

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
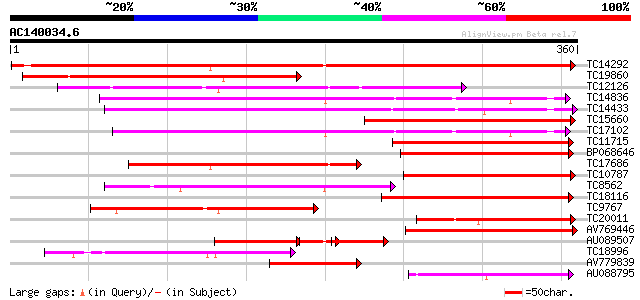
Query= AC140034.6 + phase: 0
(360 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precur... 322 5e-89
TC19860 weakly similar to UP|Q41064 (Q41064) Thiolprotease , par... 226 5e-60
TC12126 similar to PIR|S49166|S49166 cysteine proteinase precur... 223 3e-59
TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precur... 215 8e-57
TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete 212 7e-56
TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, part... 210 3e-55
TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-l... 205 8e-54
TC11715 weakly similar to UP|Q38886 (Q38886) SAG12 protein, part... 183 4e-47
BP068646 156 4e-39
TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Be... 150 4e-37
TC10787 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Be... 149 9e-37
TC8562 similar to UP|O24322 (O24322) Cysteine proteinase precurs... 143 4e-35
TC18116 similar to UP|AAP32193 (AAP32193) Cysteine protease 14, ... 142 7e-35
TC9767 similar to UP|Q7X750 (Q7X750) Cysteine proteinase , part... 132 9e-32
TC20011 similar to GB|CAA40073.1|1345573|PVEPC1 endopeptidase (E... 130 4e-31
AV769446 122 7e-29
AU089507 65 2e-27
TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precur... 111 2e-25
AV779839 96 1e-20
AU088795 87 4e-18
>TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (95%)
Length = 1792
Score = 322 bits (826), Expect = 5e-89
Identities = 168/368 (45%), Positives = 230/368 (61%), Gaps = 10/368 (2%)
Frame = +3
Query: 2 NACSHSHLHCKLAMMTSTILTTTIFILLMLCNTCVIASESECPPTHKQKSSDVEAMKKRF 61
++ S SHL M S + + L+ ++ + S +H S + +K +
Sbjct: 15 SSSSQSHL----PKMGSATMAIVLMFTLLAVSSAMDMSIISYDNSHMGNSRTDDEVKNMY 182
Query: 62 DGWVKRHGRKYKHNDEREVRFGIYQANVQYI-QCKNAQKN-SYNLTDNKFADLTNEEFQS 119
+ W+ +HG+ Y E+E RF I++ N+++I + NA N SY L N+FADLTNEE++S
Sbjct: 183 EEWLVKHGKVYNALGEKEKRFEIFKDNLKFIDEHNNADLNRSYKLGLNRFADLTNEEYRS 362
Query: 120 TYMGLST-------RLRSHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAA 172
Y G +LR+ + + LPES DWRKEGA+ + DQG CG CWAF+A
Sbjct: 363 KYFGTRVDPNRRMAKLRTKSDRYAPRVGDKLPESVDWRKEGALVGVKDQGSCGSCWAFSA 542
Query: 173 VAAVEGINKIKSGKLISLSEQELIDCDVKSGNQGCQGGLMETAYTFIIENGGLTTEQDYP 232
V AVE INKI +G L+SLS QEL+DCD +S N+GC GGLM+ A+ FII NGG+ +E+DYP
Sbjct: 543 VTAVESINKIVTGDLVSLSVQELVDCD-RSYNEGCNGGLMDYAFDFIINNGGIDSEEDYP 719
Query: 233 YEGVDGTCKMEKAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEG 292
Y+GVDG C + SI YE+VP +E LK A A+QP+SVAI+ GG FQ Y G
Sbjct: 720 YKGVDGRCDQYRKNAKVVSIDDYEDVPTYDEIALKKAVANQPISVAIEGGGREFQLYDSG 899
Query: 293 VFSGICGKQLNHGVTVVGYGKETINKYWIVKNSWGADWGESGYIRMKRDT-LSKEGMCGI 351
+F+G CG L+HGV VGYG E YWIV+NSWGA WGE GYIR++R+ ++ G CGI
Sbjct: 900 IFTGRCGTALDHGVVAVGYGTENGLDYWIVRNSWGASWGEGGYIRLERNLGNARSGKCGI 1079
Query: 352 AMQASYPL 359
A++ SYP+
Sbjct: 1080AIEPSYPI 1103
>TC19860 weakly similar to UP|Q41064 (Q41064) Thiolprotease , partial (16%)
Length = 571
Score = 226 bits (576), Expect = 5e-60
Identities = 112/180 (62%), Positives = 138/180 (76%), Gaps = 3/180 (1%)
Frame = +3
Query: 9 LHCKLAMMTSTILTTTIFILLMLCNTCVIASESECPPTHKQKSSDVEAMKKRFDGWVKRH 68
LH +LAMM +T LT T F LLM+ N +I S SECPP HK+ SS++EA +KRF+ W K+H
Sbjct: 30 LH*ELAMMIATKLTYTFFTLLMVSNLWII-SASECPPMHKKNSSNLEAKRKRFESWQKQH 206
Query: 69 GRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEFQSTYMGLSTRL 128
GRKY++ +E +VRF IYQ NV++I+C N+Q SY+LTDNKFADLTNEEF+ YMG
Sbjct: 207 GRKYENPEEWQVRFDIYQTNVEFIECINSQNRSYHLTDNKFADLTNEEFKRIYMGYGKTS 386
Query: 129 RSHNTG---FRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINKIKSG 185
S N G RY+ HGDLPES DWRK+GAVT+I DQG CG CWAF+AVAAVEGI++IKSG
Sbjct: 387 LSCNAGAGLLRYNGHGDLPESIDWRKKGAVTDIKDQGTCGSCWAFSAVAAVEGIHQIKSG 566
>TC12126 similar to PIR|S49166|S49166 cysteine proteinase precursor -
spring vetch {Vicia sativa;} , partial (71%)
Length = 839
Score = 223 bits (569), Expect = 3e-59
Identities = 124/270 (45%), Positives = 164/270 (59%), Gaps = 10/270 (3%)
Frame = +1
Query: 31 LCNTCVIASESECPPTHKQKSSDVEAMKKRFDGWVKRHGRKYKHNDEREVRFGIYQANVQ 90
L + + S +E H++ + E + ++ W + H + DE+ RF +++ANV
Sbjct: 43 LFSLALFLSVAESFDYHEKDLASEEGLWDLYERW-RSHHTVSRSLDEKHKRFNVFKANVM 219
Query: 91 YIQCKNAQKNSYNLTDNKFADLTNEEFQSTYMGLSTRLRSH---------NTGFRYDEHG 141
++ N Y L NKFADLTN EF+S Y S+++ H N F Y+
Sbjct: 220 HVHETNKLDKPYKLKLNKFADLTNYEFKSIYA--SSKIDHHRMFHGIPRGNGSFMYENVD 393
Query: 142 DLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINKIKSGKLISLSEQELIDCDVK 201
+P SKDWR GAVT + DQGQCG CWAF+A+AAVEGIN+I++ L+SLSEQEL+DCDV+
Sbjct: 394 SVPISKDWRTSGAVTPVKDQGQCGSCWAFSAIAAVEGINQIRTHHLVSLSEQELVDCDVE 573
Query: 202 SGNQGCQGGLMETAYTFIIENGGLTTEQDYPYEGVDGTCKMEKAAHY-AASISGYEEVPA 260
N+GC GG M A FI E G+T+E YPY DGTC +K + A SI GYE VP
Sbjct: 574 V-NEGCNGGFMAKALEFIKEK-GITSESIYPYTATDGTCNTQKMVNEPAVSIDGYEIVPE 747
Query: 261 DNEAKLKAAAAHQPVSVAIDAGGYSFQFYS 290
+NE L AAA+QP+SV IDAGG FQFYS
Sbjct: 748 NNEVALLKAAANQPISVGIDAGGSDFQFYS 837
>TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (92%)
Length = 1416
Score = 215 bits (548), Expect = 8e-57
Identities = 121/315 (38%), Positives = 179/315 (56%), Gaps = 16/315 (5%)
Frame = +2
Query: 58 KKRFDGWVKRHGRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEF 117
++ F + + G+ Y +E + RFG++++N++ + S KF+DLT EF
Sbjct: 176 ERHFTTFKAKFGKSYATREEHDYRFGVFKSNLRRAKVHAKLDPSAEHGVTKFSDLTPAEF 355
Query: 118 QSTYMGLST-RLRSHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAV 176
+ ++G+ RL + DLP+ DWR +GAVT + DQG CG CWAF+A A+
Sbjct: 356 RRQFLGVKALRLPADAQKAPILPTRDLPDDFDWRDKGAVTPVKDQGACGSCWAFSATGAL 535
Query: 177 EGINKIKSGKLISLSEQELIDCD-------VKSGNQGCQGGLMETAYTFIIENGGLTTEQ 229
EG + + +GKL+S+SEQ+L+DCD S + GC GGLM A+ +I+++GG+ E+
Sbjct: 536 EGSHYLTTGKLVSISEQQLVDCDHVCDPEEYNSCDSGCNGGLMNNAFEYILQSGGVMREK 715
Query: 230 DYPYEGVDGTCKMEKAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFY 289
DYPY G DGTCK +K + AAS++ + D + + P++VAI+A Q Y
Sbjct: 716 DYPYTGRDGTCKFDK-SKIAASVANFSVGSLDEDQIAANLVMNGPLAVAINA--VYMQTY 886
Query: 290 SEGVFSG-ICGKQLNHGVTVVGYGKETI------NK-YWIVKNSWGADWGESGYIRMKRD 341
GV ICGK L+HGV +VGYGK NK YWI+KNSWG WGE+GY ++ R
Sbjct: 887 VGGVSCPYICGKNLDHGVLLVGYGKAGYAPIRFKNKAYWIIKNSWGESWGENGYYKICRG 1066
Query: 342 TLSKEGMCGIAMQAS 356
+CG+ S
Sbjct: 1067----RNVCGVDSMVS 1099
>TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete
Length = 1392
Score = 212 bits (540), Expect = 7e-56
Identities = 120/305 (39%), Positives = 167/305 (54%), Gaps = 5/305 (1%)
Frame = +1
Query: 61 FDGWVKRHGRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEFQST 120
F + R+G++Y +E + RF I+ +++ I+ N ++ SY L N FADL+ +EF++
Sbjct: 232 FARFATRYGKRYDSVEEIQHRFRIFSESLELIKSTNKKRLSYKLGLNHFADLSWDEFRTQ 411
Query: 121 YMGLSTRLRSHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGIN 180
+G + + G LP KDWRKE V+E+ DQ CG CW F+ A+E
Sbjct: 412 KLGAAQNCSATLIGNHKLTDAVLPAEKDWRKESIVSEVKDQAHCGSCWTFSTTGALEAAY 591
Query: 181 KIKSGKLISLSEQELIDCDVKSGNQGCQGGLMETAYTFIIENGGLTTEQDYPYEGVDGTC 240
GK ISLSEQ+L+DC N GC GGL A+ +I NGG+ E++YPY D C
Sbjct: 592 AQAHGKNISLSEQQLVDCAGAFNNFGCNGGLPSQAFEYIKYNGGIALEKEYPYTAKDEAC 771
Query: 241 KMEKAAHYAASISGYEEVPADNEAKLKAAAAH-QPVSVAIDAGGYSFQFYSEGVF-SGIC 298
K A + A + + E +LK A A +PVSVA F+ Y EGV+ S C
Sbjct: 772 KF-TAENVAVRVLDSVNITLGAEDELKHAVAFARPVSVAFQVVD-GFRLYKEGVYTSDTC 945
Query: 299 GK---QLNHGVTVVGYGKETINKYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQA 355
G +NH V VGYG E YWI+KNSWG+ WG+ GY +M+ + MCG+A A
Sbjct: 946 GNTPMDVNHAVLAVGYGVENNVPYWIIKNSWGSTWGDHGYFKMELG----KNMCGVATCA 1113
Query: 356 SYPLV 360
SYP+V
Sbjct: 1114SYPIV 1128
>TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, partial (34%)
Length = 618
Score = 210 bits (535), Expect = 3e-55
Identities = 97/135 (71%), Positives = 114/135 (83%), Gaps = 1/135 (0%)
Frame = +1
Query: 226 TTEQDYPYEGVDGTCKMEKAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYS 285
TTE+DYPY+G DGTC EKAAH+A +ISG+++VPA NEA LKAAAAHQPVSV IDAGG+
Sbjct: 1 TTERDYPYKGKDGTCNKEKAAHHAVTISGHKKVPASNEAMLKAAAAHQPVSVGIDAGGFL 180
Query: 286 FQFYSEGVFSGICGKQLNHGVTVVGYGKETIN-KYWIVKNSWGADWGESGYIRMKRDTLS 344
FQ YS GVFSG CGK+LNH VTVVGYG+E KYW+VKNSWGADWGE+GY+++ RDT
Sbjct: 181 FQLYSGGVFSGFCGKELNHAVTVVGYGREVNGPKYWLVKNSWGADWGEAGYMKILRDTQD 360
Query: 345 KEGMCGIAMQASYPL 359
K+G CGIAM ASYP+
Sbjct: 361 KDGTCGIAMVASYPV 405
>TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-like protein
1, complete
Length = 1086
Score = 205 bits (522), Expect = 8e-54
Identities = 117/309 (37%), Positives = 173/309 (55%), Gaps = 18/309 (5%)
Frame = +1
Query: 66 KRHGRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEFQSTYMGL- 124
+R G+ Y +E RF ++++N+ + S +F+DLT EFQ + +GL
Sbjct: 148 RRFGKVYATEEEHGYRFNVFKSNMHRARRHQLLDPSAVHGVTQFSDLTPMEFQHSVLGLR 327
Query: 125 STRLRSHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINKIKS 184
L S +LP+ DWR+ GAVT + +QG CG CW+F+A A+EG + + +
Sbjct: 328 GVGLPSDADSAPILPTDNLPKDFDWREHGAVTPVKNQGSCGSCWSFSATGALEGAHFLST 507
Query: 185 GKLISLSEQELIDCD--------VKSGNQGCQGGLMETAYTFIIENGGLTTEQDYPYEGV 236
G+L+SLSEQ+L+DCD S + GC GGLM +A+ +I+ NGG+ E+DYPY G
Sbjct: 508 GELVSLSEQQLVDCDHQQCDPEEAGSCDSGCNGGLMNSAFEYILNNGGVMREEDYPYSGT 687
Query: 237 D-GTCKMEKAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFS 295
+ GTCK +K A AAS++ + V D + + P++VAI+A Q Y GV
Sbjct: 688 NGGTCKFDK-AKIAASVANFSVVSRDEDQIAANLVKNGPLAVAINA--VYMQTYVGGVSC 858
Query: 296 G-ICGKQLNHGVTVVGYGKETI-------NKYWIVKNSWGADWGESGYIRMKRDTLSKEG 347
+C K+LNHGV +VGYG E+ YWI+KNSWG +WGE+GY ++ R
Sbjct: 859 PYVCSKKLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRG----RN 1026
Query: 348 MCGIAMQAS 356
+CG+ S
Sbjct: 1027ICGVDSMVS 1053
>TC11715 weakly similar to UP|Q38886 (Q38886) SAG12 protein, partial (32%)
Length = 566
Score = 183 bits (464), Expect = 4e-47
Identities = 86/116 (74%), Positives = 99/116 (85%), Gaps = 1/116 (0%)
Frame = +1
Query: 244 KAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSGICGKQLN 303
KAAH+A SISG+E+VPA NEA LKAAAA+QPVSVAIDAGG+ FQ YS GVF G CGKQLN
Sbjct: 1 KAAHHAVSISGHEKVPASNEAMLKAAAANQPVSVAIDAGGFLFQLYSGGVFFGFCGKQLN 180
Query: 304 HGVTVVGYGKETIN-KYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYP 358
H VT+VGYG++ KYW+VKNSWGADWGESGY+++ RDTL K+G CGIAM ASYP
Sbjct: 181 HAVTIVGYGRQVNGPKYWLVKNSWGADWGESGYMKILRDTLDKDGTCGIAMDASYP 348
>BP068646
Length = 491
Score = 156 bits (395), Expect = 4e-39
Identities = 70/110 (63%), Positives = 87/110 (78%)
Frame = -3
Query: 249 AASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSGICGKQLNHGVTV 308
AAS+ G+E+VPA +E+ L A A+QP+SVAIDA G FQ YS GVF+G CG +L+HGVT
Sbjct: 483 AASLKGFEDVPAHSESALLKAVANQPISVAIDASGSEFQCYSSGVFTGSCGTELDHGVTA 304
Query: 309 VGYGKETINKYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYP 358
VGYG + +YW+VKNSWG WGE GYIRM+RD ++EG+CGIAMQASYP
Sbjct: 303 VGYGSDGGTQYWLVKNSWGEQWGEQGYIRMQRDVAAEEGLCGIAMQASYP 154
>TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Bean
endopeptidase) (Cysteine proteinase)
(Sulfhydryl-endopeptidase) (SH-EP) , partial (54%)
Length = 624
Score = 150 bits (378), Expect = 4e-37
Identities = 78/155 (50%), Positives = 98/155 (62%), Gaps = 7/155 (4%)
Frame = +3
Query: 76 DEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEFQSTYMGLST------RLR 129
DE+ RF +++ANV ++ N Y L NKFAD+TN EF+S Y R
Sbjct: 159 DEKRKRFNVFKANVMHVHDTNKLDMPYKLELNKFADMTNHEFRSIYASSKIDHHKMLRGT 338
Query: 130 SHNTG-FRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINKIKSGKLI 188
H G F Y+ +P S DWRK+GAVT I DQ C CWAF+AVAAVEGIN+IK+ KL+
Sbjct: 339 PHGKGTFMYENVDSVPPSIDWRKKGAVTAIKDQRDCSSCWAFSAVAAVEGINQIKTQKLV 518
Query: 189 SLSEQELIDCDVKSGNQGCQGGLMETAYTFIIENG 223
SLSEQEL+DCD++ N GC GG M A+ FI +NG
Sbjct: 519 SLSEQELVDCDIEH-NNGCNGGYMNNAFDFIKQNG 620
>TC10787 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Bean
endopeptidase) (Cysteine proteinase)
(Sulfhydryl-endopeptidase) (SH-EP) , partial (34%)
Length = 502
Score = 149 bits (375), Expect = 9e-37
Identities = 72/110 (65%), Positives = 84/110 (75%), Gaps = 1/110 (0%)
Frame = +1
Query: 251 SISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSGICGKQLNHGVTVVG 310
SI G+E VPA++EA L A A+ PVSVAIDAGG F FYSEGVF+G CG L+HGV VVG
Sbjct: 1 SIDGHETVPANDEAALLNAVANPPVSVAIDAGGSDFPFYSEGVFTGDCGTGLDHGVAVVG 180
Query: 311 YGKETI-NKYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYPL 359
YG YWIVKNSWG +WGE GYIRM+R+ KEG+CGIAM+ASYP+
Sbjct: 181 YGTTVDGTDYWIVKNSWGPEWGEQGYIRMQRNISDKEGLCGIAMEASYPI 330
>TC8562 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (57%)
Length = 776
Score = 143 bits (361), Expect = 4e-35
Identities = 78/196 (39%), Positives = 115/196 (57%), Gaps = 11/196 (5%)
Frame = +2
Query: 61 FDGWVKRHGRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDN--KFADLTNEEFQ 118
F + R G+ Y +E RFG+++ N+ I K+ QK + +F+DLT EF+
Sbjct: 191 FSTFKARFGKTYATEEEHNYRFGVFKKNL--ILAKSHQKLDPSAVHGVTRFSDLTPSEFR 364
Query: 119 STYMGLST-RLRSHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVE 177
++GL R DLP DWR++GAVTE+ +QG CG CW+F+A A+E
Sbjct: 365 RQFLGLKAPRHLKDAQKAPILPTNDLPTDFDWREKGAVTEVKNQGSCGSCWSFSATGALE 544
Query: 178 GINKIKSGKLISLSEQELIDC-------DVKSGNQGCQGGLMETAYTFIIENGGLTTEQD 230
G + + +G+L+SLSEQ+L+DC D +S + GC GGLM A+ + ++ GG+ E+D
Sbjct: 545 GAHYLATGELVSLSEQQLVDCDHECDPEDSRSCDSGCNGGLMTNAFEYTLKAGGVMREKD 724
Query: 231 YPYEGVD-GTCKMEKA 245
YPY G D GTCK EK+
Sbjct: 725 YPYTGRDRGTCKFEKS 772
>TC18116 similar to UP|AAP32193 (AAP32193) Cysteine protease 14, partial
(37%)
Length = 555
Score = 142 bits (359), Expect = 7e-35
Identities = 68/122 (55%), Positives = 80/122 (64%)
Frame = +3
Query: 237 DGTCKMEKAAHYAASISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSG 296
+GTC+M K +ISGY +VP +NE L A A+QP+SVAI+A G FQFYS GVF G
Sbjct: 27 EGTCEMSKEESQVVTISGYHDVPQNNEQSLLKALANQPLSVAIEASGRDFQFYSGGVFDG 206
Query: 297 ICGKQLNHGVTVVGYGKETINKYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQAS 356
CG QL+HGV VGYG Y VKNSWG WGE GYIR +R+ EGMCG+ AS
Sbjct: 207 HCGTQLDHGVAAVGYGTSKGLDYITVKNSWGTKWGEKGYIRFRRNNGKPEGMCGLYKMAS 386
Query: 357 YP 358
YP
Sbjct: 387 YP 392
>TC9767 similar to UP|Q7X750 (Q7X750) Cysteine proteinase , partial (50%)
Length = 564
Score = 132 bits (332), Expect = 9e-32
Identities = 69/156 (44%), Positives = 97/156 (61%), Gaps = 11/156 (7%)
Frame = +3
Query: 52 SDVEAMKKRFDGWVK--RHGRKYKHNDEREVRFGIYQANVQYIQCKNAQKNSYNLTDNKF 109
SDV + K +D + + H + D++ RF +++ANV ++ N Y L NKF
Sbjct: 102 SDVGSEKSLWDLYERWRSHHTVSRSLDDKHKRFNVFKANVMHVHNTNKMDKPYKLKLNKF 281
Query: 110 ADLTNEEFQSTYMGLSTRLRSH---------NTGFRYDEHGDLPESKDWRKEGAVTEIMD 160
AD+TN EF+S Y G +++ H N F Y++ +P S DWRK+GAVT++ D
Sbjct: 282 ADMTNHEFRSAYAG--SKVNHHRMFRGATRGNGSFMYEKINRVPPSVDWRKKGAVTDVKD 455
Query: 161 QGQCGGCWAFAAVAAVEGINKIKSGKLISLSEQELI 196
QGQCG CWAF+ V AVEGIN+IK+ KL+SLSEQEL+
Sbjct: 456 QGQCGSCWAFSTVVAVEGINQIKTNKLVSLSEQELV 563
>TC20011 similar to GB|CAA40073.1|1345573|PVEPC1 endopeptidase (EP-C1)
{Phaseolus vulgaris;} , partial (29%)
Length = 472
Score = 130 bits (326), Expect = 4e-31
Identities = 66/105 (62%), Positives = 74/105 (69%), Gaps = 4/105 (3%)
Frame = +1
Query: 259 PADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSG---ICGKQLNHGVTVVGYGKET 315
P +NE L AAA+QPVSVA++AG Q YSEGVF G CG QLNHGV +VGYG
Sbjct: 1 PENNETALLLAAANQPVSVAVEAG-VDMQLYSEGVFMGGGVDCGTQLNHGVAIVGYGATQ 177
Query: 316 IN-KYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYPL 359
+YWIVKNSWGA WGE GYIRMKRD K G+CGIAM SYP+
Sbjct: 178 YGTQYWIVKNSWGAKWGEEGYIRMKRDISDKRGLCGIAMAPSYPI 312
>AV769446
Length = 489
Score = 122 bits (307), Expect = 7e-29
Identities = 60/110 (54%), Positives = 72/110 (64%), Gaps = 1/110 (0%)
Frame = -1
Query: 252 ISGYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSGICGKQLNHGVTVVGY 311
I GY+ V ++NE L A A QPV V IDA G F+ YS G+F G CG LNH +T VG+
Sbjct: 483 IHGYKWVSSNNEEALIQAVAKQPVIVGIDAAGDDFKHYSGGIFRGQCGTYLNHAITFVGF 304
Query: 312 GKETIN-KYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYPLV 360
G + KYWI KNSWG DWGE GYIR+ R S+EG+CGIAM YP +
Sbjct: 303 GIDKDGLKYWIGKNSWGTDWGEGGYIRLPRGGYSEEGICGIAMSPLYPKI 154
>AU089507
Length = 379
Score = 65.1 bits (157), Expect(3) = 2e-27
Identities = 27/55 (49%), Positives = 34/55 (61%)
Frame = +3
Query: 131 HNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINKIKSG 185
H + Y+ LP DWR +GAV I DQG CG CWAF+ +A VE +NKI +G
Sbjct: 3 HEHRYAYNARDRLPAHVDWRLQGAVAPIKDQGSCGSCWAFSTIATVEAVNKIVTG 167
Score = 62.4 bits (150), Expect(3) = 2e-27
Identities = 25/36 (69%), Positives = 31/36 (85%)
Frame = +2
Query: 205 QGCQGGLMETAYTFIIENGGLTTEQDYPYEGVDGTC 240
+GC GGLM+ A+ FII NGG+ TE+DYPY+GVDGTC
Sbjct: 224 KGCNGGLMDYAFEFIIGNGGIDTEEDYPYKGVDGTC 331
Score = 31.6 bits (70), Expect(3) = 2e-27
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = +1
Query: 185 GKLISLSEQELIDCDVKSGNQGCQGG 210
GKL+ LSEQEL+DCD ++ N+ Q G
Sbjct: 166 GKLVPLSEQELVDCD-RAYNERLQWG 240
>TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (29%)
Length = 549
Score = 111 bits (278), Expect = 2e-25
Identities = 64/177 (36%), Positives = 100/177 (56%), Gaps = 18/177 (10%)
Frame = +2
Query: 23 TTIFILLMLCNTCVIAS------ESECPPTHKQKSSDVEAMKKRFDGWVKRHGRKYKHND 76
TT+ ++++ ++ + S + PP SD E M+ ++ W + G+ Y
Sbjct: 32 TTVLVVVITVSSAMDMSIISYSRDKSAPPLR----SDEEVMRM-YEEWAMKQGKVYNALG 196
Query: 77 EREVRFGIYQANVQYIQCKNAQKNSYNLTDNKFADLTNEEFQSTYMGL---STRLR---- 129
E+E RF I++ N+++I NA+ +Y + N+FADL+NEE+++ ++G+ S R R
Sbjct: 197 EKEKRFEIFKDNLKFIDEHNAENRTYKVGLNRFADLSNEEYRAKFLGIRVDSNRRRRRMA 376
Query: 130 -----SHNTGFRYDEHGDLPESKDWRKEGAVTEIMDQGQCGGCWAFAAVAAVEGINK 181
+H R + LPES DWRKEGAV + DQG+CG WA +AVAA EGINK
Sbjct: 377 KSTTSNHRYAPRVGDGEKLPESVDWRKEGAVVGVKDQGECGSAWALSAVAAGEGINK 547
>AV779839
Length = 498
Score = 95.5 bits (236), Expect = 1e-20
Identities = 42/58 (72%), Positives = 54/58 (92%)
Frame = +3
Query: 166 GCWAFAAVAAVEGINKIKSGKLISLSEQELIDCDVKSGNQGCQGGLMETAYTFIIENG 223
G WAF+AVAAVEG NKIK+GKL+SLSEQ+LIDCD+++GNQGC+GG META+T+I ++G
Sbjct: 321 GGWAFSAVAAVEGTNKIKTGKLVSLSEQQLIDCDIRNGNQGCEGGEMETAFTYISKHG 494
>AU088795
Length = 482
Score = 87.0 bits (214), Expect = 4e-18
Identities = 43/108 (39%), Positives = 57/108 (51%), Gaps = 3/108 (2%)
Frame = +2
Query: 254 GYEEVPADNEAKLKAAAAHQPVSVAIDAGGYSFQFYSEGVFSGICGKQ---LNHGVTVVG 310
GY +V A ++ L A QP+SV ID + FQ Y+ G++ G C ++H V VG
Sbjct: 5 GYTDV-AQTDSALLCATVKQPISVGIDGSSWDFQLYTGGIYDGDCSXNPDDIDHAVLXVG 181
Query: 311 YGKETINKYWIVKNSWGADWGESGYIRMKRDTLSKEGMCGIAMQASYP 358
YG YWI NSWG WG YI + +T + G+C I ASYP
Sbjct: 182 YGSXGDEXYWIXNNSWGTXWGMEXYIYITXNTDLQYGVCAINAMASYP 325
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.132 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,511,003
Number of Sequences: 28460
Number of extensions: 89060
Number of successful extensions: 541
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 502
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 503
length of query: 360
length of database: 4,897,600
effective HSP length: 91
effective length of query: 269
effective length of database: 2,307,740
effective search space: 620782060
effective search space used: 620782060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC140034.6


