHRASGRFSNGLNMPDLISEKIGSEPTLP-YLSPELDGEKLLNGANFASAGIGIL 371
Query: 125 NATS-DVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIIS 171
N T +++I + QL Y+++YQ+++ A +GEK+A+ + +AL +I+
Sbjct: 372 NDTGVQFINIIRIGAQLGYFRQYQQRVSALIGEKEAQNLVNQALVLIT 515
>AV772461
Length = 442
Score = 106 bits (264), Expect = 7e-24
Identities = 44/71 (61%), Positives = 59/71 (82%)
Frame = -3
Query: 287 VVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVA 346
++ KP YGF+ S+ACCATGMFEMGYA +R S+FSC DAS++VFWDSFHP EKT G+VA
Sbjct: 440 IISKPDLYGFESTSLACCATGMFEMGYAFTRGSMFSCTDASQFVFWDSFHPPEKTYGLVA 261
Query: 347 NYLVKNALAQF 357
+Y++++ L+ F
Sbjct: 260 HYVMQHGLSHF 228
>TC18753
Length = 562
Score = 106 bits (264), Expect = 7e-24
Identities = 65/168 (38%), Positives = 91/168 (53%), Gaps = 2/168 (1%)
Frame = +3
Query: 15 ILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPT 74
+L + L + +VP FGDS VD GNNN +S++A++N++PYG DF GG PT
Sbjct: 66 VLIVCLWSTTTTGVGAEPQVPCFFSFGDSLVDNGNNNRLSSLAKANYRPYGIDFPGG-PT 242
Query: 75 GRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVL-SV 133
GRFSNG+ + D I E G YIP Y + GV++ASAA G T L
Sbjct: 243 GRFSNGKTSVDVIGELLGFGSYIPPY--ATTRGQDILRGVNYASAAAGIREETGQQLGGR 416
Query: 134 IPLWKQLEYYKEYQKKLGAYLG-EKKAKETITKALYIISLGTNDFLEN 180
I Q++ Y+ +L LG E A ++K +Y I +G+ND+L N
Sbjct: 417 ISFRGQVQNYQRTVSQLINLLGDENTAANYLSKCIYSIGIGSNDYLNN 560
>BG662032
Length = 421
Score = 103 bits (257), Expect = 4e-23
Identities = 46/128 (35%), Positives = 74/128 (56%)
Frame = +1
Query: 210 KLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKD 269
+LY L A+K +G + P+GC+P ++T N N+CV N +AL++N +L L +L +
Sbjct: 37 RLYRLDARKFVIGNVGPIGCIPYQKTINQLNENECVDLPNKLALQYNARLKDLMAELNDN 216
Query: 270 LPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRY 329
LPG V +N YD+++ ++K +YGF ++ ACC G G + C D ++
Sbjct: 217 LPGATFVVANVYDLVMELIKNYDKYGFTTSTKACCGNGGQFAGIVPCGPTSSICTDRYKH 396
Query: 330 VFWDSFHP 337
VFWD +HP
Sbjct: 397 VFWDPYHP 420
>TC9321 similar to UP|O04320 (O04320) Proline-rich protein APG isolog,
partial (37%)
Length = 541
Score = 95.5 bits (236), Expect = 1e-20
Identities = 45/115 (39%), Positives = 67/115 (58%), Gaps = 1/115 (0%)
Frame = +2
Query: 242 NDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASM 301
N CVS +N A FN K+N KL+K PG+++V + Y L +V+ P +GF A
Sbjct: 5 NGCVSRFNTDAQAFNKKINSAAAKLQKQHPGLKIVIFDIYKPLYDLVQTPSNFGFAEARR 184
Query: 302 ACCATGMFE-MGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALA 355
CC TG E C+ S+ +C +A++YVFWDS HP+E N ++A+ L+ +A
Sbjct: 185 GCCGTGTVETTSLLCNPKSIGTCSNATQYVFWDSVHPSEAANQVLADALILQGIA 349
>TC19904 weakly similar to UP|Q94CH8 (Q94CH8) Family II lipase EXL1, partial
(20%)
Length = 478
Score = 67.8 bits (164), Expect(3) = 9e-20
Identities = 29/47 (61%), Positives = 36/47 (75%)
Frame = -1
Query: 55 TVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYL 101
T AR NF PYG+DF GG PTGRFSNG++ +D I+E G+K +PAYL
Sbjct: 253 TSARCNFPPYGKDFKGGMPTGRFSNGKVPSDIIAEELGMKELLPAYL 113
Score = 32.7 bits (73), Expect(3) = 9e-20
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = -2
Query: 102 DPSFNISQFATGVSFASAATGYDNATSDVLSVIPL 136
DP + TGV FAS GYD TS+ + I L
Sbjct: 111 DPKLQPHELPTGVCFASGGAGYDPVTSETAAAISL 7
Score = 32.3 bits (72), Expect(3) = 9e-20
Identities = 18/46 (39%), Positives = 28/46 (60%), Gaps = 3/46 (6%)
Frame = -3
Query: 9 LLYTNNILCILLLQRLDLSLVK---SAKVPAIIVFGDSSVDAGNNN 51
L Y + L I ++ ++ VK + VPA++V+GDS VD GNN+
Sbjct: 404 LPYLSVFLLIFIVTLKTMA*VKLPPNISVPALLVYGDSIVDTGNND 267
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,721,069
Number of Sequences: 28460
Number of extensions: 73364
Number of successful extensions: 481
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 434
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 436
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC140670.14


