

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.8 - phase: 0
(122 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
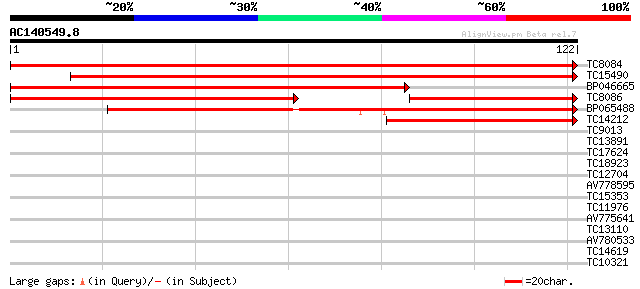
Sequences producing significant alignments: (bits) Value
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 148 3e-37
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 124 6e-30
BP046665 103 8e-24
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 77 9e-22
BP065488 82 2e-17
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 44 6e-06
TC9013 32 0.023
TC13891 homologue to UP|Q940Y6 (Q940Y6) AT4g27600/T29A15_90, par... 28 0.44
TC17624 27 1.3
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 26 1.7
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 26 1.7
AV778595 25 2.8
TC15353 weakly similar to UP|AAQ81937 (AAQ81937) SCC3, partial (7%) 25 3.7
TC11976 similar to GB|AAP12856.1|30017245|BT006207 At3g02340 {Ar... 25 4.8
AV775641 24 8.3
TC13110 similar to GB|CAC05445.1|9955561|ATT19L5 senescence-asso... 24 8.3
AV780533 24 8.3
TC14619 similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26... 24 8.3
TC10321 weakly similar to UP|Q86B51 (Q86B51) CG3525-PE, partial ... 24 8.3
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 148 bits (373), Expect = 3e-37
Identities = 70/122 (57%), Positives = 94/122 (76%)
Frame = +1
Query: 1 MVEEGGTYQLEGRVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKS 60
+V + GTY ++ VI+G+NI + +FIPRL + PSD+ PFKF+RR FPIS+ FAM INKS
Sbjct: 82 IVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCFAMTINKS 261
Query: 61 QGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFR 120
QGQSL HV +YL P+F+HGQLYVA+S+V SR GLK+L+ D+++ + NVVYREVF
Sbjct: 262 QGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFE 441
Query: 121 NV 122
N+
Sbjct: 442 NI 447
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 124 bits (310), Expect = 6e-30
Identities = 58/109 (53%), Positives = 82/109 (75%)
Frame = +2
Query: 14 VISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLP 73
VI+G++I + + IPR+ + PSD+ PFKF+RRQ PIS+ FAM INKSQG+SL HVG+YL
Sbjct: 149 VITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVGLYLS 328
Query: 74 SPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
P+ +HG LYVA+ +V SR LK+L+ D+++ + NVVYRE+F N+
Sbjct: 329 RPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFENI 475
>BP046665
Length = 524
Score = 103 bits (257), Expect = 8e-24
Identities = 49/86 (56%), Positives = 66/86 (75%)
Frame = -1
Query: 1 MVEEGGTYQLEGRVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKS 60
+V + GT ++ VI+ +NI + +FIPR+ + PSD+ PFKF+RRQFPIS+ AM INKS
Sbjct: 410 IVADLGTNVIKATVIT*TNIGDDIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKS 231
Query: 61 QGQSLEHVGVYLPSPIFSHGQLYVAI 86
QGQSL HVG+YL +F+HGQLYVA+
Sbjct: 230 QGQSLSHVGLYLSRHVFTHGQLYVAL 153
Score = 32.0 bits (71), Expect = 0.030
Identities = 15/45 (33%), Positives = 26/45 (57%)
Frame = -2
Query: 71 YLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVY 115
Y+ ++SH +S + SR GLK+L+ D+++ + NVVY
Sbjct: 202 YIFLAMYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 76.6 bits (187), Expect(2) = 9e-22
Identities = 36/62 (58%), Positives = 48/62 (77%)
Frame = +1
Query: 1 MVEEGGTYQLEGRVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKS 60
+V + GTY ++ VI+G+NI + +FIPRL + PSD+ PFKF+RR FPIS+ FAM INKS
Sbjct: 190 IVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCFAMTINKS 369
Query: 61 QG 62
QG
Sbjct: 370 QG 375
Score = 40.8 bits (94), Expect(2) = 9e-22
Identities = 18/36 (50%), Positives = 26/36 (72%)
Frame = +3
Query: 87 SQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
S+V SR GLK+L+ D+++ + NVVYREVF N+
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 476
>BP065488
Length = 439
Score = 82.4 bits (202), Expect = 2e-17
Identities = 48/104 (46%), Positives = 70/104 (67%), Gaps = 3/104 (2%)
Frame = -1
Query: 22 EKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPS--PIFSH 79
+ +FIPR+++ PS + P KF+R QFPIS+ FAM INKSQ S+ + V L S +SH
Sbjct: 376 DDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQ-XSVHYPTVALISFLACYSH 200
Query: 80 -GQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
+LYVA+S V SR GLK+L+ +++ + N VY+EVF+N+
Sbjct: 199 MDKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKNI 68
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 44.3 bits (103), Expect = 6e-06
Identities = 20/41 (48%), Positives = 28/41 (67%)
Frame = +2
Query: 82 LYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
LYVA+S+V S+ GLKILI+ D N+VY+EVF+ +
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVFQKI 124
>TC9013
Length = 766
Score = 32.3 bits (72), Expect = 0.023
Identities = 16/26 (61%), Positives = 18/26 (68%)
Frame = +3
Query: 97 ILINDDDDDDIDVASNVVYREVFRNV 122
ILI D DD + + SNVVY EVFR V
Sbjct: 627 ILICDGDDSNSNSTSNVVYTEVFRTV 704
>TC13891 homologue to UP|Q940Y6 (Q940Y6) AT4g27600/T29A15_90, partial (7%)
Length = 441
Score = 28.1 bits (61), Expect = 0.44
Identities = 18/48 (37%), Positives = 23/48 (47%), Gaps = 6/48 (12%)
Frame = +1
Query: 73 PSPIFSHGQLYVAISQVTSRG------GLKILINDDDDDDIDVASNVV 114
P HG L+ S +S G G + + DDDDDD +VA N V
Sbjct: 190 PGSASIHGGLFRVSSSFSSHGCAAFDGGSEGDLEDDDDDDDEVAENGV 333
>TC17624
Length = 615
Score = 26.6 bits (57), Expect = 1.3
Identities = 11/12 (91%), Positives = 12/12 (99%)
Frame = +1
Query: 111 SNVVYREVFRNV 122
SNVVY+EVFRNV
Sbjct: 538 SNVVYKEVFRNV 573
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 26.2 bits (56), Expect = 1.7
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Frame = -2
Query: 35 DNRIPFKFKR-RQFPISVSFAMIINKSQGQSL 65
D IPF+ R R+FP+SVSF + + G SL
Sbjct: 284 DQIIPFEPSRMRRFPLSVSFIVAVEF*TGASL 189
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 26.2 bits (56), Expect = 1.7
Identities = 15/32 (46%), Positives = 19/32 (58%), Gaps = 1/32 (3%)
Frame = -2
Query: 35 DNRIPFKFKR-RQFPISVSFAMIINKSQGQSL 65
D +PFK R R+FP SVSF + + G SL
Sbjct: 251 DQTMPFKPSRIRRFP*SVSFKVAVEFCTGASL 156
>AV778595
Length = 594
Score = 25.4 bits (54), Expect = 2.8
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 90 TSRGGLKILINDDDDDDIDVASNVV 114
+S +L DDDDDD D AS++V
Sbjct: 153 SSNSCCSLLPPDDDDDDDDDASSIV 79
Score = 23.9 bits (50), Expect = 8.3
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = -1
Query: 100 NDDDDDDIDVASNVVY 115
+DDDDDD D +S V++
Sbjct: 120 DDDDDDDDDASSIVLF 73
>TC15353 weakly similar to UP|AAQ81937 (AAQ81937) SCC3, partial (7%)
Length = 528
Score = 25.0 bits (53), Expect = 3.7
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = -1
Query: 26 IPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINK 59
+P TP+ NR PF + F + + + +INK
Sbjct: 117 LPFTGTTPTPNRNPFFLILKAFILCILVSQVINK 16
>TC11976 similar to GB|AAP12856.1|30017245|BT006207 At3g02340 {Arabidopsis
thaliana;}, partial (11%)
Length = 530
Score = 24.6 bits (52), Expect = 4.8
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 100 NDDDDDDIDVASNVVYREVFR 120
+DDDDDD D S + R++ R
Sbjct: 195 DDDDDDDDDGVSEISQRQLAR 257
>AV775641
Length = 393
Score = 23.9 bits (50), Expect = 8.3
Identities = 19/49 (38%), Positives = 25/49 (50%), Gaps = 4/49 (8%)
Frame = -3
Query: 48 PISVSFAMIINKSQGQSLEHVGVY-LPSPIF---SHGQLYVAISQVTSR 92
P SV F + S GQS H + L + + SHG L+V + VTSR
Sbjct: 298 PSSVFFNIWTGVSTGQSHMHSSLSELLNTLLTTHSHGSLFVGSTTVTSR 152
>TC13110 similar to GB|CAC05445.1|9955561|ATT19L5 senescence-associated
protein (SAG29) {Arabidopsis thaliana;} , partial (21%)
Length = 605
Score = 23.9 bits (50), Expect = 8.3
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -3
Query: 58 NKSQGQSLEHVGVYLPSPIFS 78
++ Q + H G LPSP+FS
Sbjct: 381 SRCSAQEVTHAGRSLPSPLFS 319
>AV780533
Length = 530
Score = 23.9 bits (50), Expect = 8.3
Identities = 10/31 (32%), Positives = 17/31 (54%)
Frame = -1
Query: 66 EHVGVYLPSPIFSHGQLYVAISQVTSRGGLK 96
EH +LP P+ + G+ + +SRG L+
Sbjct: 335 EHATTFLPGPMHTDGRQTLPPDS*SSRGSLR 243
>TC14619 similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (35%)
Length = 643
Score = 23.9 bits (50), Expect = 8.3
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +1
Query: 88 QVTSRGGLKILINDDDDDDID 108
+V S G+ LI+DDDD+D +
Sbjct: 265 EVRSDAGVIRLIDDDDDEDCE 327
>TC10321 weakly similar to UP|Q86B51 (Q86B51) CG3525-PE, partial (7%)
Length = 562
Score = 23.9 bits (50), Expect = 8.3
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +2
Query: 24 VFIPRLSLTPSDNRI 38
V++P LSL SDNR+
Sbjct: 458 VYLPALSLNKSDNRV 502
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.140 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,637,648
Number of Sequences: 28460
Number of extensions: 19118
Number of successful extensions: 183
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 170
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 180
length of query: 122
length of database: 4,897,600
effective HSP length: 79
effective length of query: 43
effective length of database: 2,649,260
effective search space: 113918180
effective search space used: 113918180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 49 (23.5 bits)
Medicago: description of AC140549.8


