

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
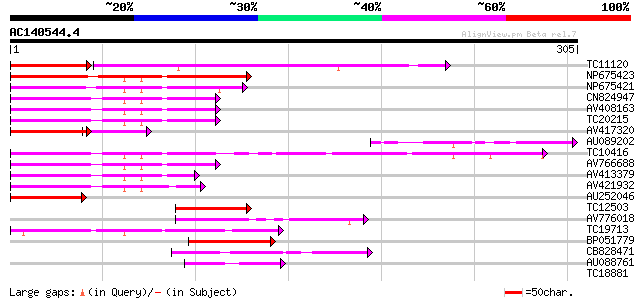
Query= AC140544.4 - phase: 0 /pseudo
(305 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 80 1e-33
NP675423 transcription factor MYB101 [Lotus japonicus] 124 2e-29
NP675421 transcription factor MYB103 [Lotus japonicus] 103 4e-23
CN824947 99 1e-21
AV408163 96 1e-20
TC20215 similar to AAS10104 (AAS10104) MYB transcription factor,... 91 2e-19
AV417320 82 1e-17
AU089202 85 1e-17
TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial... 82 1e-16
AV766688 81 2e-16
AV413379 69 8e-13
AV421932 66 8e-12
AU252046 65 1e-11
TC12503 homologue to UP|Q9ATD9 (Q9ATD9) BNLGHi233, partial (24%) 64 3e-11
AV776018 62 1e-10
TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%) 59 8e-10
BP051779 54 4e-08
CB828471 49 8e-07
AU088761 44 3e-05
TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose... 36 0.007
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 80.1 bits (196), Expect(2) = 1e-33
Identities = 59/205 (28%), Positives = 94/205 (45%), Gaps = 13/205 (6%)
Frame = +2
Query: 46 QDLRGVARAAD*DGLII*GRILRGESLVCKKNKPLFNFMHF*AT-----ATRLPKRTDNE 100
Q L V + * G I *G RG + + KK + + + AT LP RTDNE
Sbjct: 197 QVLTDVEKVVG*GGPIT*GLTSRGGNFLQKKEQTILHLHSILGNKWSTIATHLPGRTDNE 376
Query: 101 IKNYWNTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEA-RL 159
IKN+WNTHLKK+L +MG DP+TH+P+ + + + +AN++ + +A +L
Sbjct: 377 IKNFWNTHLKKKLIQMGFDPMTHQPRTDLVSTLPYLLALANMTQLMDHNHQSFDEQAVQL 556
Query: 160 VRESKLRSQLQHQFGS-------NTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHL 212
+ L LQ + + + + S N + N+KE ++ + T L
Sbjct: 557 AKLQYLNYLLQSSNNPLCMTNSYDQNALTNMEAFSLLNSISNVKENLGVDFSQMDTQTSL 736
Query: 213 LEFKDLMENSSMAFSSTLQHEMTMI 237
+ ++MA S L H M+
Sbjct: 737 FSH----DGNNMASSQPLHHPSNML 799
Score = 79.3 bits (194), Expect(2) = 1e-33
Identities = 32/44 (72%), Positives = 39/44 (87%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKA 44
MGRSPCC++ GLKKGPWT EEDQKL+ +I++HGHGSWR+LP A
Sbjct: 67 MGRSPCCDENGLKKGPWTPEEDQKLVEHIQKHGHGSWRALPKLA 198
>NP675423 transcription factor MYB101 [Lotus japonicus]
Length = 1047
Score = 124 bits (312), Expect = 2e-29
Identities = 69/140 (49%), Positives = 89/140 (63%), Gaps = 10/140 (7%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCC+++GLKKGPWT EED L+ YI++HGHGSWR+LP L G+ R L
Sbjct: 1 MGRTPCCDEIGLKKGPWTPEEDAILVDYIQKHGHGSWRALPK-----LAGLNRCGKSCRL 165
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E L+ + L N + A A LP RTDNEIKN+WNTHLK
Sbjct: 166 RWTNYLRPDIKRGKFTEEEEQLIINLHAVLGN--KWSAIAGHLPGRTDNEIKNFWNTHLK 339
Query: 111 KRLTKMGIDPVTHKPKNETL 130
K+L +MG+DPVTH+P+ + L
Sbjct: 340 KKLLQMGLDPVTHRPRLDHL 399
>NP675421 transcription factor MYB103 [Lotus japonicus]
Length = 924
Score = 103 bits (257), Expect = 4e-23
Identities = 62/140 (44%), Positives = 84/140 (59%), Gaps = 12/140 (8%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+P ++ G+KKGPW+ EED+ L+ YI++HGHGSWR+LP L G+ R L
Sbjct: 1 MMRTPSIDESGMKKGPWSQEEDKVLVDYIQKHGHGSWRALP-----QLAGLNRCGKSCRL 165
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E L+ + L N + A+ LP RTDNEIKN WNTHLK
Sbjct: 166 RWTNYLRPDIKRGKFSDEEEQLIINLHASLGN--KWATIASHLPGRTDNEIKNLWNTHLK 339
Query: 111 K--RLTKMGIDPVTHKPKNE 128
K +L +MG+DPVTH P+++
Sbjct: 340 KKLKLLQMGLDPVTHCPRSD 399
>CN824947
Length = 738
Score = 98.6 bits (244), Expect = 1e-21
Identities = 57/123 (46%), Positives = 73/123 (59%), Gaps = 10/123 (8%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCCEKVGLK+G WT+EED+ L YI+ G GSWRSLP A G+ R L
Sbjct: 151 MGRAPCCEKVGLKRGRWTAEEDELLTKYIQASGEGSWRSLPKNA-----GLLRCGKSCRL 315
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ G + RG ESL+ K + N + A+ +P RTDNEIKNYWN+HL
Sbjct: 316 RWINYLRGDLKRGNISDEEESLIVKLHASFGN--RWSLIASHMPGRTDNEIKNYWNSHLS 489
Query: 111 KRL 113
++L
Sbjct: 490 RKL 498
>AV408163
Length = 425
Score = 95.5 bits (236), Expect = 1e-20
Identities = 56/123 (45%), Positives = 70/123 (56%), Gaps = 10/123 (8%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK + KG WT EED +L+SYI HG G WRSLP A G+ R L
Sbjct: 77 MGRSPCCEKAHINKGAWTKEEDDRLISYIRAHGEGCWRSLPKAA-----GLLRCGKSCRL 241
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG + L+ K + L N + A RLP RTDNEIKNYWNTH++
Sbjct: 242 RWINYLRPDLKRGNFTEEEDELIIKLHSLLGN--KWSLIAGRLPGRTDNEIKNYWNTHIR 415
Query: 111 KRL 113
++L
Sbjct: 416 RKL 424
>TC20215 similar to AAS10104 (AAS10104) MYB transcription factor, partial
(35%)
Length = 464
Score = 91.3 bits (225), Expect = 2e-19
Identities = 52/123 (42%), Positives = 73/123 (59%), Gaps = 10/123 (8%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCCEKVGLK+G W+ EED+ L +I+++G GSW+SLP A G+ R L
Sbjct: 29 MGRAPCCEKVGLKRGRWSEEEDKILTDFIKQNGEGSWKSLPKNA-----GLLRCGKSCRL 193
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E ++ K + L N + A LP RTDNEIKNYWN+HL+
Sbjct: 194 RWINYLREDVKRGNITSEEEEIIVKLHAALGN--RWSVIAGHLPGRTDNEIKNYWNSHLR 367
Query: 111 KRL 113
+++
Sbjct: 368 RKI 376
>AV417320
Length = 369
Score = 81.6 bits (200), Expect(2) = 1e-17
Identities = 33/44 (75%), Positives = 40/44 (90%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKA 44
M R+PCCEK+GLKKGPWTSEEDQ L+SYI++HGHG+WR+LP A
Sbjct: 78 MVRAPCCEKIGLKKGPWTSEEDQILISYIQKHGHGNWRALPKHA 209
Score = 24.3 bits (51), Expect(2) = 1e-17
Identities = 16/37 (43%), Positives = 21/37 (56%)
Frame = +1
Query: 40 LPTKAVQDLRGVARAAD*DGLII*GRILRGESLVCKK 76
+P++++ V RAAD GL I*G I R E KK
Sbjct: 190 VPSQSMLGY*DVERAADYGGLTI*GLISREEISQLKK 300
>AU089202
Length = 401
Score = 85.1 bits (209), Expect = 1e-17
Identities = 58/115 (50%), Positives = 67/115 (57%), Gaps = 4/115 (3%)
Frame = +3
Query: 195 IKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMI---NVAEEGFTNLLLDN 251
+KEEGE +WKGY +S SM F+S M + +V EEGFTNLLL
Sbjct: 9 VKEEGE-QWKGYHES-------------SMTFTSGFHEFMEGVVGDDVVEEGFTNLLL-K 143
Query: 252 SNSGDLSLSPESGGECNTCDGSGSGGGSDFNE-DNKNYWNNILNLVNSSPSDSSM 305
+NS D E GGE N G G GSDF E DN NYW++ILNLVNSSPSDS M
Sbjct: 144 TNSDD----SEGGGESNN---GGRGSGSDFYEADNNNYWSSILNLVNSSPSDSPM 287
>TC10416 weakly similar to UP|O04108 (O04108) MYB factor, partial (41%)
Length = 1046
Score = 82.0 bits (201), Expect = 1e-16
Identities = 88/310 (28%), Positives = 133/310 (42%), Gaps = 21/310 (6%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+P C+K G++KG WT+EED+KL++Y+ +G +WR LP A G+AR L
Sbjct: 92 MARTPSCDKSGMRKGTWTAEEDKKLIAYVTRYGCWNWRQLPKFA-----GLARCGKSCRL 256
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ + +K L N + A LP RTD+E+KN+W+T LK
Sbjct: 257 RWLNYLRPNIKRGNFTQEEEEIIIRMHKKLGN--RWSTIAAELPGRTDSEVKNHWHTTLK 430
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQ 170
+ H+ T N V+N K + + A L+ + +
Sbjct: 431 R---------TVHEQNTVT----NEETKVSN--SKNERDPVPKSGIASLLVTPPPATSIS 565
Query: 171 HQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTL 230
GS ++ P +SS S ++ E +ED L+ D EN + F S L
Sbjct: 566 ---GSIGALSPFSTSSEFSCTTSDLTTSSSSEMLVFEDDFDFLD--DFTENVNENFWSDL 730
Query: 231 QHEMTMI----NVAE-EGFTNLLLDNSNSGDL----SLSPESGGECNTCDGSGSGGGSDF 281
+ + I N E + TN L N ++ + LSP N S G D
Sbjct: 731 EPYLANISNKPNETELQIETNCALQNQDNIQILDACELSPNQSSSENLILESDFGCFLDE 910
Query: 282 NEDN--KNYW 289
N D N+W
Sbjct: 911 NTDRTLDNFW 940
>AV766688
Length = 608
Score = 81.3 bits (199), Expect = 2e-16
Identities = 49/123 (39%), Positives = 71/123 (56%), Gaps = 10/123 (8%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+P C+K GL+KG WT+EED+KL++Y+ +G +WR LP A G+AR L
Sbjct: 77 MARTPSCDKSGLRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFA-----GLARCGKSCRL 241
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ + +K L N + A LP RTDNE+KN+W+T LK
Sbjct: 242 RWLNYLRPDIKRGHFTQEEEEIIIRMHKNLGN--RWSIIAAELPGRTDNEVKNHWHTSLK 415
Query: 111 KRL 113
+R+
Sbjct: 416 RRV 424
>AV413379
Length = 416
Score = 69.3 bits (168), Expect = 8e-13
Identities = 47/112 (41%), Positives = 57/112 (49%), Gaps = 10/112 (8%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK KG WT EED +L++YI HG G WRSLP A G+ R L
Sbjct: 102 MGRSPCCEKAHTNKGAWTKEEDDRLIAYIRAHGEGCWRSLPKSA-----GLLRCGKSCRL 266
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIK 102
+ + RG + L+ K + L N + A RL RTDNEIK
Sbjct: 267 RWINYLRPDLKRGNFTPQEDELIIKLHSLLGN--KWSLIAGRLAGRTDNEIK 416
>AV421932
Length = 402
Score = 65.9 bits (159), Expect = 8e-12
Identities = 44/116 (37%), Positives = 59/116 (49%), Gaps = 11/116 (9%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MG CC + +K+G W+ EED+KL+ YI HG+G W +P KA G+ R L
Sbjct: 78 MGHHSCCNQQKVKRGLWSPEEDEKLIRYITTHGYGCWSEVPEKA-----GLQRCGKSCRL 242
Query: 61 ----II*GRILRG------ESLVCKKNKPLFN-FMHF*ATATRLPKRTDNEIKNYW 105
+ I RG E L+ + + N + H A+ LP RTDNEIKNYW
Sbjct: 243 RWINYLRPDIRRGRFTPEEEKLIISLHGVVGNRWAHI---ASHLPGRTDNEIKNYW 401
>AU252046
Length = 276
Score = 65.5 bits (158), Expect = 1e-11
Identities = 25/41 (60%), Positives = 35/41 (84%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLP 41
MGR PCC+KVGLKKGPW++EED+KL+++I +G WR++P
Sbjct: 152 MGRQPCCDKVGLKKGPWSAEEDKKLINFILTNGQCCWRAVP 274
>TC12503 homologue to UP|Q9ATD9 (Q9ATD9) BNLGHi233, partial (24%)
Length = 957
Score = 63.9 bits (154), Expect = 3e-11
Identities = 28/41 (68%), Positives = 31/41 (75%)
Frame = +2
Query: 90 ATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNETL 130
A R+P RTDNEIKN+WNTHL K+L GIDP THKP N L
Sbjct: 167 AGRIPGRTDNEIKNFWNTHLSKKLISQGIDPRTHKPLNPEL 289
>AV776018
Length = 476
Score = 62.0 bits (149), Expect = 1e-10
Identities = 42/113 (37%), Positives = 61/113 (53%), Gaps = 9/113 (7%)
Frame = +1
Query: 90 ATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWE 149
AT+ P RTDNEIKN WN+ LKK+L + GIDPVTHKP +E EN ++ +K+Q
Sbjct: 82 ATQWPGRTDNEIKNLWNSCLKKKLRQRGIDPVTHKPLSEV---ENGKED----KNKSQ-- 234
Query: 150 SARLEAEARLVRESKLRSQLQHQFGSNTSVFP---------SQSSSSSSNQVL 193
++ E L++ +S +++S+ P S SS SS +L
Sbjct: 235 -EKVSNELNLLKSESFKSDAASYEQNSSSLAPKAYTTEMEGSSSSDCSSKDLL 390
>TC19713 homologue to UP|P93474 (P93474) Myb26, partial (55%)
Length = 532
Score = 59.3 bits (142), Expect = 8e-10
Identities = 43/155 (27%), Positives = 73/155 (46%), Gaps = 8/155 (5%)
Frame = +3
Query: 1 MGRSPC--CEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*D 58
M + PC ++KGPWT EED L++YI HG G W +L A L+ ++
Sbjct: 66 MDKKPCNSSHDPEVRKGPWTIEEDLILINYIANHGEGVWNTLAKSA--GLKRTGKSCRLR 239
Query: 59 GL------II*GRILRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKR 112
L + G I E L+ + + + A LP RTDNEIKN+W T ++K
Sbjct: 240 WLNYLRPDVRRGNITPEEQLLIMELHAKWG-NRWSKIAKHLPGRTDNEIKNFWRTRIQKH 416
Query: 113 LTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQ 147
+ + + ++ + + ++H + + +S A+
Sbjct: 417 IKQA---ESLQQQQSSSEINDHHQTSTSQMSTMAE 512
>BP051779
Length = 527
Score = 53.5 bits (127), Expect = 4e-08
Identities = 23/47 (48%), Positives = 31/47 (65%)
Frame = -2
Query: 97 TDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLS 143
TDNE+KNYWN+H++++L KMGIDP HK H +VA+ S
Sbjct: 526 TDNEVKNYWNSHIRRKLIKMGIDPNNHKLHQSLHRSHVHPLSVADAS 386
>CB828471
Length = 525
Score = 49.3 bits (116), Expect = 8e-07
Identities = 38/109 (34%), Positives = 50/109 (45%), Gaps = 1/109 (0%)
Frame = +1
Query: 88 ATATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQ 147
A A RLP RTDNEIKN W+THLKKRL P T + + L + S A H+
Sbjct: 151 AIAARLPGRTDNEIKNVWHTHLKKRL-----QPQTKQKQTNLDLDASKSNKDAKKEHQED 315
Query: 148 WESARLEAEARLVRESKLRSQLQHQFGSNTSVFPSQSSSSSS-NQVLNI 195
E R + S S +++S+ PS + S N V N+
Sbjct: 316 -EGPRSPQQC-----SSSNSDNMSSLTNSSSIAPSTNHDDMSINDVHNV 444
>AU088761
Length = 254
Score = 44.3 bits (103), Expect = 3e-05
Identities = 23/54 (42%), Positives = 30/54 (54%)
Frame = +1
Query: 95 KRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQW 148
K TDN+IKNYWNTHLKK+L K+ + +E L E S + + QW
Sbjct: 91 KETDNDIKNYWNTHLKKKLKKL-------QAGSEGNLGEGFSASSPQRISRGQW 231
>TC18881 similar to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucrose-responsive
element binding factor, partial (28%)
Length = 346
Score = 36.2 bits (82), Expect = 0.007
Identities = 31/101 (30%), Positives = 49/101 (47%), Gaps = 7/101 (6%)
Frame = +1
Query: 14 KGPWTSEEDQKLLSYIEEHGHGSW----RSLPTKAVQDLRGVARAAD*DGLII*GRILRG 69
KGPW+ EED+ L ++ HG +W +S+P ++ + R R + + R
Sbjct: 55 KGPWSPEEDEALTRLVQVHGPKNWTAISKSIPGRSGKSCR--LRWCNQLSPEVEHRPFTP 228
Query: 70 E--SLVCKKNKPLFNFMHF*ATATR-LPKRTDNEIKNYWNT 107
E S + + + N AT R L RTDN +KN+WN+
Sbjct: 229 EEDSTIIRAHARCGNKW---ATIARILNGRTDNAVKNHWNS 342
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.131 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,840,878
Number of Sequences: 28460
Number of extensions: 83432
Number of successful extensions: 694
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 670
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 685
length of query: 305
length of database: 4,897,600
effective HSP length: 90
effective length of query: 215
effective length of database: 2,336,200
effective search space: 502283000
effective search space used: 502283000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC140544.4


