

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.13 + phase: 0 /partial
(433 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
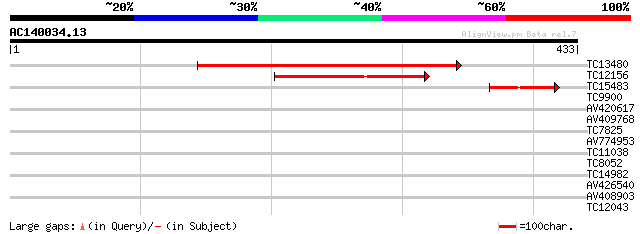
Sequences producing significant alignments: (bits) Value
TC13480 weakly similar to UP|Q9AA65 (Q9AA65) Elongation factor T... 381 e-106
TC12156 homologue to UP|Q93Y02 (Q93Y02) GTP-binding protein typA... 95 3e-20
TC15483 homologue to UP|AAR17698 (AAR17698) GTP-binding protein ... 45 2e-05
TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G, ... 31 0.35
AV420617 30 0.59
AV409768 30 0.77
TC7825 similar to UP|Q9M3Z5 (Q9M3Z5) Lipoxygenase (Fragment) , ... 29 1.7
AV774953 28 2.3
TC11038 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (69%) 28 3.8
TC8052 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {Ara... 27 6.6
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 27 6.6
AV426540 27 8.6
AV408903 27 8.6
TC12043 weakly similar to UP|Q9SND9 (Q9SND9) Anthranilate N-hydr... 27 8.6
>TC13480 weakly similar to UP|Q9AA65 (Q9AA65) Elongation factor Tu family
protein, partial (21%)
Length = 607
Score = 381 bits (979), Expect = e-106
Identities = 186/202 (92%), Positives = 199/202 (98%)
Frame = +1
Query: 144 PPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGEL 203
PPTISMTFGVNDSPLAGRDGTHLTGG+IGDRL+AESETNLAINVLPG+SET+EVQGRGEL
Sbjct: 1 PPTISMTFGVNDSPLAGRDGTHLTGGRIGDRLLAESETNLAINVLPGLSETYEVQGRGEL 180
Query: 204 QLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRA 263
QLGILIENMRREGFELSVSPPKVMYKT+KGQKLEP+EEVTIEVNDEHVG VMEALSHRRA
Sbjct: 181 QLGILIENMRREGFELSVSPPKVMYKTEKGQKLEPVEEVTIEVNDEHVGLVMEALSHRRA 360
Query: 264 EITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPLGNV 323
EITDMGPV+GTVGRTRL LTCPSRGLVGYRSVFSS+TRGTGFMHRAFL+YEK+RGPLGNV
Sbjct: 361 EITDMGPVSGTVGRTRLSLTCPSRGLVGYRSVFSSDTRGTGFMHRAFLEYEKYRGPLGNV 540
Query: 324 RKGVLVSVGFGSITLHALMSLE 345
RKGVLVS+GFG++T HALMSLE
Sbjct: 541 RKGVLVSMGFGTVTAHALMSLE 606
>TC12156 homologue to UP|Q93Y02 (Q93Y02) GTP-binding protein typA (Tyrosine
phosphorylated protein A), partial (30%)
Length = 359
Score = 94.7 bits (234), Expect = 3e-20
Identities = 53/118 (44%), Positives = 72/118 (60%)
Frame = +3
Query: 203 LQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRR 262
L L ILIENMRREG+E V PPKV+ KT + LEP E T+EV + H+G V+E L RR
Sbjct: 3 LHLTILIENMRREGYEFMVGPPKVINKTVNEKLLEPFEIATVEVPEVHMGAVVELLGKRR 182
Query: 263 AEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPL 320
++ DM V G+ G T L P+RGL+G R+ + +RGT ++ F Y + G +
Sbjct: 183 GQMFDMQGV-GSEGTTFLKYKIPTRGLLGLRNAILTASRGTAILNTIFDSYGPWAGDI 353
>TC15483 homologue to UP|AAR17698 (AAR17698) GTP-binding protein TypA,
partial (11%)
Length = 615
Score = 45.1 bits (105), Expect = 2e-05
Identities = 24/54 (44%), Positives = 33/54 (60%)
Frame = +2
Query: 367 HSRDTDLDVNPVRAKQLTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
H R DL +N + K TN+RS NK+ TV L P +L++ I Y+ DEL+EV
Sbjct: 5 HQRPGDLSLNVCKKKAATNIRS-NKEQTVILDTPLDYSLDDCIEYIQEDELVEV 163
>TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G,
chloroplast precursor (EF-G), partial (34%)
Length = 1284
Score = 31.2 bits (69), Expect = 0.35
Identities = 15/38 (39%), Positives = 22/38 (57%)
Frame = +1
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAG 273
LEPI +V + +EH+G V+ L+ RR +I G G
Sbjct: 676 LEPIMKVEVVTPEEHLGDVIGDLNSRRGQINSFGDKPG 789
>AV420617
Length = 260
Score = 30.4 bits (67), Expect = 0.59
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = +3
Query: 42 ILLPMPKICHSCLMPLYHMSPHQMQTLMHLSKC*LRNKDS 81
+LL +PK+ CL+PL + SP +M L H C + + S
Sbjct: 81 LLLSLPKVQEQCLVPLSN-SPERMVLLFHFLLCVMPRRKS 197
>AV409768
Length = 405
Score = 30.0 bits (66), Expect = 0.77
Identities = 19/57 (33%), Positives = 27/57 (47%), Gaps = 12/57 (21%)
Frame = +1
Query: 39 IPRILLPMPKICHSCLMPLYHMSPHQMQTLMHL------------SKC*LRNKDSGA 83
+PR LLP P P H+S H + + +HL S+C LR++DS A
Sbjct: 133 LPRTLLPPPPPQPPLPPPSLHLSLHHLNSNLHLQPPHLSRNSTSSSRCCLRHRDSPA 303
>TC7825 similar to UP|Q9M3Z5 (Q9M3Z5) Lipoxygenase (Fragment) , partial
(93%)
Length = 1642
Score = 28.9 bits (63), Expect = 1.7
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 199 GRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLE 237
GRG +Q+G LI+ + L S K+ K+ KG K++
Sbjct: 1289 GRGRIQIGHLIQRHYKPSKSLEPSCKKLKRKSQKGTKIQ 1405
>AV774953
Length = 579
Score = 28.5 bits (62), Expect = 2.3
Identities = 25/98 (25%), Positives = 44/98 (44%), Gaps = 2/98 (2%)
Frame = -2
Query: 115 SIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGR--DGTHLTGGKIG 172
S++GLSS S G V + SALP+ E D ++ ++F D R G+ GG +
Sbjct: 452 SVSGLSSASFGSAVFSSAADSALPSSEADSVSV-LSFSDPDDDFRDRFFGGSVSAGGGVD 276
Query: 173 DRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIE 210
L ++ +P ++ G + L +L++
Sbjct: 275 LAFTLILADRLGVSPVP--QSNSDIAGLPRMCLQVLLQ 168
>TC11038 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (69%)
Length = 596
Score = 27.7 bits (60), Expect = 3.8
Identities = 26/78 (33%), Positives = 38/78 (48%), Gaps = 4/78 (5%)
Frame = +1
Query: 268 MGPVAGTVG-RTRLCLTCPSRGLVGYRSVFSSETR---GTGFMHRAFLKYEKFRGPLGNV 323
+ P++ T+ R+ LCL P GLV ++ S + G H+ LK L +
Sbjct: 379 LDPISLTIKHRSTLCL--PISGLVHFQLRISPSLQMEEGLSIPHQEMLK-------LTTL 531
Query: 324 RKGVLVSVGFGSITLHAL 341
GVLV +GF S+TL L
Sbjct: 532 LWGVLVLLGFSSMTLSLL 585
>TC8052 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {Arabidopsis
thaliana;}, partial (55%)
Length = 1389
Score = 26.9 bits (58), Expect = 6.6
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = -2
Query: 39 IPRILLPMPKICHSCLMPLYHMSPHQMQ 66
+ R+LL P I SC Y +S HQ+Q
Sbjct: 548 VRRLLLQYPSISRSCRNEQYAVSGHQLQ 465
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 26.9 bits (58), Expect = 6.6
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +1
Query: 38 HIPRILLPMPKICHSCLMPLYHMSPHQ 64
H+P+ LLP P H L PL + PH+
Sbjct: 304 HLPQTLLPQP---HLLLNPLLQIRPHR 375
>AV426540
Length = 282
Score = 26.6 bits (57), Expect = 8.6
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = +3
Query: 37 PHIPRILLPMPKICHSCLMPLYHMSPHQMQTLMHL 71
P P +LLP+PK S L P + +S +Q+L L
Sbjct: 156 PRFPCLLLPIPK---SSLFPPFSLSAVSLQSLFSL 251
>AV408903
Length = 426
Score = 26.6 bits (57), Expect = 8.6
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -2
Query: 40 PRILLPMPKICHSCLMPLYHMSPHQMQTLMHLSKC*LR 77
P+I P P+ C S + SPHQ L H+S+ LR
Sbjct: 341 PKIQPPQPRNCIS-----HRESPHQPHELPHISQTPLR 243
>TC12043 weakly similar to UP|Q9SND9 (Q9SND9) Anthranilate
N-hydroxycinnamoyl/benzoyltransferase-like protein,
partial (11%)
Length = 421
Score = 26.6 bits (57), Expect = 8.6
Identities = 16/54 (29%), Positives = 23/54 (41%)
Frame = -2
Query: 121 SPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDR 174
+PS TT+ I S +P PPT +T + G +TGG + R
Sbjct: 177 NPSKCSNFTTIFIRSTIPNRHRLPPTKIITIKIEPR*TIGDKRHRITGGSLSRR 16
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,743,898
Number of Sequences: 28460
Number of extensions: 94485
Number of successful extensions: 512
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 506
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 510
length of query: 433
length of database: 4,897,600
effective HSP length: 93
effective length of query: 340
effective length of database: 2,250,820
effective search space: 765278800
effective search space used: 765278800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC140034.13


