

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140025.10 - phase: 0
(344 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
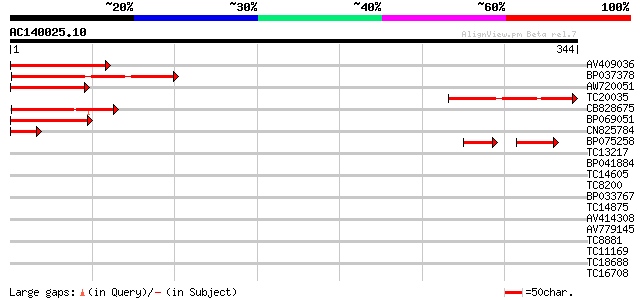
Score E
Sequences producing significant alignments: (bits) Value
AV409036 94 4e-20
BP037378 86 9e-18
AW720051 85 2e-17
TC20035 similar to UP|Q39540 (Q39540) AOBP (Ascorbate oxidase pr... 84 5e-17
CB828675 82 1e-16
BP069051 80 7e-16
CN825784 43 7e-05
BP075258 32 3e-04
TC13217 similar to UP|Q9M4G1 (Q9M4G1) Dof zinc finger protein, p... 37 0.005
BP041884 37 0.006
TC14605 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial... 35 0.018
TC8200 weakly similar to UP|Q9JGS9 (Q9JGS9) PORF2b, partial (23%) 34 0.031
BP033767 33 0.070
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 32 0.12
AV414308 32 0.12
AV779145 32 0.16
TC8881 homologue to UP|ILV5_PEA (O82043) Ketol-acid reductoisome... 32 0.16
TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial... 32 0.20
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 31 0.27
TC16708 31 0.35
>AV409036
Length = 429
Score = 93.6 bits (231), Expect = 4e-20
Identities = 39/61 (63%), Positives = 48/61 (77%)
Frame = +1
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
++TKFCYYNNYN++QPRHFCKNC+RYWT GG +RNVPVG G RK+ SS + + SE
Sbjct: 199 INTKFCYYNNYNLSQPRHFCKNCRRYWTKGGVLRNVPVGGGCRKSTKRSSKPKNSSSSET 378
Query: 61 T 61
T
Sbjct: 379 T 381
>BP037378
Length = 546
Score = 85.9 bits (211), Expect = 9e-18
Identities = 41/101 (40%), Positives = 62/101 (60%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEAT 61
+TKFCY+NNY+++QPRHFCK C+RYWT GG +RNVPVG G R+NK H + + S +T
Sbjct: 262 NTKFCYFNNYSLSQPRHFCKTCKRYWTRGGALRNVPVGGGFRRNK---KHKKSNSSSSST 432
Query: 62 LQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQ 102
++ + S N T + + ++ ++ H T+Q
Sbjct: 433 SESDKQSLS---NSTTNELNIPTNMTSTLQNLSFHGGNTIQ 546
>AW720051
Length = 564
Score = 85.1 bits (209), Expect = 2e-17
Identities = 33/48 (68%), Positives = 41/48 (84%)
Frame = +1
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSS 48
++TKFCYYNNYN++QPRHFCK C+RYWT GG +RN+ VG+G RK K S
Sbjct: 160 LNTKFCYYNNYNLSQPRHFCKGCRRYWTKGGLLRNITVGSGSRKPKRS 303
>TC20035 similar to UP|Q39540 (Q39540) AOBP (Ascorbate oxidase
promoter-binding protein), partial (16%)
Length = 434
Score = 83.6 bits (205), Expect = 5e-17
Identities = 50/79 (63%), Positives = 58/79 (73%), Gaps = 1/79 (1%)
Frame = +2
Query: 267 SLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSVPPRRLFQAFPS-KCDEKNHLVQVSS 325
S+ PKTLRIDD EA KSS W TLGIKN +S+ +F+AF S K DEKNH+ SS
Sbjct: 71 SVLTPKTLRIDDPNEAAKSSIWATLGIKN---ESLSGGSMFKAFKSTKGDEKNHV--DSS 235
Query: 326 VLQANPAALSRSLHFHETS 344
VL+ANPAALSRSL+FHE S
Sbjct: 236 VLRANPAALSRSLNFHENS 292
>CB828675
Length = 552
Score = 82.0 bits (201), Expect = 1e-16
Identities = 39/72 (54%), Positives = 49/72 (67%), Gaps = 7/72 (9%)
Frame = +3
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK-------SSSSHYRQ 54
+TKFCYYNNYN +QPRH+C+ C+R+WT GGT+RNVPVG G RKNK S SS
Sbjct: 303 NTKFCYYNNYNKSQPRHYCRACKRHWTKGGTLRNVPVG-GVRKNKKIKKSTTSPSSGTTT 479
Query: 55 ITVSEATLQNSR 66
++S T N +
Sbjct: 480 TSISTTTTSNGK 515
>BP069051
Length = 359
Score = 79.7 bits (195), Expect = 7e-16
Identities = 30/50 (60%), Positives = 42/50 (84%)
Frame = -2
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSS 50
++T+FCYYNNY++ QPR+FC C+RYWT GG++R++PVG G RKNK S+
Sbjct: 271 INTQFCYYNNYSLTQPRYFC*TCRRYWTEGGSLRHIPVGGGSRKNKHRSN 122
>CN825784
Length = 548
Score = 43.1 bits (100), Expect = 7e-05
Identities = 16/19 (84%), Positives = 19/19 (99%)
Frame = +1
Query: 1 MDTKFCYYNNYNVNQPRHF 19
MDTKFCYY+NY+VNQPR+F
Sbjct: 490 MDTKFCYYHNYHVNQPRYF 546
>BP075258
Length = 490
Score = 32.3 bits (72), Expect(2) = 3e-04
Identities = 13/21 (61%), Positives = 17/21 (80%)
Frame = -1
Query: 276 IDDSEEAEKSSFWTTLGIKNN 296
IDD EA K+S W+TLGIK++
Sbjct: 490 IDDPNEASKTSIWSTLGIKHD 428
Score = 27.7 bits (60), Expect(2) = 3e-04
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -2
Query: 308 QAFPSKCDEKNHLVQVSSVLQANPAA 333
+AF ++KNH+ + S L ANPAA
Sbjct: 399 EAFSQGFEKKNHVDEASPFLHANPAA 322
>TC13217 similar to UP|Q9M4G1 (Q9M4G1) Dof zinc finger protein, partial
(15%)
Length = 422
Score = 37.0 bits (84), Expect = 0.005
Identities = 13/16 (81%), Positives = 16/16 (99%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPR 17
+TKFCYYNNYN++QPR
Sbjct: 373 NTKFCYYNNYNLSQPR 420
>BP041884
Length = 489
Score = 36.6 bits (83), Expect = 0.006
Identities = 32/105 (30%), Positives = 45/105 (42%), Gaps = 6/105 (5%)
Frame = +3
Query: 153 QAMWNDHSF-PPQGGYFPHGTPWHLPWNPVQMSSPIPPPAFCPPG-FSMPFYPATTYWGC 210
+ ++ DH+ PP G + P TP LP +S+ P F P G PF PA
Sbjct: 42 KTLFTDHTCSPPNGAHAP--TPVTLP-----ISAGARPSFFAPLGAHGGPFTPAPPAANV 200
Query: 211 TMPSAW----NIPRQAQPSSPNGANHDSTPNSPTLGKHSREDNML 251
+ W N+ AQ S ++ PN ++ KHSR N L
Sbjct: 201 NALAGWMVNANLSTSAQSSVVAASSFSGHPNQGSILKHSRTPNTL 335
>TC14605 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial (59%)
Length = 1323
Score = 35.0 bits (79), Expect = 0.018
Identities = 21/57 (36%), Positives = 26/57 (44%)
Frame = -2
Query: 168 FPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQP 224
+PH T WH N Q + PP+ P FS P P PS++NIP Q P
Sbjct: 647 YPHKT*WH*HSNQAQST----PPSHTP--FSQPV*PPQIAASSQTPSSYNIPSQTNP 495
>TC8200 weakly similar to UP|Q9JGS9 (Q9JGS9) PORF2b, partial (23%)
Length = 930
Score = 34.3 bits (77), Expect = 0.031
Identities = 23/73 (31%), Positives = 31/73 (41%), Gaps = 3/73 (4%)
Frame = +2
Query: 170 HGTPWH---LPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSS 226
H PW P P SSP P PA C F +P P +P ++ P Q S+
Sbjct: 17 HLKPWPNSLFPSLPSSSSSPSPTPAICNNPFPIPI-PIPIPTPTPIPISFPHPNA*QTSN 193
Query: 227 PNGANHDSTPNSP 239
PN +P++P
Sbjct: 194 PNNTTPFRSPSNP 232
>BP033767
Length = 541
Score = 33.1 bits (74), Expect = 0.070
Identities = 29/100 (29%), Positives = 36/100 (36%)
Frame = +1
Query: 141 STEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMP 200
ST G++TN S + H PQ + H P N P PPP PP S P
Sbjct: 130 STGGSSTNPSHKAPF---HPLNPQTHHHHHHLPHLQTQNTHFPPPPQPPPPKMPPHLSSP 300
Query: 201 FYPATTYWGCTMPSAWNIPRQAQPSSPNGANHDSTPNSPT 240
+ PS P S P+ + PN PT
Sbjct: 301 HTHLHHHLLLPPPS----PPSPPTSPPSSSLKPPNPNPPT 408
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 32.3 bits (72), Expect = 0.12
Identities = 35/115 (30%), Positives = 52/115 (44%), Gaps = 8/115 (6%)
Frame = +1
Query: 134 SSVTSTKSTEGATTNV--SQEQAMWNDHSFPPQGGYFPHGTPWHLPWNPVQMSSPIPPPA 191
SS+TS+ ST+ A +N +Q + PP TP P +P +SP+ PP
Sbjct: 46 SSLTSSDSTKMAASNA*WAQTPPLPAPTI*PPSNPKTSTSTP--TPASPPASTSPLTPPT 219
Query: 192 FCPPGFSMPFYPATTYWGCTMP---SAWNIPRQAQPSS--PNGANHDSTPN-SPT 240
P + P+ + T P ++ PR+ PSS A+ STP+ SPT
Sbjct: 220 TSPSSSTSTAAPSASAPLTTTPTTATSTTSPRRQTPSSSPSTTASPRSTPSPSPT 384
>AV414308
Length = 389
Score = 32.3 bits (72), Expect = 0.12
Identities = 29/112 (25%), Positives = 43/112 (37%), Gaps = 5/112 (4%)
Frame = +3
Query: 130 QSNKSSVTSTKSTEGAT-----TNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWNPVQMS 184
+ N S ++ T GAT T ++E + N S PP+ +P
Sbjct: 69 RKNSKSSSNPTPTTGATPQKKSTTSNKESLLQNPPSPPPEA-------------SPSSPG 209
Query: 185 SPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGANHDSTP 236
PPP P S ATT G + P + PR +SP+ + +TP
Sbjct: 210 PGSPPPPHHAPSSSWSTATATTSPGPSKPPQSSSPRWVSLASPSTSKATATP 365
>AV779145
Length = 417
Score = 32.0 bits (71), Expect = 0.16
Identities = 29/111 (26%), Positives = 38/111 (34%), Gaps = 3/111 (2%)
Frame = +2
Query: 163 PQGGYFPHGTPWHLPWNPVQMSSPIP---PPAFCPPGFSMPFYPATTYWGCTMPSAWNIP 219
P F H P P +P +P P PP PG + PA T P
Sbjct: 65 PSPARFSHSKP---PPSPTTQPTPSPHGTPPHATAPGTASLAAPAAT-----------SP 202
Query: 220 RQAQPSSPNGANHDSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWF 270
P SP+ AN TP+ P+L + + +S SL F
Sbjct: 203 PLTSPLSPSPANSPPTPSPPSLSSPPSPSPTTRCAPATSHPRLSSLTSLRF 355
>TC8881 homologue to UP|ILV5_PEA (O82043) Ketol-acid reductoisomerase,
chloroplast precursor (Acetohydroxy-acid
reductoisomerase) (Alpha-keto-beta-hydroxylacil
reductoisomerase) , partial (52%)
Length = 1310
Score = 32.0 bits (71), Expect = 0.16
Identities = 16/64 (25%), Positives = 26/64 (40%)
Frame = -1
Query: 130 QSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWNPVQMSSPIPP 189
QS SS+ + + QA+WN+H+ P F PW + + + S + P
Sbjct: 551 QSGPSSMPQRHQQ*MDSKGLDHLQALWNEHAHQPSTSGFDQFYPWEMQADLLSHKSDVRP 372
Query: 190 PAFC 193
C
Sbjct: 371 KQHC 360
>TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial (14%)
Length = 650
Score = 31.6 bits (70), Expect = 0.20
Identities = 46/185 (24%), Positives = 73/185 (38%), Gaps = 23/185 (12%)
Frame = +3
Query: 174 WHLPWNPVQMSSPIP--PPAF-CPPGFSMPFYPATTY------W--GCTMPSAWN----- 217
W++ N ++ SP P PP P F P+ P+++ W +P +W+
Sbjct: 93 WNININSMRPPSPQPQAPPFLPTPSNFLTPYPPSSSSSLPFPSWPENQEVPESWSQLLMS 272
Query: 218 --IPRQAQPSSPNGANHDST--PNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKT 273
+ + + + N T PN+ + +E++ G G +E+ KS W PK+
Sbjct: 273 GVVAEEDKGAMAQFQNQMLTQAPNASFIDV-KQENSGNSYVYGHGNEELHSAKSCWSPKS 449
Query: 274 LRIDDSEEAEKSSFWTTLGIKNNNADSV---PPRRLFQAFPSKCDEKNHLVQVSSVLQAN 330
S + L NNN D V PP L S+C+ HLV L N
Sbjct: 450 CVTSFSSD--------MLDFSNNNRDHVRLHPPPDL----SSECNSTLHLVGQ*RKLGFN 593
Query: 331 PAALS 335
P L+
Sbjct: 594 PLHLN 608
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 31.2 bits (69), Expect = 0.27
Identities = 34/115 (29%), Positives = 46/115 (39%), Gaps = 2/115 (1%)
Frame = +3
Query: 128 EDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWNPVQMSSPI 187
E S+ SS +S+ S+ + +S A + S PP +P + S
Sbjct: 60 ESLSSSSSSSSSSSSPLSLNQISPPNAPPS*LSAPP---------------SPEERFSGT 194
Query: 188 PPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGANHD--STPNSPT 240
PPP PP PAT P+ R A P SP+ AN S+P SPT
Sbjct: 195 PPP---PP-------PATGSASTATPTPPTSSRSASPPSPSPANSHMASSPPSPT 329
>TC16708
Length = 515
Score = 30.8 bits (68), Expect = 0.35
Identities = 24/72 (33%), Positives = 29/72 (39%), Gaps = 1/72 (1%)
Frame = -3
Query: 170 HGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNG 229
H T H P + +P PP A CPP P+ CT + R+ QPSS
Sbjct: 249 HPTLQHTP-RCISHPAPSPPSATCPPS-----SPSNPRRHCTCTRSHLHRRRPQPSSDTW 88
Query: 230 AN-HDSTPNSPT 240
N S P PT
Sbjct: 87 RNPRSSLPPGPT 52
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,984,049
Number of Sequences: 28460
Number of extensions: 143388
Number of successful extensions: 893
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 845
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 876
length of query: 344
length of database: 4,897,600
effective HSP length: 91
effective length of query: 253
effective length of database: 2,307,740
effective search space: 583858220
effective search space used: 583858220
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC140025.10


