

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
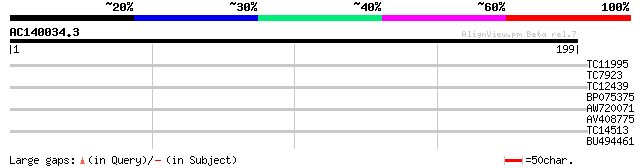
Query= AC140034.3 + phase: 0
(199 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11995 29 0.48
TC7923 similar to UP|PDI_MEDSA (P29828) Protein disulfide isomer... 28 1.4
TC12439 similar to UP|Q9FMJ2 (Q9FMJ2) Ankyrin-like protein, part... 27 3.1
BP075375 27 3.1
AW720071 25 6.9
AV408775 25 6.9
TC14513 homologue to UP|Q7M1K9 (Q7M1K9) Chlorophyll a/b-binding ... 25 9.0
BU494461 25 9.0
>TC11995
Length = 726
Score = 29.3 bits (64), Expect = 0.48
Identities = 15/45 (33%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Frame = +1
Query: 70 RDFYTSYLGRANVLAGTEDPEELEELARVRTYCVR--WYLMYLVG 112
R F T+YLGR V+ T + ++ EE+ + + +R ++L+Y G
Sbjct: 298 RSFQTNYLGRLTVIHNTLEGKKKEEILFCQVFVLRVFYFLVYRNG 432
>TC7923 similar to UP|PDI_MEDSA (P29828) Protein disulfide isomerase
precursor (PDI) , partial (89%)
Length = 1861
Score = 27.7 bits (60), Expect = 1.4
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = -1
Query: 152 LYGELADACRPGHRALGGSVTLLTVRKLKFYIYVNFIFLLYF 193
L E+ CRP L G+ LL+ L F++ +N IF L F
Sbjct: 1645 LDSEMQQRCRPPQHPLRGAALLLSKLVLGFFL-LNLIFSLLF 1523
>TC12439 similar to UP|Q9FMJ2 (Q9FMJ2) Ankyrin-like protein, partial (5%)
Length = 534
Score = 26.6 bits (57), Expect = 3.1
Identities = 11/24 (45%), Positives = 17/24 (70%)
Frame = -2
Query: 173 LLTVRKLKFYIYVNFIFLLYFSVR 196
L +RKL ++V FIFLL++ +R
Sbjct: 530 LSIIRKLLCRLFVQFIFLLFWQIR 459
>BP075375
Length = 366
Score = 26.6 bits (57), Expect = 3.1
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = -3
Query: 101 YCVRWYLMYLVGCL 114
YCV W LMYL+G L
Sbjct: 88 YCVYWRLMYLLGLL 47
>AW720071
Length = 377
Score = 25.4 bits (54), Expect = 6.9
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -1
Query: 152 LYGELADACRPGHRALGGSVTLLTVRKLKFY 182
LY +LAD+ RAL TL+ ++L+FY
Sbjct: 284 LYAKLADSVMTSRRALAARSTLID-QRLRFY 195
>AV408775
Length = 427
Score = 25.4 bits (54), Expect = 6.9
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = -1
Query: 89 PEELEELARVRTYCVRWYL 107
PE LE + R R C+RW L
Sbjct: 157 PERLERIQRRRKGCLRWVL 101
>TC14513 homologue to UP|Q7M1K9 (Q7M1K9) Chlorophyll a/b-binding protein
(cab-11), complete
Length = 1025
Score = 25.0 bits (53), Expect = 9.0
Identities = 22/67 (32%), Positives = 30/67 (43%)
Frame = +2
Query: 83 LAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKRIELIYLTTMADGYAGMRNY 142
LA EDPE L+ + RW ++ + G LL + +N I LI + D AG Y
Sbjct: 284 LALAEDPENLKWFVQAELVNGRWAMLGVAGMLLPEVLTN--IGLINVPKWYD--AGKAEY 451
Query: 143 SWGGMTL 149
TL
Sbjct: 452 FASSSTL 472
>BU494461
Length = 495
Score = 25.0 bits (53), Expect = 9.0
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = +1
Query: 170 SVTLLTVRKLKFYIYVNFIFLLYFSV 195
++T+L +R +Y+Y+NF L F +
Sbjct: 157 TITMLLIRDAFYYLYINF*LNLTFFI 234
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,561,511
Number of Sequences: 28460
Number of extensions: 47056
Number of successful extensions: 319
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 319
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 319
length of query: 199
length of database: 4,897,600
effective HSP length: 86
effective length of query: 113
effective length of database: 2,450,040
effective search space: 276854520
effective search space used: 276854520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC140034.3


