

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
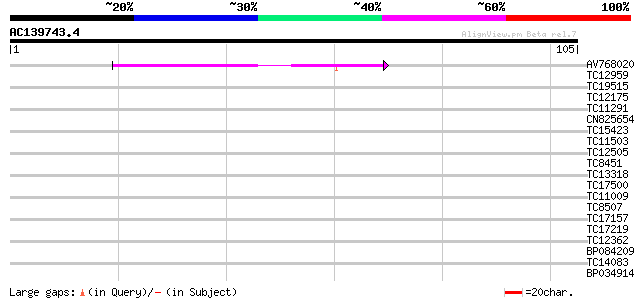
Query= AC139743.4 - phase: 0 /pseudo
(105 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV768020 40 6e-05
TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%) 32 0.022
TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, pa... 30 0.082
TC12175 similar to GB|AAA29488.1|552180|PFAANTPA parasite protei... 29 0.11
TC11291 similar to GB|BAA24377.1|2780775|AB010246 Ring3 {Mus mus... 29 0.11
CN825654 28 0.18
TC15423 28 0.18
TC11503 similar to GB|AAO63311.1|28950775|BT005247 At3g02220 {Ar... 28 0.24
TC12505 similar to UP|BAD00698 (BAD00698) Sericin 1A, partial (5%) 28 0.24
TC8451 GB|CAA61589.1|897771|LJAS1GENE asparagine synthase (gluta... 28 0.31
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 28 0.31
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 27 0.41
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 27 0.53
TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 27 0.53
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 27 0.53
TC17219 similar to UP|Q9SJ68 (Q9SJ68) Expressed protein, partial... 27 0.53
TC12362 weakly similar to UP|UBF1_HUMAN (P17480) Nucleolar trans... 27 0.53
BP084209 27 0.53
TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor... 27 0.53
BP034914 27 0.70
>AV768020
Length = 268
Score = 40.0 bits (92), Expect = 6e-05
Identities = 28/57 (49%), Positives = 33/57 (57%), Gaps = 6/57 (10%)
Frame = +3
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSM------DENEAESKYNG 70
+A + ++ + CSTTSSSSIGRNSD VSSERSM ENEA Y G
Sbjct: 114 EAAANVSPEEEEQCSTTSSSSIGRNSD------VSSERSMGDDDEGGENEA*CAYCG 266
>TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%)
Length = 517
Score = 31.6 bits (70), Expect = 0.022
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +1
Query: 38 SSSIGRNSDDDDDDEVSSERSMDENE 63
S S G +SDDDDDD+ S + D+NE
Sbjct: 106 SLSDGSSSDDDDDDDDSEDEDDDDNE 183
Score = 23.1 bits (48), Expect = 7.7
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +1
Query: 44 NSDDDDDDEVSSERSMDENEAE 65
+S DDDDD+ SE D++ +
Sbjct: 121 SSSDDDDDDDDSEDEDDDDNED 186
Score = 23.1 bits (48), Expect = 7.7
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +1
Query: 38 SSSIGRNSDDDDDDEVSSERSMDENE 63
SSS + DDD +DE + D++E
Sbjct: 121 SSSDDDDDDDDSEDEDDDDNEDDDDE 198
>TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, partial (3%)
Length = 504
Score = 29.6 bits (65), Expect = 0.082
Identities = 16/48 (33%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Frame = +3
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDD--DDEVSSERSMDENEAE 65
D D DDD G +GR SD DD D+ S +D+ +A+
Sbjct: 93 DEEDDDDDDDGDDEDPLLGPGLGRESDSDDMSHDDYDSASDIDDEDAD 236
>TC12175 similar to GB|AAA29488.1|552180|PFAANTPA parasite protein
{Plasmodium falciparum;} , partial (7%)
Length = 609
Score = 29.3 bits (64), Expect = 0.11
Identities = 19/60 (31%), Positives = 26/60 (42%)
Frame = +2
Query: 18 LFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCME 77
L + R +G+ T G SDDDDDD+ E ++ +E K GG D E
Sbjct: 200 LHECLRTGASPRGRFKFPTDVLFYGHISDDDDDDD-EEESEVESDECHGKL*GGCEDAGE 376
>TC11291 similar to GB|BAA24377.1|2780775|AB010246 Ring3 {Mus musculus;} ,
partial (4%)
Length = 326
Score = 29.3 bits (64), Expect = 0.11
Identities = 28/82 (34%), Positives = 36/82 (43%), Gaps = 1/82 (1%)
Frame = +3
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTS-SSSIGRNSDDDDDDEVSSERSM 59
M L + +G R S +D+D + S S +SSIGRNS DD S
Sbjct: 87 MPLALETDRGLGLRRSRFIPCDSIYDEDIPEEDSDCSDASSIGRNSVSSDDH--SDREDS 260
Query: 60 DENEAESKYNGGALDCMEALEE 81
E E +S + LD M LEE
Sbjct: 261 GEVEEQSTFK-SPLDTMNDLEE 323
>CN825654
Length = 580
Score = 28.5 bits (62), Expect = 0.18
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 30 GKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAE 65
G V S +S +G D+DDD+E S DE + +
Sbjct: 26 GSVFSASSFGLLGDGEDEDDDEEKSELTGEDEEDEQ 133
>TC15423
Length = 638
Score = 28.5 bits (62), Expect = 0.18
Identities = 21/54 (38%), Positives = 28/54 (50%), Gaps = 4/54 (7%)
Frame = +1
Query: 37 SSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG----GALDCMEALEEVLPIR 86
SSSSIG D ++D+E E +S G G+LDC LE+ LPI+
Sbjct: 475 SSSSIGTPDDSEEDEE----------EVQSGLKGRTGLGSLDC---LEDSLPIK 597
>TC11503 similar to GB|AAO63311.1|28950775|BT005247 At3g02220 {Arabidopsis
thaliana;}, partial (60%)
Length = 975
Score = 28.1 bits (61), Expect = 0.24
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +2
Query: 25 HDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAE 65
HDD G+VC +S N D D+D+E D+ A+
Sbjct: 584 HDDCDGEVCDDEDDNS-SENKDSDEDNEDHEADVPDQMNAQ 703
>TC12505 similar to UP|BAD00698 (BAD00698) Sericin 1A, partial (5%)
Length = 569
Score = 28.1 bits (61), Expect = 0.24
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = -3
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDEN 62
D DDD +V ++SS + DDD D+ E DE+
Sbjct: 459 DDDDDDVQVLPAAATSSHPKRKRDDDADDREEEDEDDED 343
Score = 26.6 bits (57), Expect = 0.70
Identities = 15/44 (34%), Positives = 21/44 (47%)
Frame = -3
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESK 67
D DDD + + +SS R DDD DD + ++ A SK
Sbjct: 456 DDDDDVQVLPAAATSSHPKRKRDDDADDREEEDEDDEDVVAFSK 325
>TC8451 GB|CAA61589.1|897771|LJAS1GENE asparagine synthase
(glutamine-hydrolysing) {Lotus corniculatus var.
japonicus;} , complete
Length = 2148
Score = 27.7 bits (60), Expect = 0.31
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +1
Query: 40 SIGRNSDDDDDDEVSSE 56
++ RN DDDDDDE E
Sbjct: 1867 AVARNDDDDDDDEAKHE 1917
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 27.7 bits (60), Expect = 0.31
Identities = 18/48 (37%), Positives = 20/48 (41%)
Frame = +3
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESK 67
D D DDD G+ DDDDDDE E +E E E K
Sbjct: 300 DDDDDDDDDDGE-----------EEEDDDDDDE--EEEGSEEVEVEGK 404
Score = 27.3 bits (59), Expect = 0.41
Identities = 15/59 (25%), Positives = 24/59 (40%)
Frame = +3
Query: 12 GSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
G V D + DDD+ + DDDDDD+ E D+++ + + G
Sbjct: 222 GPPKPVQTDNEEEEDDDE-------EEDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEG 377
Score = 23.5 bits (49), Expect = 5.9
Identities = 10/46 (21%), Positives = 21/46 (44%)
Frame = +3
Query: 22 PRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESK 67
P + D K T + + ++DD+DE + D+++ E +
Sbjct: 204 PEIYPDGPPKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDGEEE 341
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 27.3 bits (59), Expect = 0.41
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = -2
Query: 15 TSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDE 52
+S+ F P + GK T SS R+S DDDDDE
Sbjct: 439 SSISFLLPTTETEPYGK---TPSSRFRSRSSSDDDDDE 335
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 26.9 bits (58), Expect = 0.53
Identities = 16/55 (29%), Positives = 22/55 (39%), Gaps = 6/55 (10%)
Frame = +2
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRN------SDDDDDDEVSSERSMDENEAESKY 68
DAP DDD + G DDDDDD+ E +E + ++Y
Sbjct: 83 DAPGGGDDDDDEEDEEEEGGVEGGRGGGGDPDDDDDDDDDDDEEEEEEEDLGTEY 247
Score = 26.2 bits (56), Expect = 0.91
Identities = 17/59 (28%), Positives = 27/59 (44%)
Frame = +2
Query: 13 SRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
SRT + D + DDD G+ + D+DDDD+ D++E + + GG
Sbjct: 5 SRTRMSGD---EDDDDDGE----------DDDGDEDDDDDAPGGGDDDDDEEDEEEEGG 142
Score = 25.4 bits (54), Expect = 1.6
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +2
Query: 43 RNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEE 81
R S D+DDD+ + DE++ + GG D E EE
Sbjct: 14 RMSGDEDDDDDGEDDDGDEDDDDDAPGGGDDDDDEEDEE 130
>TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (40%)
Length = 620
Score = 26.9 bits (58), Expect = 0.53
Identities = 9/23 (39%), Positives = 18/23 (78%)
Frame = +3
Query: 44 NSDDDDDDEVSSERSMDENEAES 66
+SDD+DD++ S+ MD+ +++S
Sbjct: 15 DSDDEDDEDDDSDEEMDDADSDS 83
Score = 24.6 bits (52), Expect = 2.6
Identities = 14/44 (31%), Positives = 19/44 (42%)
Frame = +3
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENE 63
D D DDD + S S SD DD DE + + D+ +
Sbjct: 21 DDEDDEDDDSDEEMDDADSDS--DESDSDDTDEETPVKKADQGK 146
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 26.9 bits (58), Expect = 0.53
Identities = 15/49 (30%), Positives = 20/49 (40%)
Frame = +2
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGA 72
+ DD+ G DDDDDDE +E E +SK G+
Sbjct: 773 EEDDEDGDEDEDEDDDDDEEEEDDDDDDE-----DDEEGEGKSKSKSGS 904
>TC17219 similar to UP|Q9SJ68 (Q9SJ68) Expressed protein, partial (80%)
Length = 883
Score = 26.9 bits (58), Expect = 0.53
Identities = 13/21 (61%), Positives = 16/21 (75%), Gaps = 1/21 (4%)
Frame = -1
Query: 37 SSSSIGRN-SDDDDDDEVSSE 56
S SS G + +DDDDDDE+ SE
Sbjct: 166 SFSSFGSSIADDDDDDEIDSE 104
>TC12362 weakly similar to UP|UBF1_HUMAN (P17480) Nucleolar transcription
factor 1 (Upstream binding factor 1) (UBF-1)
(Autoantigen NOR-90), partial (3%)
Length = 493
Score = 26.9 bits (58), Expect = 0.53
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = -1
Query: 13 SRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDEN 62
S TSV+ + D D + CS+ SSSS ++S SS S D +
Sbjct: 211 SLTSVV*EECVDESDFSAR*CSSISSSSSSKSSSSKSRSMSSSSSSADSS 62
>BP084209
Length = 502
Score = 26.9 bits (58), Expect = 0.53
Identities = 14/41 (34%), Positives = 22/41 (53%), Gaps = 5/41 (12%)
Frame = -3
Query: 35 TTSSSSIGRNSDDDDDDEVSSER-----SMDENEAESKYNG 70
T +S S R +DDDDDE+ + + S++ S +NG
Sbjct: 485 TFTSRSSARYKEDDDDDEMPNRKRRGVQSVEPKPHRSNFNG 363
>TC14083 similar to UP|CRTC_RICCO (P93508) Calreticulin precursor, partial
(92%)
Length = 1693
Score = 26.9 bits (58), Expect = 0.53
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = +1
Query: 47 DDDDDEVSSERSMDENEAESKYNGGALDCMEALEE 81
D +D+E + E S D ++AESK G D +A EE
Sbjct: 1252 DAEDEEDTDEASHDSDDAESKTEAGE-DSADANEE 1353
>BP034914
Length = 544
Score = 26.6 bits (57), Expect = 0.70
Identities = 16/48 (33%), Positives = 21/48 (43%)
Frame = +3
Query: 25 HDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGA 72
HDD + SSSS + D D D S R E E + +GG+
Sbjct: 54 HDDTP-----SPSSSSSSSTASDSDSDSDRSSRRRREKERRRRSDGGS 182
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.129 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,467,167
Number of Sequences: 28460
Number of extensions: 15800
Number of successful extensions: 314
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 187
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 252
length of query: 105
length of database: 4,897,600
effective HSP length: 81
effective length of query: 24
effective length of database: 2,592,340
effective search space: 62216160
effective search space used: 62216160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 47 (22.7 bits)
Medicago: description of AC139743.4


