

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140030.1 + phase: 0
(336 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
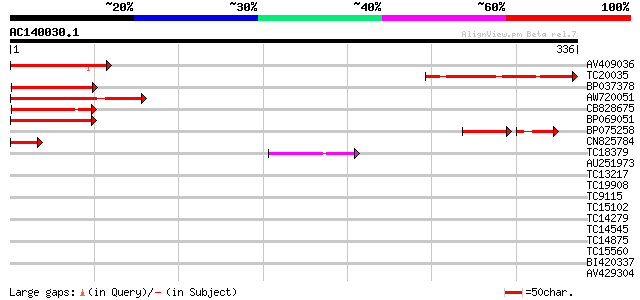
Score E
Sequences producing significant alignments: (bits) Value
AV409036 88 2e-18
TC20035 similar to UP|Q39540 (Q39540) AOBP (Ascorbate oxidase pr... 87 4e-18
BP037378 86 1e-17
AW720051 82 1e-16
CB828675 81 3e-16
BP069051 79 8e-16
BP075258 43 2e-06
CN825784 42 2e-04
TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycop... 40 6e-04
AU251973 36 0.010
TC13217 similar to UP|Q9M4G1 (Q9M4G1) Dof zinc finger protein, p... 35 0.014
TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like ... 35 0.018
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 35 0.018
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 34 0.030
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 34 0.030
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 34 0.030
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 34 0.030
TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich... 34 0.030
BI420337 34 0.040
AV429304 34 0.040
>AV409036
Length = 429
Score = 88.2 bits (217), Expect = 2e-18
Identities = 39/67 (58%), Positives = 49/67 (72%), Gaps = 7/67 (10%)
Frame = +1
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKN-------KNLSTSHY 53
++TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G RK+ KN S+S
Sbjct: 199 INTKFCYYNNYNLSQPRHFCKNCRRYWTKGGVLRNVPVGGGCRKSTKRSSKPKNSSSSET 378
Query: 54 RHLIIPE 60
L +PE
Sbjct: 379 TDLQLPE 399
>TC20035 similar to UP|Q39540 (Q39540) AOBP (Ascorbate oxidase
promoter-binding protein), partial (16%)
Length = 434
Score = 87.0 bits (214), Expect = 4e-18
Identities = 50/91 (54%), Positives = 62/91 (67%), Gaps = 1/91 (1%)
Frame = +2
Query: 247 EKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKANSLNGGGMFNGFQS-K 305
E+P + N SVL PKTLR DDP+EAAKSS+W+ LGI N+ SL+GG MF F+S K
Sbjct: 44 EEPSKQRNG---SVLTPKTLRIDDPNEAAKSSIWATLGIKNE---SLSGGSMFKAFKSTK 205
Query: 306 GKDMNHSVGTSPLMYANPAALSRSRTFHEET 336
G + NH S ++ ANPAALSRS FHE +
Sbjct: 206 GDEKNHV--DSSVLRANPAALSRSLNFHENS 292
>BP037378
Length = 546
Score = 85.5 bits (210), Expect = 1e-17
Identities = 34/51 (66%), Positives = 42/51 (81%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSH 52
+TKFCYFNNY+++QPRHFCK C+RYWT GG +RNVPVG G R+NK S+
Sbjct: 262 NTKFCYFNNYSLSQPRHFCKTCKRYWTRGGALRNVPVGGGFRRNKKHKKSN 414
>AW720051
Length = 564
Score = 82.4 bits (202), Expect = 1e-16
Identities = 37/81 (45%), Positives = 50/81 (61%)
Frame = +1
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
++TKFCY+NNYN++QPRHFCK C+RYWT GG +RN+ VG+G RK K + +P
Sbjct: 160 LNTKFCYYNNYNLSQPRHFCKGCRRYWTKGGLLRNITVGSGSRKPKRSINKN----TVPS 327
Query: 61 GAKLNSPNGIHSLGNGAAVLT 81
P S G++ LT
Sbjct: 328 APPPPPPPEPQSSSGGSSTLT 390
>CB828675
Length = 552
Score = 80.9 bits (198), Expect = 3e-16
Identities = 33/50 (66%), Positives = 42/50 (84%)
Frame = +3
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTS 51
+TKFCY+NNYN +QPRH+C+ C+R+WT GGT+RNVPVG G RKNK + S
Sbjct: 303 NTKFCYYNNYNKSQPRHYCRACKRHWTKGGTLRNVPVG-GVRKNKKIKKS 449
>BP069051
Length = 359
Score = 79.3 bits (194), Expect = 8e-16
Identities = 30/51 (58%), Positives = 43/51 (83%)
Frame = -2
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTS 51
++T+FCY+NNY++ QPR+FC C+RYWT GG++R++PVG G RKNK+ S S
Sbjct: 271 INTQFCYYNNYSLTQPRYFC*TCRRYWTEGGSLRHIPVGGGSRKNKHRSNS 119
>BP075258
Length = 490
Score = 43.1 bits (100), Expect(2) = 2e-06
Identities = 20/30 (66%), Positives = 25/30 (82%), Gaps = 1/30 (3%)
Frame = -1
Query: 269 DDPSEAAKSSLWSKLGINND-KANSLNGGG 297
DDP+EA+K+S+WS LGI +D K NSL GGG
Sbjct: 487 DDPNEASKTSIWSTLGIKHDEKGNSLYGGG 398
Score = 24.3 bits (51), Expect(2) = 2e-06
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = -2
Query: 301 GFQSKGKDMNHSVGTSPLMYANPAA 325
GF+ K NH SP ++ANPAA
Sbjct: 384 GFEKK----NHVDEASPFLHANPAA 322
>CN825784
Length = 548
Score = 41.6 bits (96), Expect = 2e-04
Identities = 15/19 (78%), Positives = 19/19 (99%)
Frame = +1
Query: 1 MDTKFCYFNNYNVNQPRHF 19
MDTKFCY++NY+VNQPR+F
Sbjct: 490 MDTKFCYYHNYHVNQPRYF 546
>TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycoprotein
DZ-HRGP precursor, partial (8%)
Length = 584
Score = 40.0 bits (92), Expect = 6e-04
Identities = 22/55 (40%), Positives = 26/55 (47%), Gaps = 1/55 (1%)
Frame = +1
Query: 154 PPQLQHIPSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPP-PWNIP 207
P ++ P P LPY + PPP PP P A P PP YW PP P+ P
Sbjct: 178 PYGMEFSPYPSLPYPQHQFYPPPMAPPPPPPPA-PAPPAPPPYWAAPPPRPYGPP 339
>AU251973
Length = 205
Score = 35.8 bits (81), Expect = 0.010
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +2
Query: 154 PPQLQHIPSPFLPYTWNSAMPPPT--FCPPNYPLAFYTPVTPPAYWGCMPPP 203
PP ++ P PY ++S PPP PP P + +P PP + PPP
Sbjct: 47 PPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPP 202
Score = 35.8 bits (81), Expect = 0.010
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +2
Query: 154 PPQLQHIPSPFLPYTWNSAMPPPT--FCPPNYPLAFYTPVTPPAYWGCMPPP 203
PP ++ P PY ++S PPP PP P + +P PP + PPP
Sbjct: 17 PPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPPPYKYSSPPPP 172
>TC13217 similar to UP|Q9M4G1 (Q9M4G1) Dof zinc finger protein, partial
(15%)
Length = 422
Score = 35.4 bits (80), Expect = 0.014
Identities = 12/16 (75%), Positives = 16/16 (100%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPR 17
+TKFCY+NNYN++QPR
Sbjct: 373 NTKFCYYNNYNLSQPR 420
>TC19908 similar to GB|CAA60022.1|791150|VURNEXT26 extensin-like protein
{Vigna unguiculata;} , partial (49%)
Length = 472
Score = 35.0 bits (79), Expect = 0.018
Identities = 27/83 (32%), Positives = 35/83 (41%)
Frame = +2
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESD 220
PSP PY + S PP PP Y Y PP++ PPP+ P S+S
Sbjct: 47 PSPPPPYVYKSPPPPSPSPPPPY---IYKSPPPPSH--SPPPPYVYKSPPPPSSSHPHHS 211
Query: 221 SAHSSVPTLGKHSRDGNIITSVN 243
++S P RD I +VN
Sbjct: 212 YLYNSPPPPRY*PRD*GNIVAVN 280
Score = 29.3 bits (64), Expect = 0.98
Identities = 24/67 (35%), Positives = 27/67 (39%), Gaps = 3/67 (4%)
Frame = +2
Query: 162 SPFLPYTWNSAMPPPTFCPPNYPLAF---YTPVTPPAYWGCMPPPWNIPCISPGSASVNE 218
SP PY + S PP PP Y +P PP Y PPP P SP V +
Sbjct: 2 SPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYIYKSPPP---PSHSPPPPYVYK 172
Query: 219 SDSAHSS 225
S SS
Sbjct: 173 SPPPPSS 193
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 35.0 bits (79), Expect = 0.018
Identities = 38/140 (27%), Positives = 53/140 (37%), Gaps = 5/140 (3%)
Frame = +1
Query: 100 ERTQNCVSNGFHHASSYGGENDHSIGVSVTASNSSERKSHT----STNGLVDKGVEGFPP 155
+R ++ H + + + HS S +S SS SH + N K PP
Sbjct: 97 QRKTTTLTRNHSHQNHHHSQ*HHSPFHSPPSSPSSNPNSHHLLPHNNNPTRSKSQPKPPP 276
Query: 156 Q-LQHIPSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSA 214
H P P P N PP+ P+ PL TP P A PP + S GSA
Sbjct: 277 SPTSHSPPPPHPRRPNPPSNPPSPPTPSAPL---TP*APTAPSTLPTPPHSAAPPSSGSA 447
Query: 215 SVNESDSAHSSVPTLGKHSR 234
+ + S + S P S+
Sbjct: 448 TTSASATTRLSTPPTTNRSQ 507
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 34.3 bits (77), Expect = 0.030
Identities = 40/181 (22%), Positives = 65/181 (35%), Gaps = 15/181 (8%)
Frame = +1
Query: 130 ASNSSERKSHTST------NGLVDKGVEGFPPQLQHIPSPFLPYTWNSAMPPPTFCPPNY 183
AS + + SH+ST + K P P P + P P+ PPN
Sbjct: 283 ASPPTPKSSHSSTPPPSRSSSTTPKSKNATSLTAPSCPPPASPA---ATAPAPSKTPPNP 453
Query: 184 PLAFYTP---------VTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTLGKHSR 234
P P TP + P P P +P S E + S+ P+ S
Sbjct: 454 PTNSSKPSSCKSSLPNTTPSSLSSAPPSPTTSPISTPSSPENQEKPA--STPPSAPPRST 627
Query: 235 DGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKANSLN 294
+ + +N+P++ S +L PS + KSS K+ +++ ++S
Sbjct: 628 SSSSSSPINTPRKPSSATKAPASSPPGSELSLHHSPPSASRKSSTTPKIS-SSEPSHSRR 804
Query: 295 G 295
G
Sbjct: 805 G 807
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 34.3 bits (77), Expect = 0.030
Identities = 26/73 (35%), Positives = 33/73 (44%), Gaps = 6/73 (8%)
Frame = +1
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFY------TPVTPPAYWGCMPPPWNIPCISPGSA 214
PSP PY + S PPP+ PP P +Y +P PP Y+ PPP P SP
Sbjct: 112 PSPPPPYYYQSP-PPPSHSPP--PPYYYKSPPPPSPSPPPPYYYKSPPP---PSPSPPPP 273
Query: 215 SVNESDSAHSSVP 227
+S S +P
Sbjct: 274 YYYKSPPPPSPIP 312
Score = 33.1 bits (74), Expect = 0.068
Identities = 24/76 (31%), Positives = 31/76 (40%), Gaps = 9/76 (11%)
Frame = +1
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVT----PPAYWGCMPPPWNIP-----CISP 211
PSP PY + S PP PP Y P + PP Y+ PPP + P +SP
Sbjct: 208 PSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHPPYYYKSPPPPTSSPPPPYHYVSP 387
Query: 212 GSASVNESDSAHSSVP 227
S + H + P
Sbjct: 388 PPPSPSPPPPYHYASP 435
Score = 32.3 bits (72), Expect = 0.12
Identities = 22/62 (35%), Positives = 27/62 (43%)
Frame = +1
Query: 154 PPQLQHIPSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGS 213
PP IP P PY + S PP + PP Y V+PP PPP++ P S
Sbjct: 289 PPPPSPIPHP--PYYYKSPPPPTSSPPPPYHY-----VSPPPPSPSPPPPYHYASPPPPS 447
Query: 214 AS 215
S
Sbjct: 448 PS 453
Score = 32.0 bits (71), Expect = 0.15
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +1
Query: 166 PYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPP 203
PY + S PPP P Y + +P PP Y+ PPP
Sbjct: 4 PYWYKSPPPPPPVPKPYY---YKSPPPPPYYYKSPPPP 108
Score = 30.4 bits (67), Expect = 0.44
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 4/66 (6%)
Frame = +1
Query: 154 PPQLQHI----PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCI 209
PP ++ PSP P ++ A PPP P P Y +PP PPP++
Sbjct: 361 PPPYHYVSPPPPSPSPPPPYHYASPPP---PSPSPAPTYIYKSPPPPVKLPPPPYHYTSP 531
Query: 210 SPGSAS 215
P S S
Sbjct: 532 PPPSPS 549
Score = 27.3 bits (59), Expect = 3.7
Identities = 19/55 (34%), Positives = 24/55 (43%), Gaps = 5/55 (9%)
Frame = +1
Query: 154 PPQLQHIPSPFLPYTWNSAMPP-----PTFCPPNYPLAFYTPVTPPAYWGCMPPP 203
PP +P P PY + S PP PT+ + P +P PP Y PPP
Sbjct: 481 PPPPVKLPPP--PYHYTSPPPPSPSPAPTYIYKSPPPPTKSP-PPPVYIYASPPP 636
Score = 26.2 bits (56), Expect = 8.3
Identities = 14/36 (38%), Positives = 16/36 (43%)
Frame = +1
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAY 196
PSP Y + S PPP P P+ Y PP Y
Sbjct: 544 PSPAPTYIYKS--PPPPTKSPPPPVYIYASPPPPIY 645
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 34.3 bits (77), Expect = 0.030
Identities = 33/139 (23%), Positives = 54/139 (38%), Gaps = 7/139 (5%)
Frame = -2
Query: 155 PQLQHIPSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIP------- 207
P I S +P ++PP PPN P+ P+ P PP IP
Sbjct: 664 PSKSSIASSPIPPNLEKSIPPNPSIPPNPPIPPIPPIPPIPPSPPRPPSPPIPIPPNFEK 485
Query: 208 CISPGSASVNESDSAHSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLR 267
I P S S S S+ S+ + + G + + N+ P + +TE+ + P +
Sbjct: 484 SIPPISGSATTSSSSSSTHLSQSTFT*CGGLPSPRNNLFHHPLSAFCTTENMLRTPILIL 305
Query: 268 FDDPSEAAKSSLWSKLGIN 286
+KSS + G++
Sbjct: 304 LSSTLTLSKSSSLTSYGLS 248
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 34.3 bits (77), Expect = 0.030
Identities = 26/90 (28%), Positives = 41/90 (44%), Gaps = 3/90 (3%)
Frame = +1
Query: 173 MPPPTFCPPNYP---LAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTL 229
+P PT PP+ P + TP +PPA + PP P S +A+ + S + ++ PT
Sbjct: 115 LPAPTI*PPSNPKTSTSTPTPASPPASTSPLTPPTTSPSSSTSTAAPSAS-APLTTTPTT 291
Query: 230 GKHSRDGNIITSVNSPKEKPETEHNSTESS 259
TS SP+ + + ST +S
Sbjct: 292 A---------TSTTSPRRQTPSSSPSTTAS 354
>TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich protein 1
(NDPP-1). {Mus musculus;}, partial (8%)
Length = 1134
Score = 34.3 bits (77), Expect = 0.030
Identities = 45/177 (25%), Positives = 70/177 (39%), Gaps = 9/177 (5%)
Frame = +3
Query: 116 YGGENDHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQL---------QHIPSPFLP 166
+ ++ H ++ TAS SS S S++ PP L H+PSP
Sbjct: 84 FTNKSHHHYPLTTTASASSSSSSSPSSS*PSPSLPPPSPPPLPPPQAPPLPSHLPSPEPS 263
Query: 167 YTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSV 226
T + PPP P PL TP PP+ P +P S A +
Sbjct: 264 STMPPSTPPPQPTKP-CPLPKSTPSQPPS-----SAPHRVP---------TSSSLASPTS 398
Query: 227 PTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKL 283
P+ G HS T++ +P +ST +S P + ++P+ A+K + +S L
Sbjct: 399 PSSGPHS------TTMAAP-------FSSTRTSTPSPNS---NNPTPASKPTTFSSL 521
>BI420337
Length = 397
Score = 33.9 bits (76), Expect = 0.040
Identities = 21/57 (36%), Positives = 25/57 (43%), Gaps = 7/57 (12%)
Frame = +1
Query: 154 PPQLQHIPSPFLPYTWNSAMPP-------PTFCPPNYPLAFYTPVTPPAYWGCMPPP 203
PP H P P PY ++S PP P+ PP Y P TP Y+ PPP
Sbjct: 139 PPPPMHSPPP--PYHYSSPPPPPKKPYKYPSPPPPVYKYKSPPPRTPAYYYKSPPPP 303
Score = 27.3 bits (59), Expect = 3.7
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 2/45 (4%)
Frame = +1
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPP--AYWGCMPPP 203
P P PY ++S PPP P P + +P PP Y PPP
Sbjct: 106 PPPPKPYYYHS--PPPPMHSPPPPYHYSSPPPPPKKPYKYPSPPP 234
Score = 26.9 bits (58), Expect = 4.8
Identities = 14/47 (29%), Positives = 20/47 (41%)
Frame = +1
Query: 161 PSPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIP 207
P P P + + PPP P P + +P P + PPP+ P
Sbjct: 256 PPPRTPAYYYKSPPPP----PKNPYKYPSPPPPVYKYKSPPPPFKYP 384
>AV429304
Length = 355
Score = 33.9 bits (76), Expect = 0.040
Identities = 26/84 (30%), Positives = 33/84 (38%)
Frame = +3
Query: 175 PPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSAHSSVPTLGKHSR 234
PPT+ PPN P TP TP + +PP P SP + VPT K
Sbjct: 9 PPTYSPPNVPSPPQTPHTP---YVPIPPKTPSPVYSPPNVPSPPQTPHAPYVPTPPKTPS 179
Query: 235 DGNIITSVNSPKEKPETEHNSTES 258
+V SP + P + T S
Sbjct: 180 PVYSPPNVPSPPQTPHAPYVPTPS 251
Score = 33.5 bits (75), Expect = 0.052
Identities = 27/79 (34%), Positives = 30/79 (37%), Gaps = 1/79 (1%)
Frame = +3
Query: 154 PPQLQHIPS-PFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPG 212
PPQ H P P P T P P + PPN P P TP A + PP P SP
Sbjct: 42 PPQTPHTPYVPIPPKT-----PSPVYSPPNVP---SPPQTPHAPYVPTPPKTPSPVYSPP 197
Query: 213 SASVNESDSAHSSVPTLGK 231
+ VPT K
Sbjct: 198 NVPSPPQTPHAPYVPTPSK 254
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.130 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,938,189
Number of Sequences: 28460
Number of extensions: 159844
Number of successful extensions: 1160
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 986
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1083
length of query: 336
length of database: 4,897,600
effective HSP length: 91
effective length of query: 245
effective length of database: 2,307,740
effective search space: 565396300
effective search space used: 565396300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC140030.1


