

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
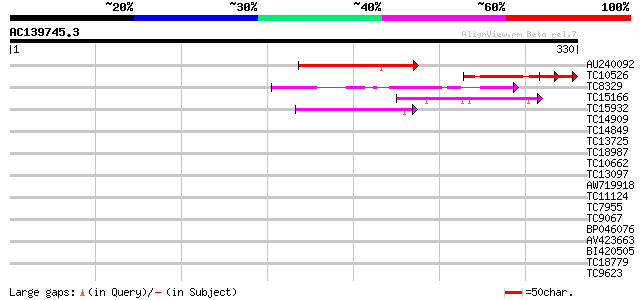
Query= AC139745.3 - phase: 0 /pseudo
(330 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU240092 62 1e-10
TC10526 homologue to UP|WR47_ARATH (Q9ZSI7) Probable WRKY transc... 57 4e-09
TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription fact... 56 1e-08
TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-bind... 51 2e-07
TC15932 similar to UP|WR61_ARATH (Q8VWV6) Probable WRKY transcri... 46 8e-06
TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcri... 39 0.001
TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%) 37 0.003
TC13725 similar to UP|Q39658 (Q39658) SPF1-like DNA-binding prot... 36 0.008
TC18987 similar to UP|Q8GYT1 (Q8GYT1) OBP33pep like protein, par... 34 0.030
TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CW... 33 0.066
TC13097 similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein (At2g413... 32 0.11
AW719918 32 0.15
TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding prot... 32 0.19
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 32 0.19
TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcrip... 31 0.25
BP046076 31 0.33
AV423663 30 0.56
BI420505 30 0.56
TC18779 similar to UP|AAS07149 (AAS07149) Expressed protein, par... 30 0.56
TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxid... 30 0.56
>AU240092
Length = 300
Score = 62.4 bits (150), Expect = 1e-10
Identities = 38/72 (52%), Positives = 44/72 (60%), Gaps = 2/72 (2%)
Frame = +2
Query: 169 HNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPY--ATMATSTLSAS 226
H+HPLPP A A+A TTSAAA+MLLS S S L + + A +TLSAS
Sbjct: 5 HSHPLPPTAQAMASTTSAAASMLLSGSMPSADHGLINPTILETASLHSSAQQNMATLSAS 184
Query: 227 QPFPTITLDFTQ 238
PFPTITLD TQ
Sbjct: 185 APFPTITLDLTQ 220
>TC10526 homologue to UP|WR47_ARATH (Q9ZSI7) Probable WRKY transcription
factor 47 (WRKY DNA-binding protein 47), partial (7%)
Length = 411
Score = 57.0 bits (136), Expect = 4e-09
Identities = 34/56 (60%), Positives = 41/56 (72%)
Frame = +2
Query: 265 LGQPPPSSMVESVSAAISSDPNFTTALAAAISSIIGPQRSGDGNNNLAGVVPGSPQ 320
LGQ PP+ VE++SAAI+SDPNFT ALAAAISSII R DG+NN + V + Q
Sbjct: 2 LGQRPPT--VETMSAAIASDPNFTAALAAAISSII---RGSDGHNNSSSDVNNNSQ 154
Score = 42.4 bits (98), Expect = 1e-04
Identities = 17/22 (77%), Positives = 20/22 (90%)
Frame = +3
Query: 309 NNLAGVVPGSPQLPQSCTTFST 330
NN + ++PGSPQLPQSCTTFST
Sbjct: 153 NNGSAIIPGSPQLPQSCTTFST 218
>TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription factor 1,
partial (41%)
Length = 1717
Score = 55.8 bits (133), Expect = 1e-08
Identities = 48/145 (33%), Positives = 66/145 (45%), Gaps = 1/145 (0%)
Frame = +1
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS 212
R +D+++L+ TYEG HNHP P A TS + R S G S
Sbjct: 700 RSVDDQSVLVATYEGEHNHPQPSQMEA----------------TSGSGRNVSLVG----S 819
Query: 213 FPYATMATSTLSASQPFP-TITLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPS 271
P S+ S S P P +TLD T++ + N P S KLP +L
Sbjct: 820 MP------SSKSLSSPAPAVVTLDLTKSRCSNDSKNAEPRK--DSAKLPQVL-------- 951
Query: 272 SMVESVSAAISSDPNFTTALAAAIS 296
VE ++ ++++DPNF AL AAIS
Sbjct: 952 --VEQMATSLTTDPNFRAALVAAIS 1020
>TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-binding protein
6 {Arabidopsis thaliana;} , partial (26%)
Length = 668
Score = 51.2 bits (121), Expect = 2e-07
Identities = 43/126 (34%), Positives = 53/126 (41%), Gaps = 41/126 (32%)
Frame = +3
Query: 226 SQPFPTITLDFTQNHN------------LSMHHNRVPLPLFFSHKLPPL-------LQLG 266
S PFPT+TLD T N N + N + P F+ P LQL
Sbjct: 3 SAPFPTVTLDLTHNPNPLQFSRPQHSAPFQIPQNFMSGPASFAQAAPLYNQSKFSGLQLS 182
Query: 267 -------------------QPPPSSMVESVSAAISSDPNFTTALAAAISSIIG---PQRS 304
QP + V + +AAI++DPNFT LAAAISSIIG +
Sbjct: 183 SQQEVGSSHQLASQQPPQQQPSLADTVSAATAAITADPNFTAVLAAAISSIIGGSHANNN 362
Query: 305 GDGNNN 310
G NNN
Sbjct: 363 GSHNNN 380
>TC15932 similar to UP|WR61_ARATH (Q8VWV6) Probable WRKY transcription
factor 61 (WRKY DNA-binding protein 61), partial (6%)
Length = 454
Score = 46.2 bits (108), Expect = 8e-06
Identities = 32/83 (38%), Positives = 42/83 (50%), Gaps = 12/83 (14%)
Frame = -2
Query: 167 GNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSAS 226
G HNHPLP +ATA+ +TSAA +ML S S +S A S S + + S+ + S
Sbjct: 336 GTHNHPLPVSATAMDSSTSAALSML*SGSYTSQPTGHCAASMGSASTMFIGLNFSSYNQS 157
Query: 227 QP------------FPTITLDFT 237
+ FPTITLD T
Sbjct: 156 RATQAFLPTPTPHLFPTITLDLT 88
>TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcription
factor 21 (WRKY DNA-binding protein 21), partial (22%)
Length = 590
Score = 38.9 bits (89), Expect = 0.001
Identities = 17/31 (54%), Positives = 20/31 (63%)
Frame = +3
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHT 183
RC E+ T+L+ TYEG HNHP P A A T
Sbjct: 177 RCLEEPTMLLVTYEGEHNHPKIPTQPANA*T 269
>TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%)
Length = 1389
Score = 37.4 bits (85), Expect = 0.003
Identities = 19/54 (35%), Positives = 30/54 (55%)
Frame = +2
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESAT 206
R + D +ITTYEG HNH +P A + H+ + L ++T++ +R S T
Sbjct: 518 RASHDLRAVITTYEGKHNHDVPAARGSGNHSVTRP----LPNNTANAIRPSSVT 667
Score = 26.6 bits (57), Expect = 6.2
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 1/69 (1%)
Frame = +2
Query: 157 DKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKES-ATGYLSNSFPY 215
D I Y+G HNHP P AT ++S++ + S+ S+ ++ +S AT
Sbjct: 11 DGQITEIVYKGTHNHP-KPQATRRNSSSSSSLPIPHSNHISNDIQDQSYATHGSGQMDSV 187
Query: 216 ATMATSTLS 224
AT S++S
Sbjct: 188 ATPENSSIS 214
>TC13725 similar to UP|Q39658 (Q39658) SPF1-like DNA-binding protein,
partial (11%)
Length = 578
Score = 36.2 bits (82), Expect = 0.008
Identities = 17/38 (44%), Positives = 25/38 (65%)
Frame = +2
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAM 190
R + D +ITTYEG HNH + PAA +H T+++ +M
Sbjct: 29 RASTDAKAVITTYEGKHNHDV-PAARNSSHNTASSNSM 139
>TC18987 similar to UP|Q8GYT1 (Q8GYT1) OBP33pep like protein, partial (48%)
Length = 367
Score = 34.3 bits (77), Expect = 0.030
Identities = 28/90 (31%), Positives = 43/90 (47%), Gaps = 2/90 (2%)
Frame = +2
Query: 169 HNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMAT--STLSAS 226
H+H PP + + + ++SA LS++ + SAT LS + P T +T + +SA
Sbjct: 98 HSHEAPPPSASRSSSSSAGETSTLSAAMAPPTTSASATSTLSKT-PATTSSTRGTVISAV 274
Query: 227 QPFPTITLDFTQNHNLSMHHNRVPLPLFFS 256
PT +LS + V PLFFS
Sbjct: 275 SRSPTTL-------SLSTTNLSVRTPLFFS 343
>TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CWP1
precursor, partial (6%)
Length = 937
Score = 33.1 bits (74), Expect = 0.066
Identities = 30/114 (26%), Positives = 49/114 (42%)
Frame = +2
Query: 173 LPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTI 232
+P + + + +T++ A++ LSSS+SST S + SFP A+ +S S + F +
Sbjct: 74 MPSDSESESESTTSIASLTLSSSSSSTRSPTSKP*VAAASFPAASTPSSLKSKTSSFAST 253
Query: 233 TLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSSMVESVSAAISSDPN 286
T H LP PP PS + S+A S +P+
Sbjct: 254 ASSPTTTH----------LP-------PPPPPTNPAAPSGTSSASSSAASPNPS 364
>TC13097 similar to UP|Q9ZVB8 (Q9ZVB8) At2g41330 protein
(At2g41330/F13H10.12), partial (7%)
Length = 660
Score = 32.3 bits (72), Expect = 0.11
Identities = 38/128 (29%), Positives = 53/128 (40%), Gaps = 9/128 (7%)
Frame = -2
Query: 154 CAEDKTILITTYEGNHN--HPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSN 211
C+ KT TT N N +P P A+ A + ++S+ +A SSS+ S S +
Sbjct: 395 CSASKT---TTMVANRNPENPSPSASLASSSSSSSFSASSSSSSSKSQCSNSSPKPPRAL 225
Query: 212 SFPYATMATSTLSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHK---LPPLL----Q 264
S P + ++L T +L M +VP FSHK L PLL Q
Sbjct: 224 SLPMPLVHHPPTKKGDTHHLVSLTSTTYGSLLMIDQKVPK---FSHKDPHLEPLLTKSFQ 54
Query: 265 LGQPPPSS 272
G P S
Sbjct: 53 KGSNPKES 30
>AW719918
Length = 506
Score = 32.0 bits (71), Expect = 0.15
Identities = 29/85 (34%), Positives = 37/85 (43%), Gaps = 16/85 (18%)
Frame = +1
Query: 166 EGNHNH----PL-------PPAATAIAHTTSAAAAMLLSSSTSSTLRK-----ESATGYL 209
+ HNH PL PP +TA + TTS AA+ +S TSS S T
Sbjct: 94 QNRHNHQWKPPLNSPHLLHPPPSTARSATTSLAASSRSASGTSSPFPATST*LSSTTSSA 273
Query: 210 SNSFPYATMATSTLSASQPFPTITL 234
S S + ATS A+ P T+ L
Sbjct: 274 SLSSTSSAAATSLTPATPPTATLVL 348
>TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding protein,
partial (45%)
Length = 517
Score = 31.6 bits (70), Expect = 0.19
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +1
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAH 182
R +D+ +++TTYEG H HP+ H
Sbjct: 112 RLTQDEGVVVTTYEGVHTHPIEKTTDNFEH 201
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 31.6 bits (70), Expect = 0.19
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = +3
Query: 153 RCAEDKTILITTYEGNHNHPLPPA 176
R +D T+LI TYEG H H L A
Sbjct: 207 RATDDPTMLIVTYEGEHRHALQAA 278
>TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcription
factor 15 (WRKY DNA-binding protein 15), partial (31%)
Length = 660
Score = 31.2 bits (69), Expect = 0.25
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +3
Query: 153 RCAEDKTILITTYEGNHNHPLPPA 176
R +D +L+ TYEG HNH L A
Sbjct: 240 RALDDAAMLVVTYEGEHNHALSAA 311
>BP046076
Length = 593
Score = 30.8 bits (68), Expect = 0.33
Identities = 12/19 (63%), Positives = 15/19 (78%)
Frame = -2
Query: 153 RCAEDKTILITTYEGNHNH 171
R +ED ++ITTYEG HNH
Sbjct: 343 RLSEDCRMVITTYEGRHNH 287
>AV423663
Length = 447
Score = 30.0 bits (66), Expect = 0.56
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +1
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTT 184
R + + I TY G HNHP P ++A +T
Sbjct: 196 RNKSEPAMFIVTYTGEHNHPAPTHRNSLAGST 291
>BI420505
Length = 474
Score = 30.0 bits (66), Expect = 0.56
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = -1
Query: 223 LSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHKLPP 261
LSA+ P T+ LD+ + HH++ PL + H PP
Sbjct: 318 LSANPPTQTLCLDYQKTPTS*PHHHQPPLHSWHHHSPPP 202
>TC18779 similar to UP|AAS07149 (AAS07149) Expressed protein, partial (24%)
Length = 640
Score = 30.0 bits (66), Expect = 0.56
Identities = 20/53 (37%), Positives = 27/53 (50%)
Frame = +2
Query: 220 TSTLSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSS 272
T+T S SQP P TL ++ + + R P F +PP L L +PPP S
Sbjct: 101 TTTQSPSQPDPVNTLHLLRSSSTA----RSASP--FVQNMPPRLLLPEPPPQS 241
>TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxidase,
partial (42%)
Length = 640
Score = 30.0 bits (66), Expect = 0.56
Identities = 20/55 (36%), Positives = 32/55 (57%)
Frame = +3
Query: 172 PLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSAS 226
P P + T +A AAA M+ S++T +TL S++ Y S SF + ++S+ S S
Sbjct: 30 PFPSSPTKMAALGGAAARMIPSTATRATL-SLSSSSYSSRSFFSFSSSSSSSSRS 191
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.126 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,018,132
Number of Sequences: 28460
Number of extensions: 95404
Number of successful extensions: 806
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 783
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 800
length of query: 330
length of database: 4,897,600
effective HSP length: 91
effective length of query: 239
effective length of database: 2,307,740
effective search space: 551549860
effective search space used: 551549860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC139745.3


