

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
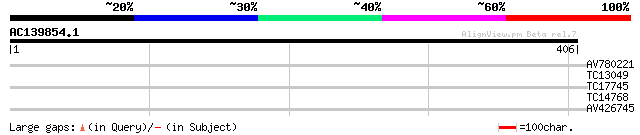
Query= AC139854.1 - phase: 0 /pseudo
(406 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780221 30 0.55
TC13049 28 2.8
TC17745 28 3.6
TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 27 4.7
AV426745 27 8.0
>AV780221
Length = 461
Score = 30.4 bits (67), Expect = 0.55
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = +1
Query: 2 FFGEGLVEGGKFVGLNGIRCVTRGTRVD*EFKM*SWST*ASWQNGNGVCSKK 53
F G EG K +G NG+R V TR F+ + ST W +G+G C ++
Sbjct: 112 FLGGRWREGRK*LG*NGVRFVDPRTRGVWGFRTWACSTRLFWASGDGGCLRR 267
>TC13049
Length = 604
Score = 28.1 bits (61), Expect = 2.8
Identities = 9/25 (36%), Positives = 17/25 (68%)
Frame = -2
Query: 158 IIFTYNQFWRKGRTRGEENQEKRLP 182
I+ ++N +WRKGR R ++ + +P
Sbjct: 531 IV*SFNGYWRKGRARPDQRKRSSVP 457
>TC17745
Length = 570
Score = 27.7 bits (60), Expect = 3.6
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = +2
Query: 36 SWST*ASWQNGNGVCSKKINLFGRGCWKINMVINVMPSLII*LF 79
SWS + + G G+ S+ +F R C KIN +I V +I *LF
Sbjct: 104 SWSFIRALEYGLGIWSQLPIVFRRSCEKINKIILV--GIIF*LF 229
>TC14768 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (21%)
Length = 1036
Score = 27.3 bits (59), Expect = 4.7
Identities = 14/24 (58%), Positives = 16/24 (66%)
Frame = -3
Query: 176 NQEKRLPNRNTKVRVGLTFNLLKP 199
N+EK+L NTKV LT NLL P
Sbjct: 692 NKEKQLHIPNTKVVPNLTINLLAP 621
>AV426745
Length = 420
Score = 26.6 bits (57), Expect = 8.0
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = +3
Query: 48 GVCSKKINLFGRGC 61
GVCSK+ FGRGC
Sbjct: 99 GVCSKRCGGFGRGC 140
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.369 0.170 0.610
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,403,255
Number of Sequences: 28460
Number of extensions: 81797
Number of successful extensions: 1144
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 844
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 292
Number of HSP's gapped (non-prelim): 879
length of query: 406
length of database: 4,897,600
effective HSP length: 92
effective length of query: 314
effective length of database: 2,279,280
effective search space: 715693920
effective search space used: 715693920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC139854.1


